合成生物学 ›› 2023, Vol. 4 ›› Issue (5): 966-979.DOI: 10.12211/2096-8280.2023-033
微液滴高通量筛选方法的研究与应用进展
秦伟彤, 杨广宇
- 上海交通大学生命科学技术学院,微生物代谢国家重点实验室,上海 200240
-
收稿日期:2023-04-24修回日期:2023-06-20出版日期:2023-10-31发布日期:2023-11-15 -
通讯作者:杨广宇 -
作者简介:秦伟彤 (1992—),女,博士。研究方向为酶的定向进化、微液滴超高通量筛选方法的建立。E-mail:qinweitong122@163.com杨广宇 (1980—),男,研究员,博士生导师。研究方向为酶分子改造、超高通量筛选方法的建立;酶分子机制解析;体外合成生物学研究。E-mail:yanggy@sjtu.edu.cn -
基金资助:国家重点研发计划(2018YFE0200501);国家自然科学基金(32030063);广东省重点领域研发计划(2022B1111050001);天津市合成生物技术创新能力提升行动;上海交通大学特区计划(21TQ1400210)
Research and application progress of microdroplets high throughput screening methods
QIN Weitong, YANG Guangyu
- State Key Laboratory of Microbial Metabolism,School of Life Science and Technology,Shanghai Jiaotong University,Shanghai 200240,China
-
Received:2023-04-24Revised:2023-06-20Online:2023-10-31Published:2023-11-15 -
Contact:YANG Guangyu
摘要:
在单细胞层面对生物功能进行高通量的分析和分选是对关键基因、元件、途径与细胞工厂进行优化的重要技术。基于微液滴的筛选方法因其低成本、超高通量等优势,已被广泛应用于生物、医药、食品和工业等各个领域。本文针对目前主流的荧光激活的液滴分选、吸光度激活的液滴分选,以及无标记液滴分选等微液滴筛选设备的进展进行综述,主要包括基于质谱、拉曼、核磁共振、电化学、图像识别等。并总结了近年来微液滴筛选设备在酶进化、微生物育种等领域应用成功的案例。此外还对不同的微液滴筛选设备的优势与面临的挑战进行了讨论,未来各种新的荧光探针的开发以及质谱等非标记检测方法的进一步发展,将是微液滴筛选设备的主要发展方向,在蛋白质工程、抗体工程、细胞分选及临床研究等方面具有重要的应用潜力。
中图分类号:
引用本文
秦伟彤, 杨广宇. 微液滴高通量筛选方法的研究与应用进展[J]. 合成生物学, 2023, 4(5): 966-979.
QIN Weitong, YANG Guangyu. Research and application progress of microdroplets high throughput screening methods[J]. Synthetic Biology Journal, 2023, 4(5): 966-979.

图2 标记液滴分选技术分选原理FADS—荧光激活的液滴分选技术;AADS—吸光度激活的液滴分选技术;S—底物;P—产物;E—酶分子;Ex—发射光谱;Em—吸收光谱
Fig. 2 The principle of labeled droplet sorting technologyFADS—Fluorescence activated droplet sorting technology; AADS—Absorbance activated droplet separation technology; S—Substrate; P—Product; E: Enzyme molecules; Ex—Emission spectrum; Em—Absorption spectrum
| 分选方法 | 最高分选效率 | 最高分选灵敏度 | 最小液滴体积 | 优点 | 缺点 |
|---|---|---|---|---|---|
| FADS | 5 kHz[ | 2.5 nmol/L[ | 2 pL[ | 检测灵敏度高、分选速度快、平台发展成熟 | 大部分检测靶标缺乏适合的荧光耦联方法 |
| AADS | 1 kHz[ | 10 μmol/L[ | 100 pL[ | 普适性较FADS高 | 检测灵敏度待提高 |
| MADS | 35 Hz[ | 5 μmol/L[ | 0.8 nL[ | 无损伤、普适性高 | 分选速度慢、灵敏度待提高 |
| RADS | 4.3 Hz[ | 50 μmol/L[ | 65 pL[ | 无损伤、普适性高 | 更适用于较大的细胞 |
| NMR-ADS | — | 1 mmol/L[ | 130 pL[ | 无损伤、提供信息广泛 | NMR与液滴分选系统的整合较困难、检测灵敏度低 |
| IBDS | 10 Hz[ | — | 35 pL[ | 无损伤 | 适用范围窄、分选速度慢 |
| EADS | 10 Hz[ | 1 μmol/L[ | 30 nL[ | 无损伤 | 适用范围窄、分选速度慢 |
表1 不同微流控分选设备比较
Table 1 Comparison of different microfluidic sorting equipment
| 分选方法 | 最高分选效率 | 最高分选灵敏度 | 最小液滴体积 | 优点 | 缺点 |
|---|---|---|---|---|---|
| FADS | 5 kHz[ | 2.5 nmol/L[ | 2 pL[ | 检测灵敏度高、分选速度快、平台发展成熟 | 大部分检测靶标缺乏适合的荧光耦联方法 |
| AADS | 1 kHz[ | 10 μmol/L[ | 100 pL[ | 普适性较FADS高 | 检测灵敏度待提高 |
| MADS | 35 Hz[ | 5 μmol/L[ | 0.8 nL[ | 无损伤、普适性高 | 分选速度慢、灵敏度待提高 |
| RADS | 4.3 Hz[ | 50 μmol/L[ | 65 pL[ | 无损伤、普适性高 | 更适用于较大的细胞 |
| NMR-ADS | — | 1 mmol/L[ | 130 pL[ | 无损伤、提供信息广泛 | NMR与液滴分选系统的整合较困难、检测灵敏度低 |
| IBDS | 10 Hz[ | — | 35 pL[ | 无损伤 | 适用范围窄、分选速度慢 |
| EADS | 10 Hz[ | 1 μmol/L[ | 30 nL[ | 无损伤 | 适用范围窄、分选速度慢 |

图3 无标记液滴分选技术分选原理RE—参考电极;WE—工作电极;CE—对电极
Fig. 3 The principle of unlabeled droplet sorting technologyRE—Reference electrode; WE—Working electrode; CE—Counter electrode
| 发表时间 | 分选系统 | 目标酶 | 分选结果 | 参考文献 |
|---|---|---|---|---|
| 2018 | FADS | 酯酶 | 对S-布洛芬的对映选择性提高600倍 | [ |
| 2019 | FADS | 硫酸酯酶 | Kcat/Km值提高30倍 | [ |
| 2019 | FADS | 纤维素酶 | 筛选出产量提升46%的高纤维素酶菌株 | [ |
| 2020 | FADS | 葡萄糖氧化酶 | Kcat值比野生型高2.1倍 | [ |
| 2020 | AADS | 胺脱氢酶 | 转化率提高3.3倍 | [ |
| 2022 | FADS | α-淀粉酶 | 产量提升50%的地衣芽孢杆菌突变株 | [ |
| 2023 | FADS | 二乙酰壳二糖脱乙酰酶 | 催化效率提高1.8倍 | [ |
| 2022 | FADS | 塑料降解酶 | 2株可降解塑料的菌株 | [ |
| 2022 | FADS | 产鼠李糖脂的微生物 | 产量提升54%~208%的菌株 | [ |
| 2022 | AADS | 葡萄糖脱氢酶 | 催化速度和效率提升10倍以上 | [ |
表2 近五年微流控分选装置成功应用的案例
Table 2 Cases of successful application of microfluidic sorting devices in the past five years
| 发表时间 | 分选系统 | 目标酶 | 分选结果 | 参考文献 |
|---|---|---|---|---|
| 2018 | FADS | 酯酶 | 对S-布洛芬的对映选择性提高600倍 | [ |
| 2019 | FADS | 硫酸酯酶 | Kcat/Km值提高30倍 | [ |
| 2019 | FADS | 纤维素酶 | 筛选出产量提升46%的高纤维素酶菌株 | [ |
| 2020 | FADS | 葡萄糖氧化酶 | Kcat值比野生型高2.1倍 | [ |
| 2020 | AADS | 胺脱氢酶 | 转化率提高3.3倍 | [ |
| 2022 | FADS | α-淀粉酶 | 产量提升50%的地衣芽孢杆菌突变株 | [ |
| 2023 | FADS | 二乙酰壳二糖脱乙酰酶 | 催化效率提高1.8倍 | [ |
| 2022 | FADS | 塑料降解酶 | 2株可降解塑料的菌株 | [ |
| 2022 | FADS | 产鼠李糖脂的微生物 | 产量提升54%~208%的菌株 | [ |
| 2022 | AADS | 葡萄糖脱氢酶 | 催化速度和效率提升10倍以上 | [ |
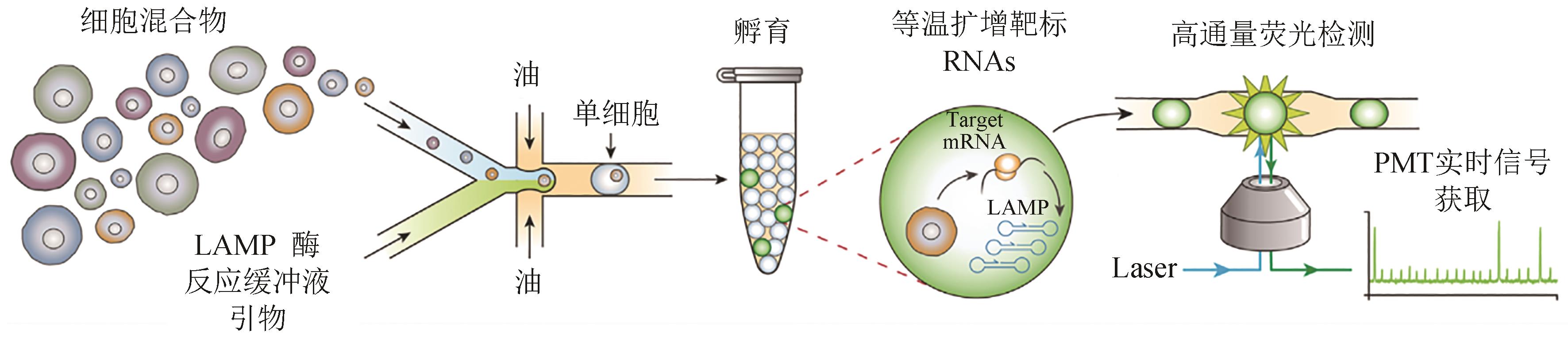
图4 SNAPD工作流程图[96](将单细胞和分析试剂共包裹到微滴中,收集并体外孵育,随后测量每个液滴的荧光以指示靶RNA的扩增)
Fig. 4 Schematic of the SNAPD workflow[96](Single cells are encapsulated into microdroplets with assay reagents, collected and incubated offline, and the fluorescence of each droplet is subsequently measured to indicate amplification of target RNAs)
| 1 | ZENG W Z, GUO L K, XU S, et al. High-throughput screening technology in industrial biotechnology[J]. Trends in Biotechnology, 2020, 38(8): 888-906. |
| 2 | XIONG W, LIU B, SHEN Y J, et al. Protein engineering design from directed evolution to de novo synthesis[J]. Biochemical Engineering Journal, 2021, 174: 108096. |
| 3 | LONGWELL C K, LABANIEH L, COCHRAN J R. High-throughput screening technologies for enzyme engineering[J]. Current Opinion in Biotechnology, 2017, 48: 196-202. |
| 4 | MACARRON R, BANKS M N, BOJANIC D, et al. Impact of high-throughput screening in biomedical research[J]. Nature Reviews Drug Discovery, 2011, 10(3): 188-195. |
| 5 | QIN W T, LI L, YANG F, et al. High-throughput iSpinach fluorescent aptamer-based real-time monitoring of in vitro transcription[J].Bioresources and Bioprocessing, 2022, 9: 112. |
| 6 | LLOYD M D. High-throughput screening for the discovery of enzyme inhibitors[J]. Journal of Medicinal Chemistry, 2020, 63(19): 10742-10772. |
| 7 | SARNAIK A, LIU A, NIELSEN D, et al. High-throughput screening for efficient microbial biotechnology[J]. Current Opinion in Biotechnology, 2020, 64: 141-150. |
| 8 | CAEN O, SCHÜTZ S, JAMMALAMADAKA M S S, et al. High-throughput multiplexed fluorescence-activated droplet sorting[J]. Microsystems & Nanoengineering, 2018, 4: 33. |
| 9 | POTYRAILO R, RAJAN K, STOEWE K, et al. Combinatorial and high-throughput screening of materials libraries: review of state of the art[J]. ACS Combinatorial Science, 2011, 13(6): 579-633. |
| 10 | CHIN C D, LINDER V, SIA S K. Lab-on-a-chip devices for global health: past studies and future opportunities[J]. Lab on a Chip, 2007, 7(1): 41-57. |
| 11 | AZIZIPOUR N, AVAZPOUR R, ROSENZWEIG D H, et al. Evolution of biochip technology: a review from lab-on-a-chip to organ-on-a-chip[J]. Micromachines, 2020, 11(6): 599. |
| 12 | BARET J C, MILLER O J, TALY V, et al. Fluorescence-activated droplet sorting (FADS): efficient microfluidic cell sorting based on enzymatic activity[J]. Lab on a Chip, 2009, 9(13): 1850-1858. |
| 13 | TU R, ZHANG Y, HUA E B, et al. Droplet-based microfluidic platform for high-throughput screening of Streptomyces[J]. Communications Biology, 2021, 4: 647. |
| 14 | SJOSTROM S L, BAI Y P, HUANG M T, et al. High-throughput screening for industrial enzyme production hosts by droplet microfluidics[J]. Lab on a Chip, 2014, 14(4): 806-813. |
| 15 | FU X Z, ZHANG Y Y, XU Q, et al. Recent advances on sorting methods of high-throughput droplet-based microfluidics in enzyme directed evolution[J]. Frontiers in Chemistry, 2021, 9: 666867. |
| 16 | OBEXER R, POTT M, ZEYMER C, et al. Efficient laboratory evolution of computationally designed enzymes with low starting activities using fluorescence-activated droplet sorting[J]. Protein Engineering, Design and Selection, 2017, 30(7): 531. |
| 17 | MA F Q, CHUNG M T, YAO Y, et al. Efficient molecular evolution to generate enantioselective enzymes using a dual-channel microfluidic droplet screening platform[J]. Nature Communications, 2018, 9: 1030. |
| 18 | TIEMEIJER B M, DESCAMPS L, HULLEMAN J, et al. A microfluidic approach for probing heterogeneity in cytotoxic T-cells by cell pairing in hydrogel droplets[J]. Micromachines, 2022, 13(11): 1910. |
| 19 | WU L, CHEN P, DONG Y S, et al. Encapsulation of single cells on a microfluidic device integrating droplet generation with fluorescence-activated droplet sorting[J]. Biomedical Microdevices, 2013, 15(3): 553-560. |
| 20 | SUN G Y, QU L S, AZI F, et al. Recent progress in high-throughput droplet screening and sorting for bioanalysis[J]. Biosensors and Bioelectronics, 2023, 225: 115107. |
| 21 | GIELEN F, HOURS R, EMOND S, et al. Ultrahigh-throughput-directed enzyme evolution by absorbance-activated droplet sorting (AADS)[J]. Proceedings of the National Academy of Sciences of the United States of America, 2016, 113(47): E7383-E7389. |
| 22 | LINK D R, GRASLAND-MONGRAIN E, DURI A, et al. Electric control of droplets in microfluidic devices[J]. Angewandte Chemie International Edition, 2006, 45(16): 2556-2560. |
| 23 | DIEFENBACH X W, FARASAT I, GUETSCHOW E D, et al. Enabling biocatalysis by high-throughput protein engineering using droplet microfluidics coupled to mass spectrometry[J]. ACS Omega, 2018, 3(2): 1498-1508. |
| 24 | LEE K S, PALATINSZKY M, PEREIRA F C, et al. An automated Raman-based platform for the sorting of live cells by functional properties[J]. Nature Microbiology, 2019, 4(6): 1035-1048. |
| 25 | GOTO H, KANAI Y, YOTSUI A, et al. Microfluidic screening system based on boron-doped diamond electrodes and dielectrophoretic sorting for directed evolution of NAD(P)-dependent oxidoreductases[J]. Lab on a Chip, 2020, 20(4): 852-861. |
| 26 | DAVOODI H, NORDIN N, BORDONALI L, et al. An NMR-compatible microfluidic platform enabling in situ electrochemistry[J]. Lab on a Chip, 2020, 20(17): 3202-3212. |
| 27 | LABELLE C A, MASSARO A, CORTÉS-LLANOS B, et al. Image-based live cell sorting[J]. Trends in Biotechnology, 2021, 39(6): 613-623. |
| 28 | SCIAMBI A, ABATE A R. Accurate microfluidic sorting of droplets at 30 kHz[J]. Lab on a Chip, 2015, 15(1): 47-51. |
| 29 | LI S X, DING X Y, GUO F, et al. An on-chip, multichannel droplet sorter using standing surface acoustic waves[J]. Analytical Chemistry, 2013, 85(11): 5468-5474. |
| 30 | NAVI M, ABBASI N, SALARI A, et al. Magnetic water-in-water droplet microfluidics: systematic experiments and scaling mathematical analysis[J]. Biomicrofluidics, 2020, 14(2): 024101. |
| 31 | ZHONG R Y, YANG S J, STEFANO UGOLINI G, et al. Acoustofluidic droplet sorter based on single phase focused transducers (small 46/2021)[J]. Small, 2021, 17(46): e2103848. |
| 32 | ROBERT DE SAINT VINCENT M, WUNENBURGER R, DELVILLE J P. Laser switching and sorting for high speed digital microfluidics[J]. Applied Physics Letters, 2008, 92(15): 154105. |
| 33 | QIAO Y X, ZHAO X Y, ZHU J, et al. Fluorescence-activated droplet sorting of lipolytic microorganisms using a compact optical system[J]. Lab on a Chip, 2018, 18(1): 190-196. |
| 34 | TABUCHI T, YOKOBAYASHI Y. High-throughput screening of cell-free riboswitches by fluorescence-activated droplet sorting[J]. Nucleic Acids Research, 2022, 50(6): 3535-3550. |
| 35 | BECKER S. Ultra-high-throughput screening based on cell-surface display and fluorescence-activated cell sorting for the identification of novel biocatalysts[J]. Current Opinion in Biotechnology, 2004, 15(4): 323-329. |
| 36 | BASU S, CAMPBELL H M, DITTEL B N, et al. Purification of specific cell population by fluorescence activated cell sorting (FACS)[J]. Journal of Visualized Experiments: JoVE, 2010(41): 1546. |
| 37 | YUAN H L, TU R, TONG X W, et al. Ultrahigh-throughput screening of industrial enzyme-producing strains by droplet-based microfluidic system[J]. Journal of Industrial Microbiology and Biotechnology, 2022, 49(3): kuac007. |
| 38 | LARSEN A C, DUNN M R, HATCH A, et al. A general strategy for expanding polymerase function by droplet microfluidics[J]. Nature Communications, 2016, 7: 11235. |
| 39 | HASAN S, GEISSLER D, WINK K, et al. Fluorescence lifetime-activated droplet sorting in microfluidic chip systems[J]. Lab on a Chip, 2019, 19(3): 403-409. |
| 40 | HASAN S, BLAHA M E, PIENDL S K, et al. Two-photon fluorescence lifetime for label-free microfluidic droplet sorting[J].Analytical and Bioanalytical Chemistry, 2022, 414(1): 721-730. |
| 41 | HUNG S T, MUKHERJEE S, JIMENEZ R. Enrichment of rare events using a multi-parameter high throughput microfluidic droplet sorter[J]. Lab on a Chip, 2020, 20(4): 834-843. |
| 42 | NEUN S, KAMINSKI T S, HOLLFELDER F. Single-cell activity screening in microfluidic droplets[M]// Methods in enzymology: enzyme activity in single cells. Amsterdam: Elsevier, 2019: 95-112. |
| 43 | COLIN P Y, KINTSES B, GIELEN F, et al. Ultrahigh-throughput discovery of promiscuous enzymes by picodroplet functional metagenomics[J]. Nature Communications, 2015, 6: 10008. |
| 44 | MEDCALF E J, GANTZ M, KAMINSKI T S, et al. Ultra-high-throughput absorbance-activated droplet sorting for enzyme screening at kilohertz frequencies[J]. Analytical Chemistry, 2023, 95(10): 4597-4604. |
| 45 | MACEICZYK R M, HESS D, CHIU F W Y, et al. Differential detection photothermal spectroscopy: towards ultra-fast and sensitive label-free detection in picoliter & femtoliter droplets[J]. Lab on a Chip, 2017, 17(21): 3654-3663. |
| 46 | KEMPA E E, SMITH C A, LI X, et al. Coupling droplet microfluidics with mass spectrometry for ultrahigh-throughput analysis of complex mixtures up to and above 30 Hz[J]. Analytical Chemistry, 2020, 92(18): 12605-12612. |
| 47 | WANG X X, REN L H, SU Y T, et al. Raman-activated droplet sorting (RADS) for label-free high-throughput screening of microalgal single-cells[J]. Analytical Chemistry, 2017, 89(22): 12569-12577. |
| 48 | WANG X X, XIN Y, REN L H, et al. Positive dielectrophoresis-based Raman-activated droplet sorting for culture-free and label-free screening of enzyme function in vivo [J]. Science Advances, 2020, 6(32): eabb3521. |
| 49 | BEMETZ J, WEGEMANN A, SAATCHI K, et al. Microfluidic-based synthesis of magnetic nanoparticles coupled with miniaturized NMR for online relaxation studies[J]. Analytical Chemistry, 2018, 90(16): 9975-9982. |
| 50 | ANAGNOSTIDIS V, SHERLOCK B, METZ J, et al. Deep learning guided image-based droplet sorting for on-demand selection and analysis of single cells and 3D cell cultures[J]. Lab on a Chip, 2020, 20(5): 889-900. |
| 51 | RIEMER J, HOEPKEN H H, CZERWINSKA H, et al. Colorimetric ferrozine-based assay for the quantitation of iron in cultured cells[J]. Analytical Biochemistry, 2004, 331(2): 370-375. |
| 52 | HANSEN S K, JAMALI B, HUBBUCH J. Selective high throughput protein quantification based on UV absorption spectra[J]. Biotechnology and Bioengineering, 2013, 110(2): 448-460. |
| 53 | DUNCOMBE T A, PONTI A, SEEBECK F P, et al. UV-vis spectra-activated droplet sorting for label-free chemical identification and collection of droplets[J]. Analytical Chemistry, 2021, 93(38): 13008-13013. |
| 54 | HOLLAND-MORITZ D A, WISMER M K, MANN B F, et al. Mass activated droplet sorting (MADS) enables high-throughput screening of enzymatic reactions at nanoliter scale[J]. Angewandte Chemie International Edition, 2020, 59(11): 4470-4477. |
| 55 | EL-ANEED A, COHEN A, BANOUB J. Mass spectrometry, review of the basics: electrospray, MALDI, and commonly used mass analyzers[J]. Applied Spectroscopy Reviews, 2009, 44(3): 210-230. |
| 56 | HA N S, DE RAAD M, HAN L Z, et al. Faster, better, and cheaper: harnessing microfluidics and mass spectrometry for biotechnology[J]. RSC Chemical Biology, 2021, 2(5): 1331-1351. |
| 57 | SUN S W, KENNEDY R T. Droplet electrospray ionization mass spectrometry for high throughput screening for enzyme inhibitors[J]. Analytical Chemistry, 2014, 86(18): 9309-9314. |
| 58 | SUN S W, BUER B C, MARSH E N G, et al. A label-free Sirtuin 1 assay based on droplet-electrospray ionization mass spectrometry[J]. Analytical Methods, 2016, 8(17): 3458-3465. |
| 59 | STEYER D J, KENNEDY R T. High-throughput nanoelectrospray ionization-mass spectrometry analysis of microfluidic droplet samples[J]. Analytical Chemistry, 2019, 91(10): 6645-6651. |
| 60 | KUDELSKI A. Analytical applications of Raman spectroscopy[J]. Talanta, 2008, 76(1): 1-8. |
| 61 | SAFIR F, VU N, TADESSE L F, et al. Combining acoustic bioprinting with AI-assisted Raman spectroscopy for high-throughput identification of bacteria in blood[J]. Nano Letters, 2023, 23(6): 2065-2073. |
| 62 | LIU Z S, LIU D M, CAI Y D, et al. Application of nuclear magnetic resonance (NMR) in coalbed methane and shale reservoirs: a review[J]. International Journal of Coal Geology, 2020, 218: 103261. |
| 63 | CAO X Y, YANG J, MAO J D. Characterization of kerogen using solid-state nuclear magnetic resonance spectroscopy: a review[J]. International Journal of Coal Geology, 2013, 108: 83-90. |
| 64 | VAN MEERTEN S G J, VAN BENTUM P J M, KENTGENS A P M. Shim-on-chip design for microfluidic NMR detectors[J]. Analytical Chemistry, 2018, 90(17): 10134-10138. |
| 65 | SWYER I, SOONG R, DRYDEN M D M, et al. Interfacing digital microfluidics with high-field nuclear magnetic resonance spectroscopy[J]. Lab on a Chip, 2016, 16(22): 4424-4435. |
| 66 | PAN C W, HORVATH D G, BRAZA S, et al. Sorting by interfacial tension (SIFT): label-free selection of live cells based on single-cell metabolism[J]. Lab on a Chip, 2019, 19(8): 1344-1351. |
| 67 | DOBSON C, ZIELKE C, PAN C, et al. Method for passive droplet sorting after photo-tagging[J]. Micromachines, 2020, 11(11): 964. |
| 68 | ZIELKE C, PAN C W, GUTIERREZ RAMIREZ A J, et al. Microfluidic platform for the isolation of cancer-cell subpopulations based on single-cell glycolysis[J]. Analytical Chemistry, 2020, 92(10): 6949-6957. |
| 69 | WATTERSON W J, TANYERI M, WATSON A R, et al. Droplet-based high-throughput cultivation for accurate screening of antibiotic resistant gut microbes[J]. eLife, 2020, 9: e56998. |
| 70 | DOAN M, VOROBJEV I, REES P, et al. Diagnostic potential of imaging flow cytometry[J]. Trends in Biotechnology, 2018, 36(7): 649-652. |
| 71 | SUTHERLAND J D. Evolutionary optimisation of enzymes[J]. Current Opinion in Chemical Biology, 2000, 4(3): 263-269. |
| 72 | CHERRY J R, FIDANTSEF A L. Directed evolution of industrial enzymes: an update[J]. Current Opinion in Biotechnology, 2003, 14(4): 438-443. |
| 73 | TURNER N J. Directed evolution drives the next generation of biocatalysts[J]. Nature Chemical Biology, 2009, 5(8): 567-573. |
| 74 | QU G, LI A T, ACEVEDO-ROCHA C G, et al. The crucial role of methodology development in directed evolution of selective enzymes[J]. Angewandte Chemie International Edition, 2020, 59(32): 13204-13231. |
| 75 | OTTEN R, PÁDUA R A P, BUNZEL H A, et al. How directed evolution reshapes the energy landscape in an enzyme to boost catalysis[J]. Science, 2020, 370(6523): 1442-1446. |
| 76 | WANG Y J, XUE P, CAO M F, et al. Directed evolution: methodologies and applications[J]. Chemical Reviews, 2021, 121(20): 12384-12444. |
| 77 | PACKER M S, LIU D R. Methods for the directed evolution of proteins[J]. Nature Reviews Genetics, 2015, 16(7): 379-394. |
| 78 | CHIU F W Y, STAVRAKIS S. High-throughput droplet-based microfluidics for directed evolution of enzymes[J]. ELECTROPHORESIS, 2019, 40(21): 2860-2872. |
| 79 | MADHAVAN A, ARUN K B, BINOD P, et al. Design of novel enzyme biocatalysts for industrial bioprocess: harnessing the power of protein engineering, high throughput screening and synthetic biology[J]. Bioresource Technology, 2021, 325: 124617. |
| 80 | VAN LOO B, HEBERLEIN M, MAIR P, et al. High-throughput, lysis-free screening for sulfatase activity using Escherichia coli autodisplay in microdroplets[J]. ACS Synthetic Biology, 2019, 8(12): 2690-2700. |
| 81 | HE R L, DING R H, HEYMAN J A, et al. Ultra-high-throughput picoliter-droplet microfluidics screening of the industrial cellulase-producing filamentous fungus Trichoderma reesei [J]. Journal of Industrial Microbiology and Biotechnology, 2019, 46(11): 1603-1610. |
| 82 | PRODANOVIĆ R, UNG W L, ILIĆ ĐURĐIĆ K, et al. A high-throughput screening system based on droplet microfluidics for glucose oxidase gene libraries[J]. Molecules, 2020, 25(10): 2418. |
| 83 | ZUREK P J, KNYPHAUSEN P, NEUFELD K, et al. UMI-linked consensus sequencing enables phylogenetic analysis of directed evolution[J]. Nature Communications, 2020, 11: 6023. |
| 84 | SUN G Y, WU Y K, HUANG Z Y, et al. Directed evolution of diacetylchitobiose deacetylase via high-throughput droplet sorting with a novel, bacteria-based biosensor[J]. Biosensors and Bioelectronics, 2023, 219: 114818. |
| 85 | QIAO Y, HU R, CHEN D, et al. Fluorescence-activated droplet sorting of PET degrading microorganisms[J]. Journal of Hazardous Materials, 2022, 424(pt b): 127417. |
| 86 | XU A M, ZHANG X X, CAO S X, et al. Transcription-associated fluorescence-activated droplet sorting for di-rhamnolipid hyperproducers[J]. ACS Synthetic Biology, 2022, 11(6): 1992-2000. |
| 87 | ZACHOS I, GENTH R, SUTIONO S, et al. Hot flows: evolving an archaeal glucose dehydrogenase for ultrastable carba-NADP+ using microfluidics at elevated temperatures[J]. ACS Catalysis, 2022, 12(3): 1841-1846. |
| 88 | TAN Y M, ZHANG Y, HAN Y B, et al. Directed evolution of an α1,3-fucosyltransferase using a single-cell ultrahigh-throughput screening method[J]. Science Advances, 2019, 5(10): eaaw8451. |
| 89 | KAMINSKI T S, SCHELER O, GARSTECKI P. Droplet microfluidics for microbiology: techniques, applications and challenges[J]. Lab on a Chip, 2016, 16(12): 2168-2187. |
| 90 | BOWMAN E K, NGUYEN HOANG P T, GORDILLO SIERRA A R, et al. Temporal sorting of microdroplets can identify productivity differences of itaconic acid from libraries of Yarrowia lipolytica [J]. Lab on a Chip, 2023, 23(9): 2249-2256. |
| 91 | AN X S, ZUO P, YE B C. A single cell droplet microfluidic system for quantitative determination of food-borne pathogens[J]. Talanta, 2020, 209: 120571. |
| 92 | NEETHIRAJAN S, KOBAYASHI I, NAKAJIMA M, et al. Microfluidics for food, agriculture and biosystems industries[J]. Lab on a Chip, 2011, 11(9): 1574-1586. |
| 93 | XING G W, ZHANG W F, LI N, et al. Recent progress on microfluidic biosensors for rapid detection of pathogenic bacteria[J]. Chinese Chemical Letters, 2022, 33(4): 1743-1751. |
| 94 | LI S X, MA F, BACHMAN H, et al. Acoustofluidic bacteria separation[J]. Journal of Micromechanics and Microengineering, 2017, 27(1): 015031. |
| 95 | OHLSSON P, PETERSSON K, AUGUSTSSON P, et al. Acoustic impedance matched buffers enable separation of bacteria from blood cells at high cell concentrations[J]. Scientific Reports, 2018, 8: 9156. |
| 96 | HYMAN L B, CHRISTOPHER C R, ROMERO P A. Single-cell nucleic acid profiling in droplets (SNAPD) enables high-throughput analysis of heterogeneous cell populations[J]. Nucleic Acids Research, 2021, 49(18): e103. |
| 97 | SEAH Y F S, HU H X, MERTEN C A. Microfluidic single-cell technology in immunology and antibody screening[J]. Molecular Aspects of Medicine, 2018, 59: 47-61. |
| 98 | MAZUTIS L, GILBERT J, UNG W L, et al. Single-cell analysis and sorting using droplet-based microfluidics[J]. Nature Protocols, 2013, 8(5): 870-891. |
| 99 | BARANOVA M N, BABIKOVA P A, KUDZHAEV A M, et al. Live biosensors for ultrahigh-throughput screening of antimicrobial activity against gram-negative bacteria[J]. Antibiotics, 2021, 10(10): 1161. |
| 100 | SILTANEN C A, COLE R H, POUST S, et al. An oil-free picodrop bioassay platform for synthetic biology[J]. Scientific Reports, 2018, 8: 7913. |
| [1] | 郭肖杰, 剪兴金, 王立言, 张翀, 邢新会. 合成生物学表型测试生物反应器及其装备化研究进展[J]. 合成生物学, 2024, 5(1): 16-37. |
| [2] | 刘欢, 崔球. 原位电离质谱技术在微生物菌株筛选中的应用进展[J]. 合成生物学, 2023, 4(5): 980-999. |
| [3] | 吴玉洁, 刘欣欣, 刘健慧, 杨开广, 随志刚, 张丽华, 张玉奎. 基于高通量液相色谱质谱技术的菌株筛选与关键分子定量分析研究进展[J]. 合成生物学, 2023, 4(5): 1000-1019. |
| [4] | 赵国淼, 杨鑫, 张媛, 王靖, 谭剑, 魏超, 周娜娜, 李凡, 王小艳. 生物设施平台及其工业应用[J]. 合成生物学, 2023, 4(5): 892-903. |
| [5] | 涂然, 李世新, 李昊霓, 王猛. 液滴微流控技术在微生物工程菌株选育中的应用进展[J]. 合成生物学, 2023, 4(1): 165-184. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||


