Synthetic Biology Journal ›› 2022, Vol. 3 ›› Issue (1): 53-65.DOI: 10.12211/2096-8280.2021-046
• Invited Review • Previous Articles Next Articles
Construction of tumor gene circuits using CRISPR/Cas tool and their applications
SONG Fei1,2, LIU Yuchen1,2, CAI Zhiming1,2, HUANG Weiren1,2
- 1.Department of Urology,The first Affiliated Hospital of Shenzhen University,Shenzhen Second People′s Hospital,Shenzhen Institute of Translational Medicine,Shenzhen 518035,Guangdong,China
2.Guangdong Provincial Key Laboratory of Systems Biology and Synthetic Biology for Urogenital Tumors,Shenzhen 518035,Guangdong,China
-
Received:2021-04-15Revised:2021-11-24Online:2022-03-14Published:2022-02-28 -
Contact:HUANG Weiren
基于CRISPR/Cas工具的肿瘤基因线路构建及应用
宋斐1,2, 刘宇辰1,2, 蔡志明1,2, 黄卫人1,2
- 1.深圳大学第一附属医院,深圳市第二人民医院,泌尿外科,广东 深圳 518035
2.广东省泌尿生殖肿瘤系统生物学与合成生物学重点实验室,广东 深圳 518035
-
通讯作者:黄卫人 -
作者简介:宋斐 (1979—),女,副研究员,硕士生导师。E-mail:lst121@outlook.com黄卫人 (1980—),男,研究员,博士生导师。主要研究方向:(1)肿瘤基因组学,应用多组学手段鉴定肿瘤及微环境诊疗标志物,开发相关临床应用;(2)肿瘤类器官,利用体外培养系统还原肿瘤体内生长,药物筛选及耐药机制研究;(3)医学合成生物学,创新肿瘤治疗新方法。E-mail:pony8980@163.com -
基金资助:国家重点研发项目(2019YFA0906000);广东省自然科学基金(2021A1515012488)
CLC Number:
Cite this article
SONG Fei, LIU Yuchen, CAI Zhiming, HUANG Weiren. Construction of tumor gene circuits using CRISPR/Cas tool and their applications[J]. Synthetic Biology Journal, 2022, 3(1): 53-65.
宋斐, 刘宇辰, 蔡志明, 黄卫人. 基于CRISPR/Cas工具的肿瘤基因线路构建及应用[J]. 合成生物学, 2022, 3(1): 53-65.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2021-046
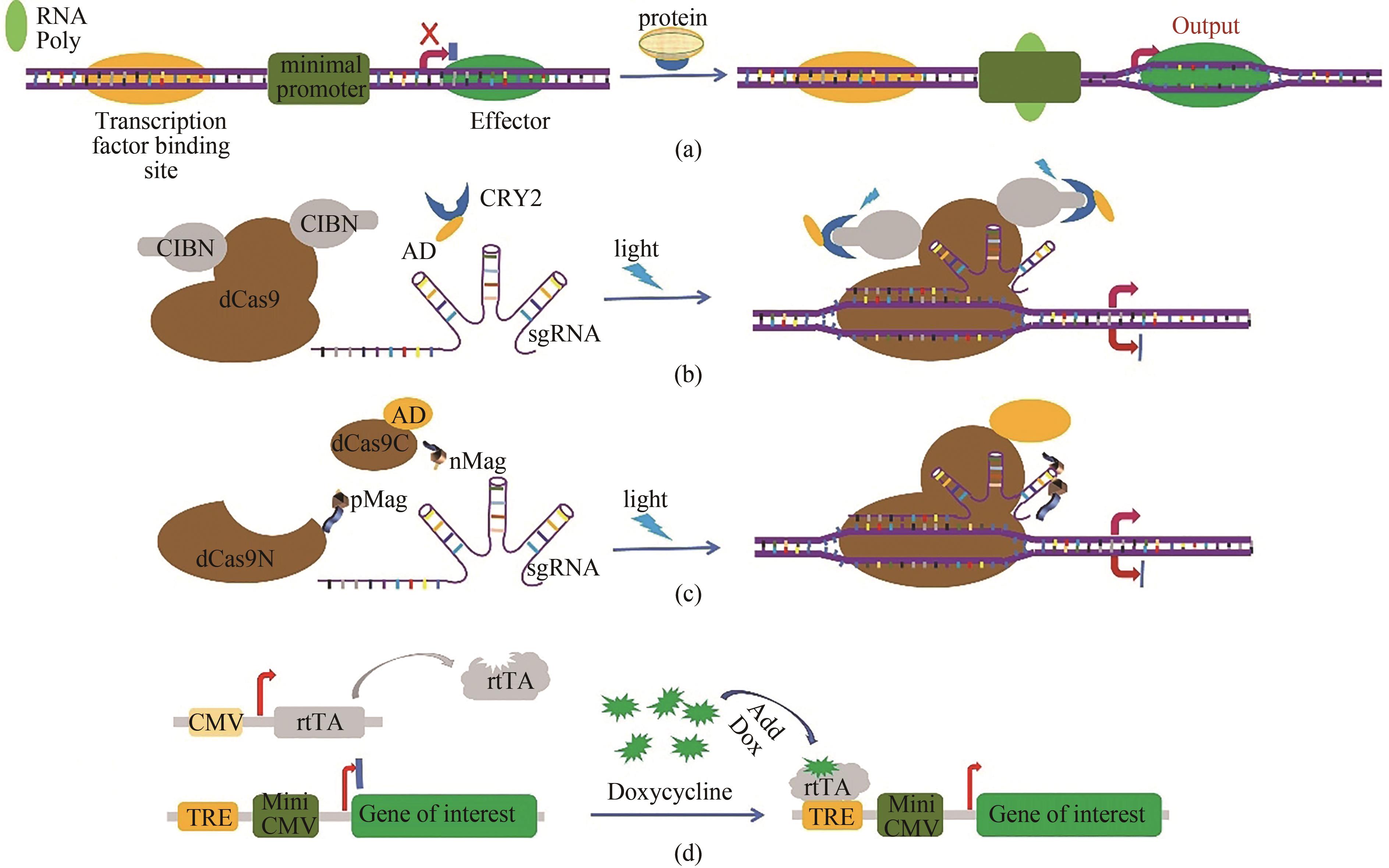
Fig. 1 Schematic representation for the artificial gene circuits developed based on CRISPR/Cas technology(a) artificial sequences of transcription factor binding site are inserted into the upstream of the Cas9 coding sequences to control gene expression by sensing intracellular signal proteins. In malignant tumor cells, some specific transcription factors such as β-catenin and NF-κB are abnormally activated. The expression of the downstream CRISPR/Cas genes is turned on when abnormal signal proteins in malignant cells bind to the transcription factor binding site and RNA polymerase (RNA poly) is recruited to the TATA box. Effector represents the Cas9 protein. (b) the light-inducible gene expression regulation system in cancer cells developed based on CRISPR/Cas9 technology. Blue light stimulation induces heterodimerization between A. thaliana cryptochrome 2 (CRY2) and its binding partner CIBN (cryptochrome-interacting basic helix-loop-helix protein 1). Therefore, the transcriptional activation domain (AD) fused with the CRY2 protein is targeted to the specific region and promotes the expression of downstream genes. (c) schematic diagram for the light inducible CRISPR/dCas9 system. The dCas9 is split into two fragments lacking nuclease activity, and the dCas9 fragments are fused with light-inducible dimerization domains (pMag and nMag). Blue light stimulation induces heterodimerization between pMag and nMag, which enables split dCas9 fragments to reassociate, thereby reconstituting RNA-guided transcriptional activation activity. (d) schematic diagram for a small molecular artificial switch system. The binding of doxycycline results in the conformational change of reverse tetracycline transcriptional activator rtTA, and then the activated rtTA could bind to the Tet-responsive element (TRE) to drive the expression of the target genes, such as Cas9 gene.
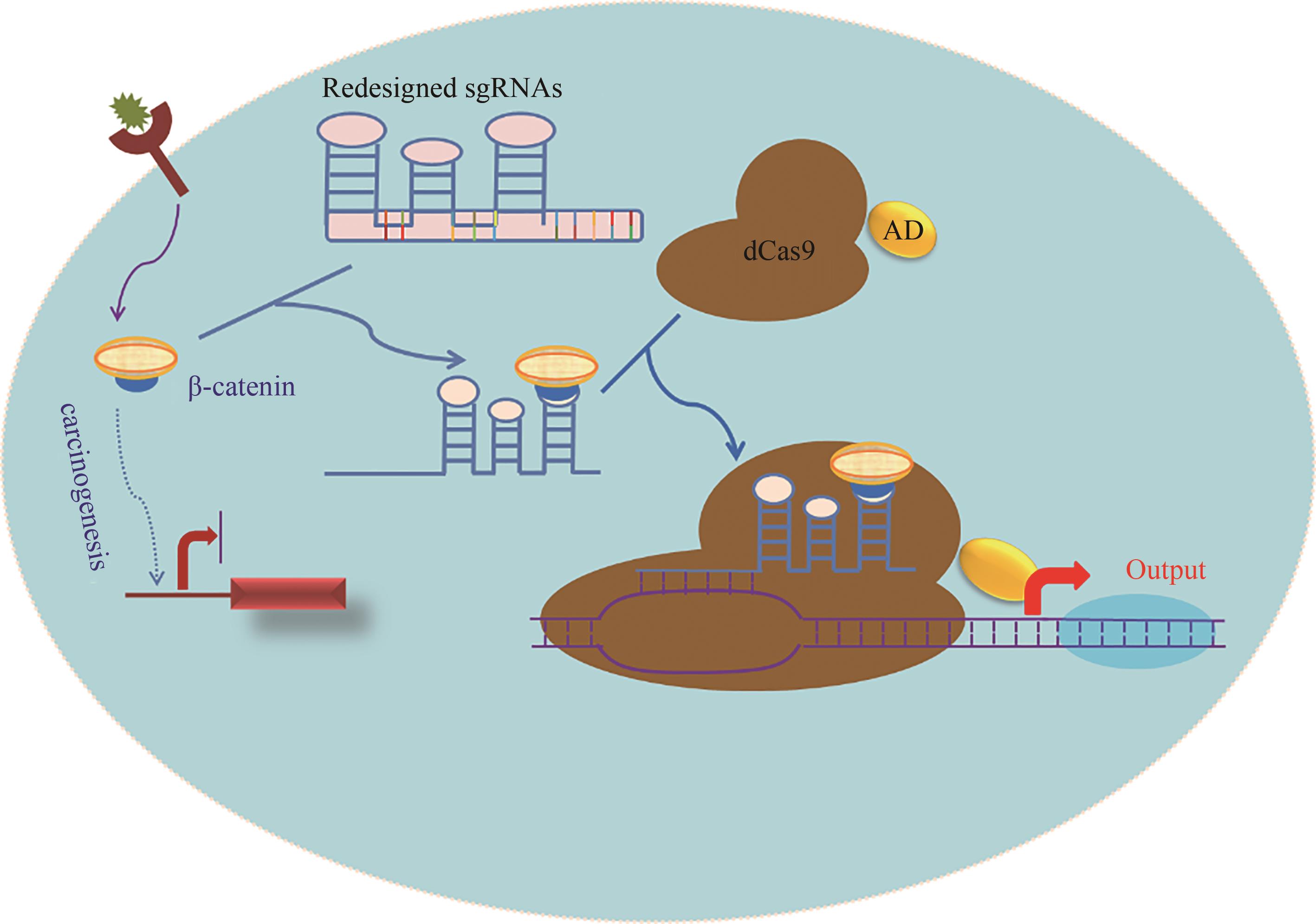
Fig. 2 Schematic diagram for the signal conductor that links one signal with another developed based on CRISPR/Cas technology[The β-catenin activates the Wnt pathway, and promotes the proliferation of tumor cells. The redesigned sgRNA preferentially binds to the endogenous β-catenin, and then couples with dCas9-AD protein to activate the output genes, such as the endogenous tumor suppressor genes (eg, p53) or apoptosis genes (eg, p21 and caspase 3), enabling the tumor cells to redirect oncogenic signaling to an anti-oncogenic pathway.]
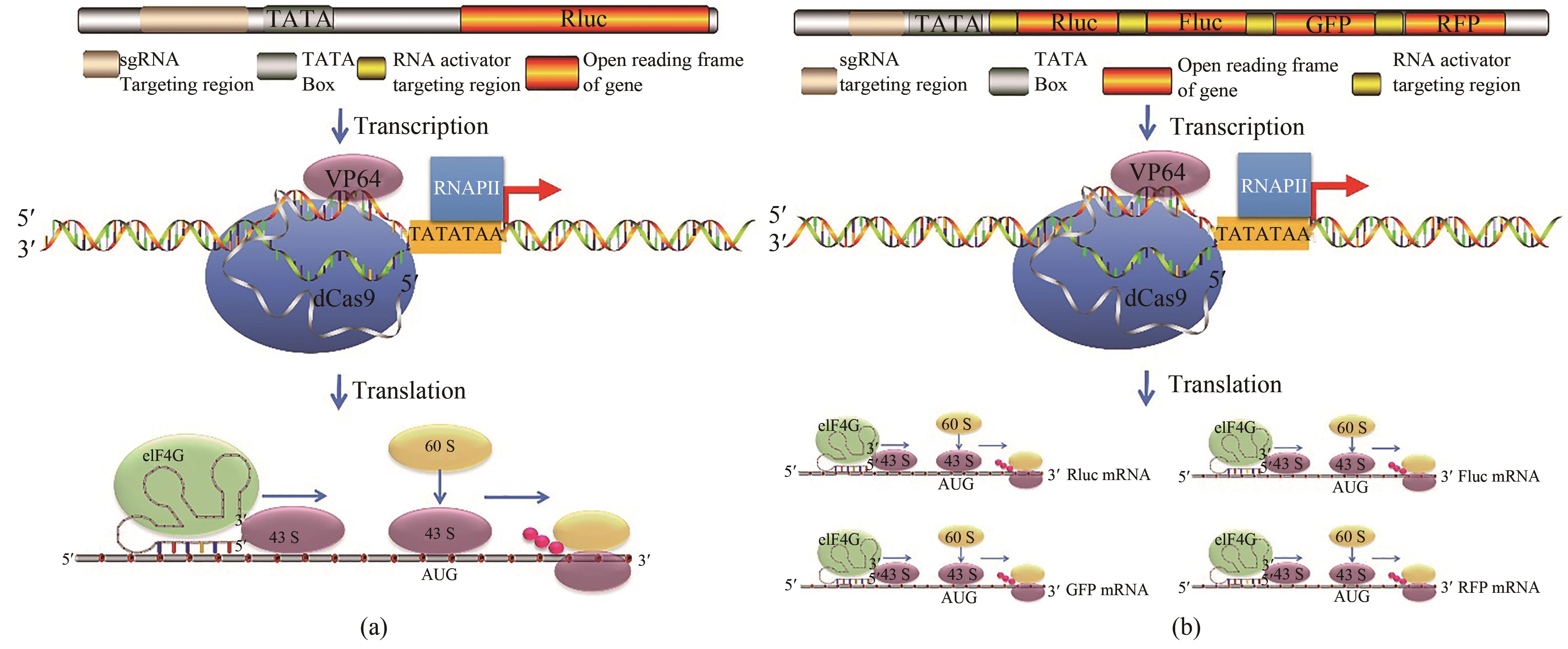
Fig. 3 CRISPReader drives gene expression by coupling the transcriptional and translational mechanisms[52](a) CRISPReader is constructed by combining transcriptional and translational platforms. The dCas9-VP64 protein robustly activated transcription of reporter is constructed when combined with sgRNA targeting sequences near the TATA box. Then, the RNA activator leads to the formation of initiation factor complexes involving eIF4G and recruits ribosomes to initiate translation. (b) mechanisms of the CRISPReader designed to drive the gene cluster expression. After dCas9-VP64-mediated transcription, the RNA activators bind to each targeted mRNA and independently initiates mRNA translation.
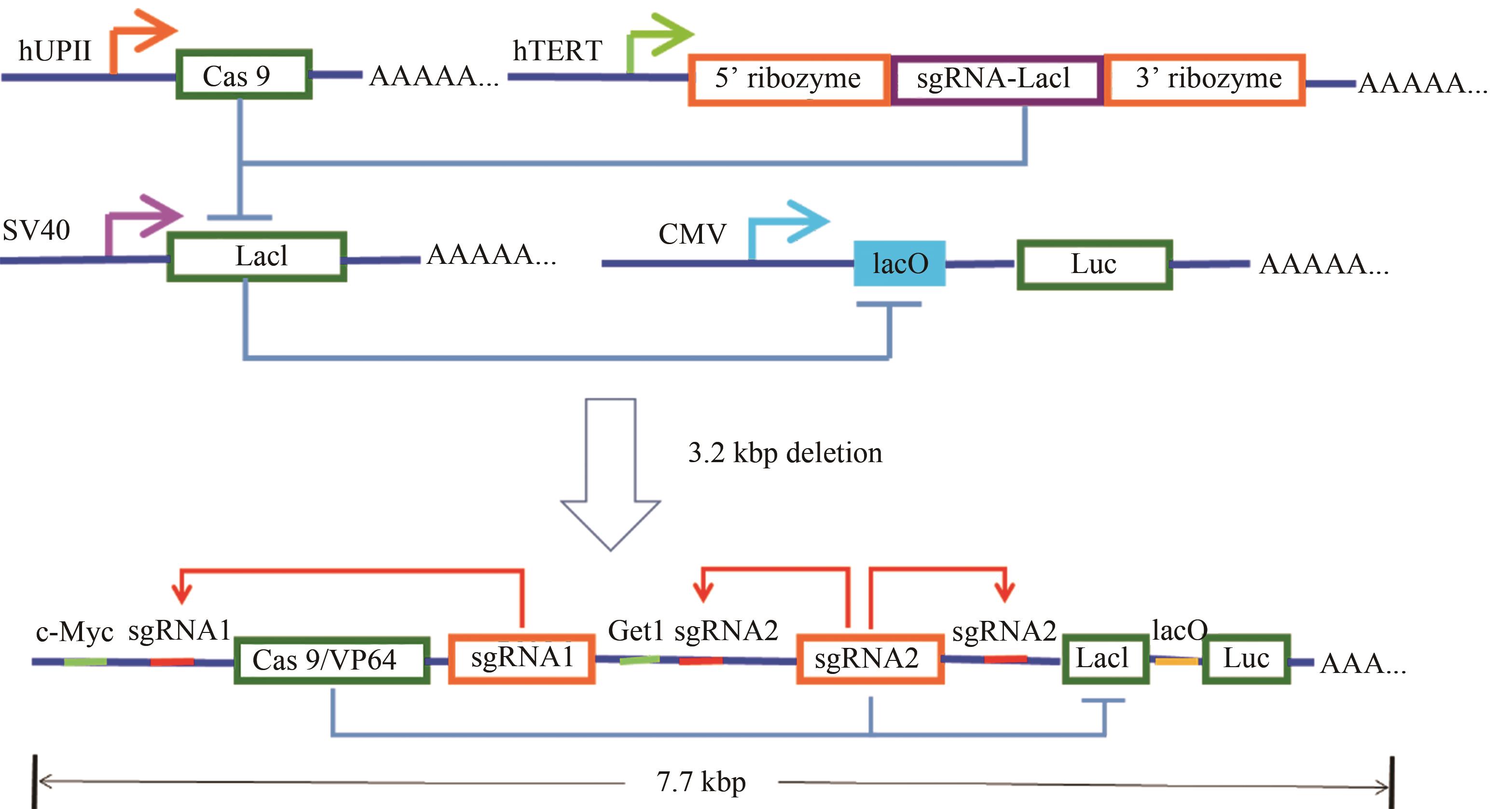
Fig. 4 Design and construction of the AND gate minigene circuits[54](The UPII promoter drives the transcription of Cas9 mRNA, while the TERT promoter is used to promote the transcription of sgRNA targeting LacI. The output Renilla luciferase gene is regulated by a LacI-controlled CMV promoter. The luciferase is expressed only when both UPII promoter and TERT promoter are activated. In the design of the minigene circuit, the UPII and TERT promoters are replaced by their respective transcription factor binding elements. Both c-Myc and Get1, only in bladder cancer cells, have a relative high expression level at the same time. After initial expression of sgRNA1 and sgRNA2, they could further bind to the upstream of their own transcription initiation sites, and amplify the transcription signals of c-Myc and Get1 through the positive feedback mechanism to amplify their downstream gene transcription. Furthermore, the LacI gene is knocked out by sgRNA2, and luciferase reporter gene is activated by transcription. In normal bladder epithelial cells, luciferase could not be effectively transcribed and further silenced by a trace amount of LacI expressed at the background level.)
| 1 | HANAHAN D, WEINBERG R A. The hallmarks of cancer [J]. Cell, 2000, 100(1): 57-70. |
| 2 | BROPHY J A N, VOIGT C A. Principles of genetic circuit design[J]. Nature Methods, 2014, 11(5): 508-520. |
| 3 | ROYBAL K T, RUPP L J, MORSUT L, et al. Precision tumor recognition by T cells with combinatorial antigen-sensing circuits[J]. Cell, 2016, 164(4): 770-779. |
| 4 | NISSIM L, WU M R, PERY E, et al. Synthetic RNA-based immunomodulatory gene circuits for cancer immunotherapy[J]. Cell, 2017, 171(5): 1138-1150. |
| 5 | XIE M Q, YE H F, WANG H, et al. β-Cell-mimetic designer cells provide closed-loop glycemic control[J]. Science, 2016, 354(6317): 1296-1301. |
| 6 | ZETSCHE B, GOOTENBERG J S, ABUDAYYEH O O, et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system[J]. Cell, 2015, 163(3): 759-771. |
| 7 | TANG X, LOWDER L G, ZHANG T, et al. A CRISPR-Cpf1 system for efficient genome editing and transcriptional repression in plants[J]. Nature Plants, 2017, 3: 17018. |
| 8 | YAMANO T, ZETSCHE B, ISHITANI R, et al. Structural basis for the canonical and non-canonical PAM recognition by CRISPR-Cpf1[J]. Molecular Cell, 2017, 67(4): 633-645. |
| 9 | WANG L P, WANG H J, LIU H Y, et al. Improved CRISPR-Cas12a-assisted one-pot DNA editing method enables seamless DNA editing[J]. Biotechnology and Bioengineering, 2019, 116(6): 1463-1474. |
| 10 | NIHONGAKI Y, OTABE T, UEDA Y, et al. A split CRISPR-Cpf1 platform for inducible genome editing and gene activation[J]. Nature Chemical Biology, 2019, 15(9): 882-888. |
| 11 | FREIJE C A, MYHRVOLD C, BOEHM C K, et al. Programmable inhibition and detection of RNA viruses using Cas13[J]. Molecular Cell, 2019, 76(5): 826-837. |
| 12 | YANG L Z, WANG Y, LI S Q, et al. Dynamic imaging of RNA in living cells by CRISPR-Cas13 systems[J]. Molecular Cell, 2019, 76(6): 981-997. |
| 13 | NANDAGOPAL N, ELOWITZ M B. Synthetic biology: Integrated gene circuits[J]. Science, 2011, 333(6047): 1244-1248. |
| 14 | FRIEDLAND A E, LU T K, WANG X, et al. Synthetic gene networks that count[J]. Science, 2009, 324(5931): 1199-1202. |
| 15 | DANINO T, MONDRAGÓN-PALOMINO O, TSIMRING L, et al. A synchronized quorum of genetic clocks[J]. Nature, 2010, 463(7279): 326-330. |
| 16 | NIELSEN J, KEASLING J D. Engineering cellular metabolism[J]. Cell, 2016, 164(6): 1185-1197. |
| 17 | GUPTA S, BRAM E E, WEISS R. Genetically programmable pathogen sense and destroy[J]. ACS Synthetic Biology, 2013, 2(12): 715-723. |
| 18 | CULLER S J, HOFF K G, SMOLKE C D. Reprogramming cellular behavior with RNA controllers responsive to endogenous proteins[J]. Science, 2010, 330(6008): 1251-1255. |
| 19 | ZHAN H J, XIE H B, ZHOU Q, et al. Synthesizing a genetic sensor based on CRISPR-Cas9 for specifically killing p53-deficient cancer cells[J]. ACS Synthetic Biology, 2018, 7(7): 1798-1807. |
| 20 | MEGRAW M, CUMBIE J S, IVANCHENKO M G, et al. Small genetic circuits and microRNAs: Big players in polymerase II transcriptional control in plants[J]. Plant Cell, 2016, 28(2): 286-303. |
| 21 | JANSING J, SACK M, AUGUSTINE S M, et al. CRISPR/Cas9-mediated knockout of six glycosyltransferase genes in Nicotiana benthamiana for the production of recombinant proteins lacking β-1,2-xylose and core α-1,3-fucose[J]. Plant Biotechnology Journal, 2019, 17(2): 350-361. |
| 22 | NIHONGAKI Y, YAMAMOTO S, KAWANO F, et al. CRISPR-Cas9-based photoactivatable transcription system[J]. Chemistry & Biology, 2015, 22(2): 169-174. |
| 23 | LIN F, DONG L, WANG W M, et al. An efficient light-inducible P53 expression system for inhibiting proliferation of bladder cancer cell[J]. International Journal of Biological Sciences, 2016, 12(10): 1273-1278. |
| 24 | ZHOU X X, ZOU X Z, CHUNG H K, et al. A single-chain photoswitchable CRISPR-Cas9 architecture for light-inducible gene editing and transcription[J]. ACS Chemical Biology, 2018, 13(2): 443-448. |
| 25 | STROVAS T J, ROSENBERG A B, KUYPERS B E, et al. MicroRNA-based single-gene circuits buffer protein synthesis rates against perturbations[J]. ACS Synthetic Biology, 2014, 3(5): 324-331. |
| 26 | KIPNISS N H, DINGAL P C D P, ABBOTT T R, et al. Engineering cell sensing and responses using a GPCR-coupled CRISPR-Cas system[J]. Nature Communications, 2017, 8(1): 2212. |
| 27 | KIANI S, BEAL J, EBRAHIMKHANI M R, et al. CRISPR transcriptional repression devices and layered circuits in mammalian cells[J]. Nature Methods, 2014, 11(7): 723-726. |
| 28 | WANG Y D, LIAO S Y, GUAN N Z, et al. A versatile genetic control system in mammalian cells and mice responsive to clinically licensed sodium ferulate[J]. Science Advances, 2020, 6(32): eabb9484. |
| 29 | ZHOU Q, ZHAN H, LIAO X, et al. A revolutionary tool: CRISPR technology plays an important role in construction of intelligentized gene circuits[J]. Cell Proliferation, 2019, 52(2): e12552. |
| 30 | SHAO J W, XUE S, YU G L, et al. Smartphone-controlled optogenetically engineered cells enable semiautomatic glucose homeostasis in diabetic mice[J]. Science Translational Medicine, 2017, 9(387): eaal2298. |
| 31 | YU Y H, WU X, GUAN N Z, et al. Engineering a far-red light-activated split-Cas9 system for remote-controlled genome editing of internal organs and tumors[J]. Science Advances, 2020, 6(28): eabb1777. |
| 32 | ZHANG Y, LING X Y, SU X X, et al. Optical control of a CRISPR/Cas9 system for gene editing by using photolabile crRNA[J]. Angewandte Chemie International Edtion, 2020, 59(47): 20895-20899. |
| 33 | ZOU R S, LIU Y, WU B, et al. Cas9 deactivation with photocleavable guide RNAs[J]. Molecular Cell, 2021, 81(7): 1553-1565. |
| 34 | BUBECK F, HOFFMANN M D, HARTEVELD Z, et al. Engineered anti-CRISPR proteins for optogenetic control of CRISPR-Cas9[J]. Nature Methods, 2018, 15(11): 924-927. |
| 35 | NISSIM L, BAR-ZIV R H. A tunable dual-promoter integrator for targeting of cancer cells[J]. Molecular Systems Biology, 2010, 6: 444. |
| 36 | XIE Z, WROBLEWSKA L, PROCHAZKA L, et al. Multi-input RNAi-based logic circuit for identification of specific cancer cells[J]. Science, 2011, 333(6047): 1307-1311. |
| 37 | LI Y Q, JIANG Y, CHEN H, et al. Modular construction of mammalian gene circuits using TALE transcriptional repressors[J]. Nature Chemical Biology, 2015, 11(3): 207-213. |
| 38 | MA D C, PENG S G, XIE Z. Integration and exchange of split dCas9 domains for transcriptional controls in mammalian cells[J]. Nature Communications, 2016, 7: 13056. |
| 39 | MORSUT L, ROYBAL K T, XIONG X, et al. Engineering customized cell sensing and response behaviors using synthetic Notch receptors[J]. Cell, 2016, 164(4): 780-791. |
| 40 | LIU Y C, ZHAN Y H, CHEN Z C, et al. Directing cellular information flow via CRISPR signal conductors[J]. Nature Methods, 2016, 13(11): 938-944. |
| 41 | LIU Y C, LI J F, CHEN Z C, et al. Synthesizing artificial devices that redirect cellular information at will[J]. eLife, 2018, 7: e31936. |
| 42 | STERNBERG S H, REDDING S, JINEK M, et al. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9[J]. Nature, 2014, 507(7490): 62-67. |
| 43 | CHENG A W, WANG H Y, YANG H, et al. Multiplexed activation of endogenous genes by CRISPR-on, an RNA-guided transcriptional activator system[J]. Cell Research, 2013, 23(10): 1163-1171. |
| 44 | ZALATAN J G, LEE M E, ALMEIDA R, et al. Engineering complex synthetic transcriptional programs with CRISPR RNA scaffolds[J]. Cell, 2015, 160(1/2): 339-350. |
| 45 | RAN F A, CONG L, YAN W X, et al. In vivo genome editing using Staphylococcus aureus Cas9[J]. Nature, 2015, 520(7546): 186-191. |
| 46 | BURSTEIN D, HARRINGTON L B, STRUTT S C, et al. New CRISPR-Cas systems from uncultivated microbes[J]. Nature, 2017, 542(7640): 237-241. |
| 47 | NISHIMASU H, RAN F A, HSU P D, et al. Crystal structure of Cas9 in complex with guide RNA and target DNA[J]. Cell, 2014, 156(5): 935-949. |
| 48 | NISHIMASU H, CONG L, YAN W X, et al. Crystal structure of Staphylococcus aureus Cas9[J]. Cell, 2015, 162(5): 1113-1126. |
| 49 | STERNBERG S H, LAFRANCE B, KAPLAN M, et al. Conformational control of DNA target cleavage by CRISPR-Cas9[J]. Nature, 2015, 527(7576): 110-113. |
| 50 | LIU Y C, HAN J H, CHEN Z C, et al. Engineering cell signaling using tunable CRISPR-Cpf1-based transcription factors[J]. Nature Communications, 2017, 8: 2095. |
| 51 | KIANI S, CHAVEZ A, TUTTLE M, et al. Cas9 gRNA engineering for genome editing, activation and repression[J]. Nature Methods, 2015, 12(11): 1051-1054. |
| 52 | ZHAN H J, ZHOU Q, GAO Q J, et al. Multiplexed promoterless gene expression with CRISPReader[J]. Genome Biology, 2019, 20(1): 113. |
| 53 | WEI S, ZOU Q J, LAI S S, et al. Conversion of embryonic stem cells into extraembryonic lineages by CRISPR-mediated activators[J]. Scientific Reports, 2016, 6: 19648. |
| 54 | LIU Y C, HUANG W R, CAI Z M. Synthesizing and gate minigene circuits based on CRISPReader for identification of bladder cancer cells[J]. Nature Communications, 2020, 11: 5486. |
| 55 | KOPINSKI P K, SINGH L N, ZHANG S P, et al. Mitochondrial DNA variation and cancer[J]. Nature Reviews Cancer, 2021, 21(7): 431-445. |
| 56 | CHOUDHURY A R, SINGH K K. Mitochondrial determinants of cancer health disparities[J]. Seminars in Cancer Biology, 2017, 47: 125-146. |
| 57 | JO A, HAM S, LEE G H, et al. Efficient mitochondrial genome editing by CRISPR/Cas9[J]. Biomed Research International, 2015, 2015: 305716. |
| 58 | WANG G, SHIMADA E, ZHANG J, et al. Correcting human mitochondrial mutations with targeted RNA import[J]. Proceedings of the National Academy of Sciences of the United States of America, 2012, 109(13): 4840-4845. |
| 59 | GAMMAGE P A, MORAES C T, MINCZUK M. Mitochondrial genome engineering: the revolution may not be CRISPR-ized[J]. Trends in Genetics, 2018, 34(2): 101-110. |
| 60 | SMIRNOV A, TARASSOV I, MAGER-HECKEL A M, et al. Two distinct structural elements of 5S rRNA are needed for its import into human mitochondria[J]. RNA, 2008, 14(4): 749-759. |
| 61 | YANG X B, JIANG J C, LI Z Y, et al. Strategies for mitochondrial gene editing[J]. Computational and Structural Biotechnology Journal, 2021, 19: 3319-3329. |
| 62 | YOO B C, YADAV N S, OROZCO E M JR, et al. Cas9/gRNA-mediated genome editing of yeast mitochondria and Chlamydomonas chloroplasts[J]. PeerJ, 2020, 8: e8362. |
| 63 | YUAN P Y, MAO X, WU X F, et al. Mitochondria-targeting, intracellular delivery of native proteins using biodegradable silica nanoparticles[J]. Angewandte Chemie International Edtion, 2019, 58(23): 7657-7661. |
| 64 | TORCHILIN V P. Recent approaches to intracellular delivery of drugs and DNA and organelle targeting[J]. Annual Review of Biomedical Engineering, 2006, 8: 343-375. |
| 65 | HADDAD S, ABÁNADES LÁZARO I, FANTHAM M, et al. Design of a functionalized metal-organic framework system for enhanced targeted delivery to mitochondria[J]. Journal of the American Chemical Society, 2020, 142(14): 6661-6674. |
| 66 | LOUTRE R, HECKEL A M, SMIRNOVA A, et al. Can mitochondrial DNA Be CRISPRized: Pro and Contra[J]. IUBMB Life, 2018, 70(12): 1233-1239. |
| 67 | BIAN W P, CHEN Y L, LUO J J, et al. Knock-in strategy for editing human and zebrafish mitochondrial DNA using Mito-CRISPR/Cas9 system[J]. ACS Synthetic Biology, 2019, 8(4): 621-632. |
| 68 | SWARTS D C, HEGGE J W, HINOJO I, et al. Argonaute of the archaeon Pyrococcus furiosus is a DNA-guided nuclease that targets cognate DNA[J]. Nucleic Acids Research, 2015, 43(10): 5120-5129. |
| 69 | DUNBAR C E, HIGH K A, JOUNG J K, et al. Gene therapy comes of age[J]. Science, 2018, 359(6372): eaan4672. |
| 70 | GRIMM D, BÜNING H. Small but increasingly mighty: latest advances in AAV vector research, design, and evolution[J]. Human Gene Therapy, 2017, 28(11): 1075-1086. |
| 71 | LENIS T L, SARFARE S, JIANG Z C, et al. Complement modulation in the retinal pigment epithelium rescues photoreceptor degeneration in a mouse model of Stargardt disease[J]. Proceedings of the National Academy of Sciences of the United States of America, 2017, 114(15): 3987-3992. |
| 72 | KAUFMAN H L, KOHLHAPP F J, ZLOZA A. Oncolytic viruses: a new class of immunotherapy drugs[J]. Nature Reviews Drug Discovery, 2015, 14(9): 642-662. |
| 73 | YOON A R, JUNG B K, CHOI E, et al. CRISPR-Cas12a with an oAd induces precise and cancer-specific genomic reprogramming of EGFR and efficient tumor regression[J]. Molecular Therapy, 2020, 28(10): 2286-2296. |
| 74 | PHELPS M P, YANG H, PATEL S, et al. Oncolytic virus-mediated RAS targeting in rhabdomyosarcoma[J]. Molecular Therapy-Oncolytics, 2018, 11: 52-61. |
| 75 | HUANG H Y, LIU Y Q, LIAO W X, et al. Oncolytic adenovirus programmed by synthetic gene circuit for cancer immunotherapy[J]. Nature Communications, 2019, 10: 4801. |
| 76 | SEGEL M, LASH B, SONG J W, et al. Mammalian retrovirus-like protein PEG10 packages its own mRNA and can be pseudotyped for mRNA delivery[J]. Science, 2021, 373(6557): 882-889. |
| 77 | SHIN J, JIANG F G, LIU J J, et al. Disabling Cas9 by an anti-CRISPR DNA mimic[J]. Science Advances, 2017, 3(7): e1701620. |
| 78 | CHEN J S, DAGDAS Y S, KLEINSTIVER B P, et al. Enhanced proofreading governs CRISPR-Cas9 targeting accuracy[J]. Nature, 2017, 550(7676): 407-410. |
| 79 | HARRINGTON L B, DOXZEN K W, MA E, et al. A broad-spectrum inhibitor of CRISPR-Cas9[J]. Cell, 2017, 170(6): 1224-1233.e1215. |
| [1] | MA Mengdan, SHANG Mengyu, LIU Yuchen. Application and prospect of CRISPR-Cas9 system in tumor biology [J]. Synthetic Biology Journal, 2023, 4(4): 703-719. |
| [2] | Ke XU, Jingnan WANG, Chun LI. Intelligent microbial cell factory with tolerance for green biological manufacturing [J]. Synthetic Biology Journal, 2020, 1(4): 427-439. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||