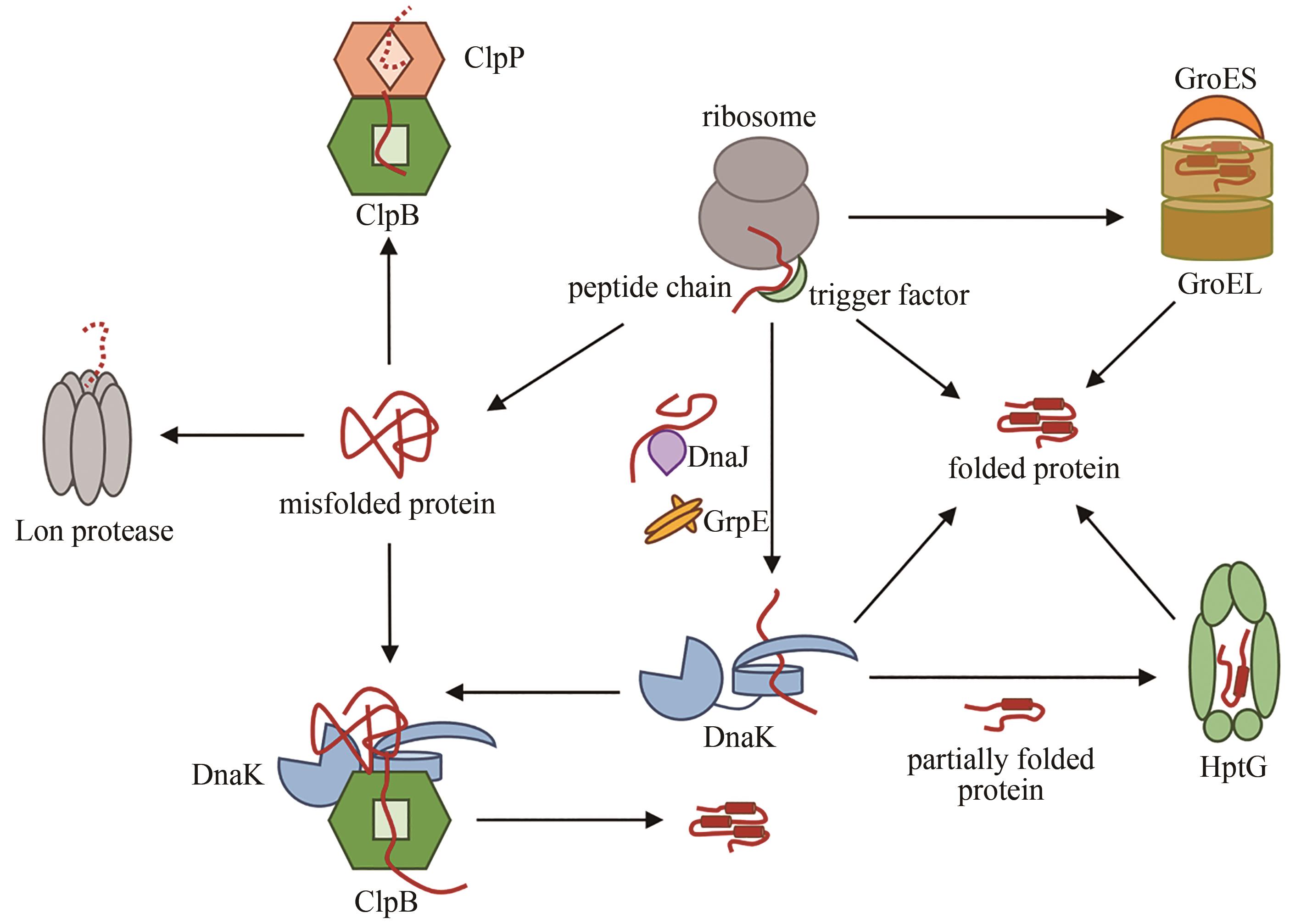
Synthetic Biology Journal ›› 2022, Vol. 3 ›› Issue (3): 445-464.DOI: 10.12211/2096-8280.2022-013
• Invited Review • Previous Articles Next Articles
Molecular chaperones promote protein stability and evolution
TANG Yuqi1,2, YE Songtao1,2, LIU Jia1,2, ZHANG Xin1,2
- 1.Department of Chemistry,School of Science,Westlake University,Hangzhou 310030,Zhejiang,China
2.Institute of Natural Sciences,Westlake Institute for Advanced Study,Hangzhou 310024,Zhejiang,China
-
Received:2022-02-12Revised:2022-05-07Online:2022-07-13Published:2022-06-30 -
Contact:ZHANG Xin
分子伴侣作用下的蛋白质稳定与进化
唐宇琦1,2, 叶松涛1,2, 刘嘉1,2, 张鑫1,2
- 1.西湖大学理学院化学系,浙江 杭州 310030
2.浙江西湖高等研究院,理学研究所,浙江 杭州 310024
-
通讯作者:张鑫 -
作者简介:唐宇琦 (1998 —),女,科研助理。研究方向是对体外蛋白质相分离的理解。E-mail: tangyuqi@westlake.edu.cn张鑫 (1978 —),男,教授,博士生导师。张鑫 课题组聚焦于化学和生物的交叉领域,以“生物聚集体化学”为研究中心,瞄准此研究领域亟需解决的重要科学和技术问题,为基础研究和生物医药产业提供重要科学支持。E-mail: zhangxin@westlake.edu.cn
Cite this article
TANG Yuqi, YE Songtao, LIU Jia, ZHANG Xin. Molecular chaperones promote protein stability and evolution[J]. Synthetic Biology Journal, 2022, 3(3): 445-464.
唐宇琦, 叶松涛, 刘嘉, 张鑫. 分子伴侣作用下的蛋白质稳定与进化[J]. 合成生物学, 2022, 3(3): 445-464.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2022-013
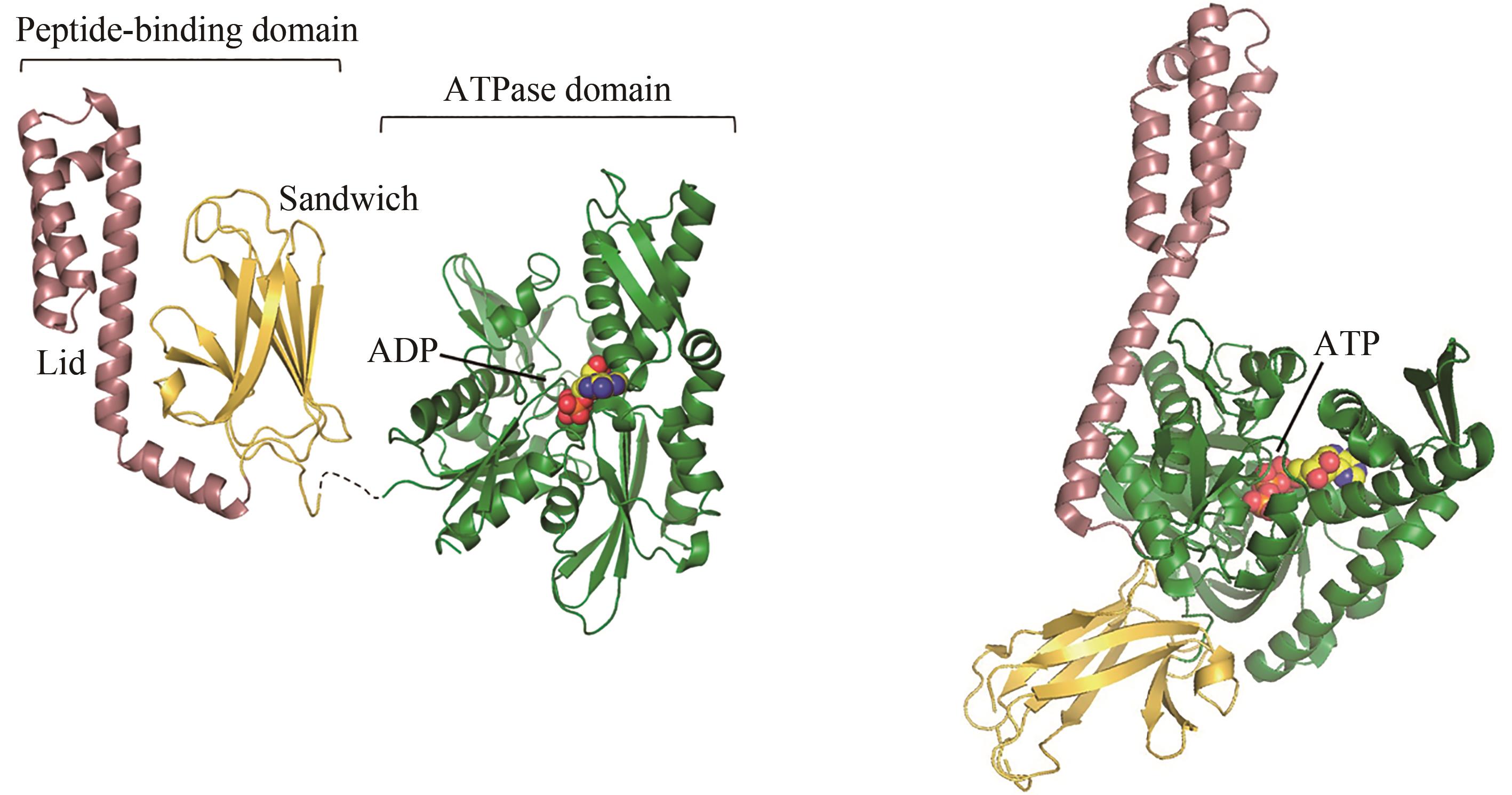
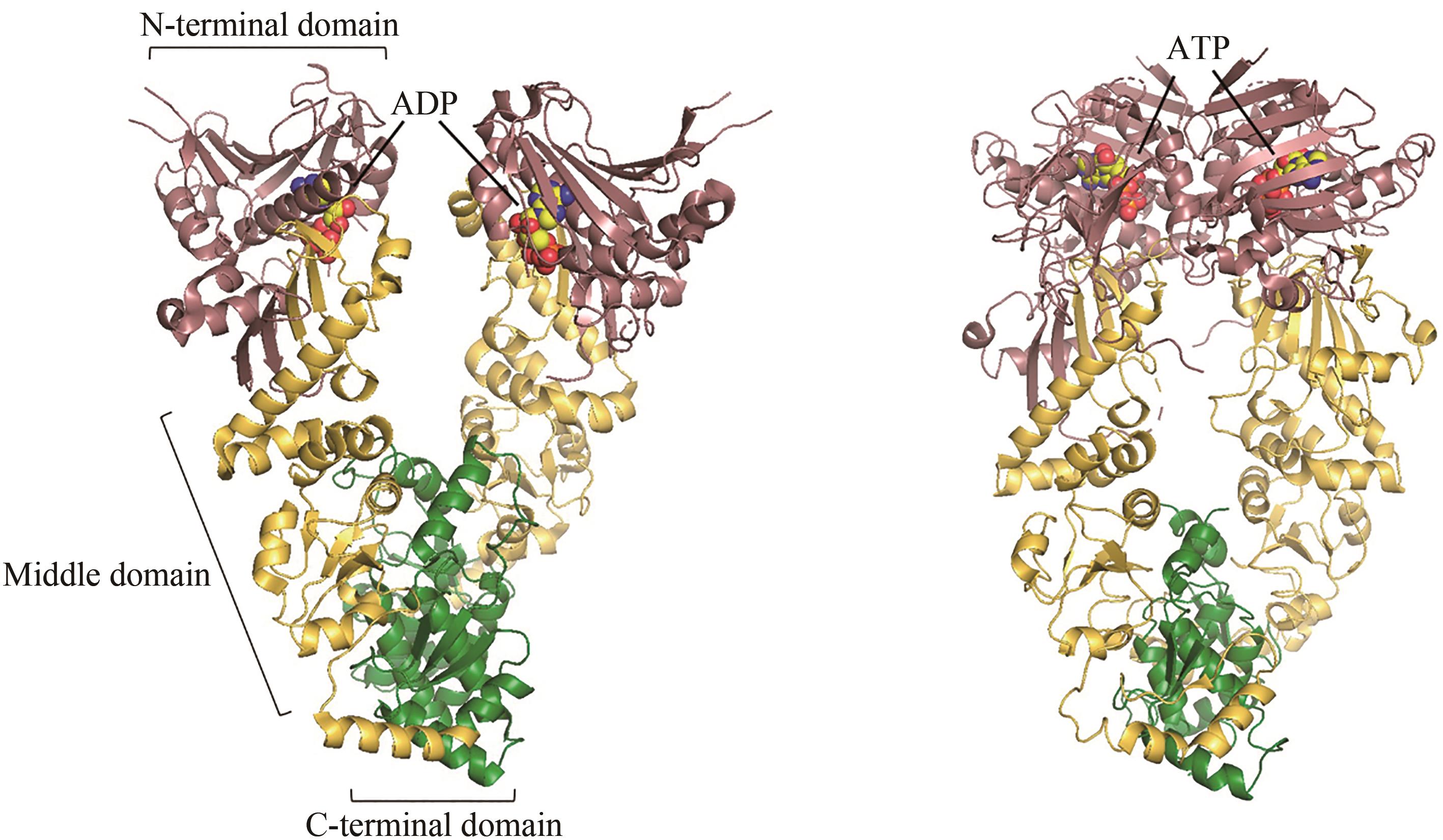
Fig. 2 ADP-bound (left) and ATP-bound (right, PDB code: 4B9Q)[55] conformations of Hsp70[In the ADP-bound state, the peptide-binding domain (PDB code: 1DKZ)[50] and ATPase domain (PDB code: 3HSC)[51] are connected by a flexible linker]
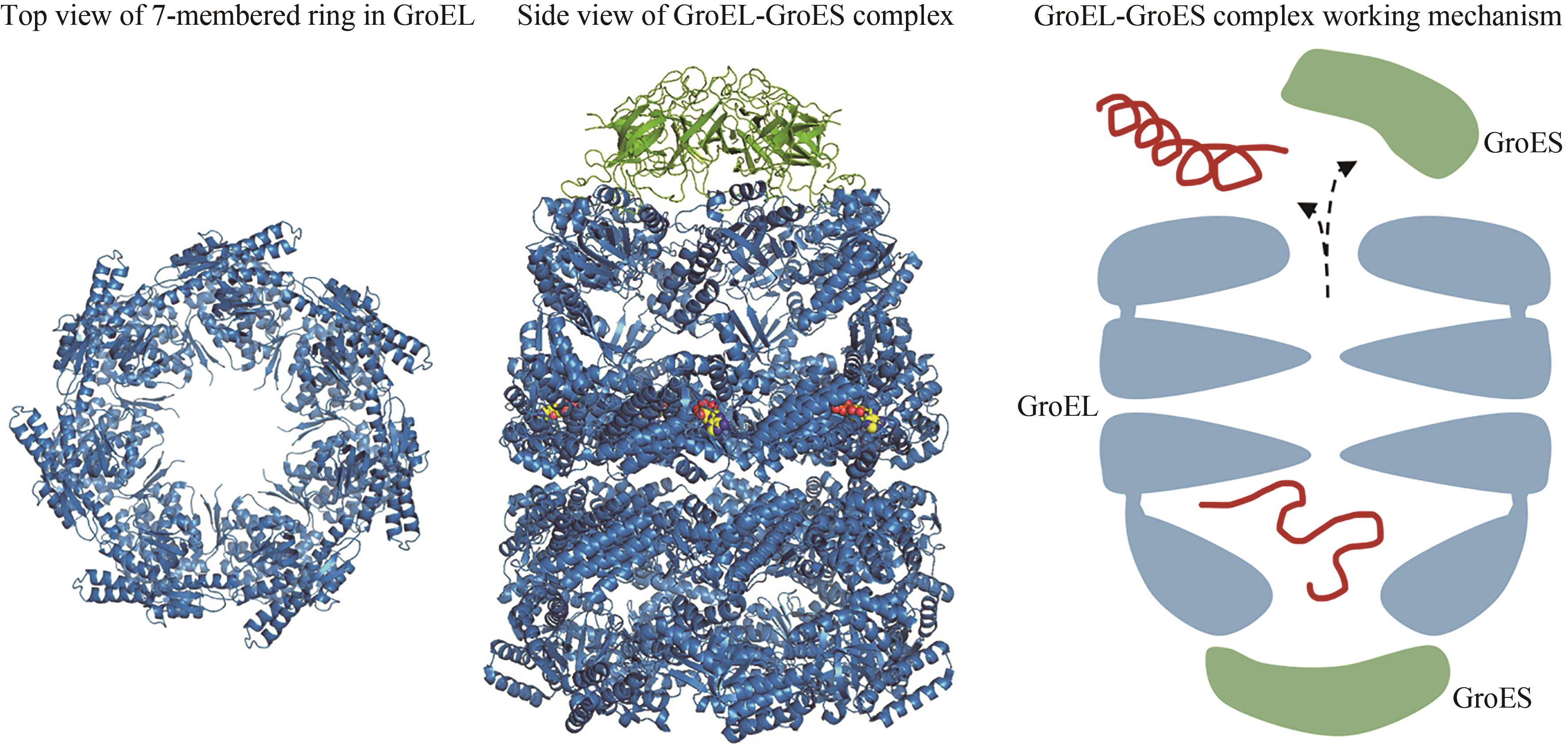
Fig. 3 Structures of the 7-membered ring in GroEL (left, PDB code: 1IOK)[65] and the Hsp60/Hsp10(middle, PDB code: 1AON)[67], as well as the working mechanism of Hsp60/Hsp10 (right)
| 1 | GIVER L, GERSHENSON A, FRESKGARD P O, et al. Directed evolution of a thermostable esterase[J]. Proceedings of the National Academy of Sciences of the United States of America, 1998, 95(22): 12809-12813. |
| 2 | VAN DEN BURG B, VRIEND G, VELTMAN O R, et al. Engineering an enzyme to resist boiling[J]. Proceedings of the National Academy of Sciences of the United States of America, 1998, 95(5): 2056-2060. |
| 3 | MIYAZAKI K, WINTRODE P L, GRAYLING R A, et al. Directed evolution study of temperature adaptation in a psychrophilic enzyme[J]. Journal of Molecular Biology, 2000, 297(4): 1015-1026. |
| 4 | GONZÁLEZ M, BAGATOLLI L A, ECHABE I, et al. Interaction of biotin with streptavidin: Thermostability and conformational changes upon binding[J]. Journal of Biological Chemistry, 1997, 272(17): 11288-11294. |
| 5 | SANCHEZ DE GROOT N, ARMAOS A, GRAÑA-MONTES R, et al. RNA structure drives interaction with proteins[J]. Nature Communications, 2019, 10: 32-46. |
| 6 | LAGIER-TOURENNE C, POLYMENIDOU M, CLEVELAND D W. TDP-43 and FUS/TLS: Emerging roles in RNA processing and neurodegeneration[J]. Human Molecular Genetics, 2010, 19(R1): R46-R64. |
| 7 | KAPELI K, PRATT G A, VU A Q, et al. Distinct and shared functions of ALS-associated proteins TDP-43, FUS and TAF15 revealed by multisystem analyses[J]. Nature Communications, 2016, 7: 12143. |
| 8 | STRICKLER S S, GRIBENKO A V, GRIBENKO A V, et al. Protein stability and surface electrostatics: A charged relationship[J]. Biochemistry, 2006, 45(9): 2761-2766. |
| 9 | CURTIS R A, PRAUSNITZ J M, BLANCH H W. Protein-protein and protein-salt interactions in aqueous protein solutions containing concentrated electrolytes[J]. Biotechnology and Bioengineering, 1998, 57(1): 11-21. |
| 10 | LABER J R, DEAR B J, MARTINS M L, et al. Charge shielding prevents aggregation of supercharged GFP variants at high protein concentration[J]. Molecular Pharmaceutics, 2017, 14(10): 3269-3280. |
| 11 | TAVERNA D M, GOLDSTEIN R A. Why are proteins marginally stable? [J]. Proteins: Structure, Function, and Genetics, 2002, 46(1): 105-109. |
| 12 | VOGL T, JATZKE C, HINZ H J, et al. Thermodynamic stability of annexin V E17G: Equilibrium parameters from an irreversible unfolding reaction[J]. Biochemistry, 1997, 36(7): 1657-1668. |
| 13 | GHOSH K, DILL K. Cellular proteomes have broad distributions of protein stability[J]. Biophysical Journal, 2010, 99(12): 3996-4002. |
| 14 | ROSS C A, POIRIER M A. Protein aggregation and neurodegenerative disease[J]. Nature Medicine, 2004, 10(7): S10-S17. |
| 15 | AGUZZI A, O'CONNOR T. Protein aggregation diseases: Pathogenicity and therapeutic perspectives[J]. Nature Reviews Drug Discovery, 2010, 9(3): 237-248. |
| 16 | 陈冬冬, 石琦, 董小平. 蛋白质体外扩增技术在神经退行性疾病诊断中的应用[J]. 疾病监测, 2022, 37(2): 180-184. |
| CHEN D D, SHI Q, DONG X P. Application of protein amplification in vitro in the diagnosis of neurodegenerative diseases[J]. Disease Surveillance, 2022, 37(2): 180-184. | |
| 17 | 张肖楠, 廉姜芳. 内质网应激与蛋白质错误折叠相关疾病的研究进展[J]. 浙江医学, 2020, 42(1): 86-90. |
| ZHANG X N, LIAN J F. Research progress of endoplasmic reticulum stress and protein misfolding-related diseases [J]. Zhejiang Medical Journal, 2020, 42(1): 86-90. | |
| 18 | 熊秋宏, 李文静, 吴长新. 自噬和泛素-蛋白酶体系统之间的交联[J]. 中国生物化学与分子生物学报, 2018, 34(11): 1154-1159. |
| XIONG Q H, LI W J, WU C X. The crosstalk between autophagy and ubiquitin-proteasome system[J]. Chinese Journal of Biochemistry and Molecular Biology, 2018, 34(11): 1154-1159. | |
| 19 | OTTENS F, FRANZ A, HOPPE T. Build-UPS and break-downs: Metabolism impacts on proteostasis and aging[J]. Cell Death & Differentiation, 2021, 28(2): 505-521. |
| 20 | DONG Z, CUI H J. The autophagy-lysosomal pathways and their emerging roles in modulating proteostasis in tumors[J]. Cells, 2018, 8(1): 4. |
| 21 | HARTL F U, MARTIN J. Molecular chaperones in cellular protein folding[J]. Current Opinion in Structural Biology, 1995, 5(1): 92-102. |
| 22 | MORÁN LUENGO T, MAYER M P, RÜDIGER S G D. The Hsp70-Hsp90 chaperone cascade in protein folding[J]. Trends in Cell Biology, 2019, 29(2): 164-177. |
| 23 | HARTL F U, BRACHER A, HAYER-HARTL M. Molecular chaperones in protein folding and proteostasis[J]. Nature, 2011, 475(7356): 324-332. |
| 24 | KAISER C M, CHANG H C, AGASHE V R, et al. Real-time observation of trigger factor function on translating ribosomes[J]. Nature, 2006, 444(7118): 455-460. |
| 25 | DOYLE S M, GENEST O, WICKNER S. Protein rescue from aggregates by powerful molecular chaperone machines[J]. Nature Reviews Molecular Cell Biology, 2013, 14(10): 617-629. |
| 26 | TAVARIA M, GABRIELE T, KOLA I, et al. A hitchhiker's guide to the human Hsp70 family[J]. Cell Stress & Chaperones, 1996, 1(1): 23-28. |
| 27 | THOMAS J G, BANEYX F. Protein misfolding and inclusion body formation in recombinant escherichia coli cells overexpressing heat-shock proteins[J]. Journal of Biological Chemistry, 1996, 271(19): 11141-11147. |
| 28 | HALDAR S, TAPIA-ROJO R, ECKELS E C, et al. Trigger factor chaperone acts as a mechanical foldase[J]. Nature Communications, 2017, 8: 668. |
| 29 | NUNES J M, MAYER-HARTL M, HARTL F U, et al. Action of the Hsp70 chaperone system observed with single proteins[J]. Nature Communications, 2015, 6: 6307. |
| 30 | ASRAFUZZAMAN S, PATRA M, ROY S S, et al. The holding and folding chaperone properties of two small heat-shock pair proteins IbpA and IbPB of Escherichia coli [J]. Biophysical Journal, 2011, 100(3): 209a-210a. |
| 31 | DEVILLE C, FRANKE K, MOGK A, et al. Two-step activation mechanism of the ClpB disaggregase for sequential substrate threading by the main ATPase motor[J]. Cell Reports, 2019, 27(12): 3433-3446.e4. |
| 32 | 郑岩.不同分子伴侣活性对新生肽链的影响[D].哈尔滨: 哈尔滨工业大学,2020. |
| ZHENG Yan. Effects of different molecular chaperone activities on nascent peptide chains[D].Harbin: Harbin Institute of Technology,2020. | |
| 33 | RIZZOLO K, YU A Y H, OLOGBENLA A, et al. Functional cooperativity between the trigger factor chaperone and the ClpXP proteolytic complex[J]. Nature Communications, 2021, 12: 281. |
| 34 | WRUCK F, AVELLANEDA M J, KOERS E J, et al. Protein folding mediated by trigger factor and Hsp70: new insights from single-molecule approaches[J]. Journal of Molecular Biology, 2018, 430(4): 438-449. |
| 35 | BHANDARI V, HOURY W A. Substrate interaction networks of the Escherichia coli chaperones: Trigger factor, DnaK and GroEL[J]. Advances in Experimental Medicine and Biology, 2015, 883: 271-294. |
| 36 | AGASHE V R, GUHA S, CHANG H C, et al. Function of trigger factor and DnaK in multidomain protein folding: increase in yield at the expense of folding speed[J]. Cell, 2004, 117(2): 199-209. |
| 37 | LASKOWSKA E, WAWRZYNÓW A, TAYLOR A. IbpA and IbpB, the new heat-shock proteins, bind to endogenous Escherichia coli proteins aggregated intracellularly by heat shock[J]. Biochimie, 1996, 78(2): 117-122. |
| 38 | HASLBECK M, VIERLING E. A first line of stress defense: small heat shock proteins and their function in protein homeostasis[J]. Journal of Molecular Biology, 2015, 427(7): 1537-1548. |
| 39 | ALLEN S P, POLAZZI J O, GIERSE J K, et al. Two novel heat shock genes encoding proteins produced in response to heterologous protein expression in Escherichia coli [J]. Journal of Bacteriology, 1992, 174(21): 6938-6947. |
| 40 | MOGK A, DEUERLING E, VORDERWÜLBECKE S, et al. Small heat shock proteins, ClpB and the DnaK system form a functional triade in reversing protein aggregation[J]. Molecular Microbiology, 2003, 50(2): 585-595. |
| 41 | SCHOPF F H, BIEBL M M, BUCHNER J. The HSP90 chaperone machinery[J]. Nature Reviews Molecular Cell Biology, 2017, 18(6): 345-360. |
| 42 | SAIBIL H. Chaperone machines for protein folding, unfolding and disaggregation[J]. Nature Reviews Molecular Cell Biology, 2013, 14(10): 630-642. |
| 43 | CALLONI G, CHEN T T, SCHERMANN S M, et al. DnaK functions as a central hub in the E. coli chaperone network[J]. Cell Reports, 2012, 1(3): 251-264. |
| 44 | MISSELWITZ B, STAECK O, RAPOPORT T A. J proteins catalytically activate Hsp70 molecules to trap a wide range of peptide sequences[J]. Molecular Cell, 1998, 2(5): 593-603. |
| 45 | JIANG Y J, ROSSI P, KALODIMOS C G. Structural basis for client recognition and activity of Hsp40 chaperones[J]. Science, 2019, 365(6459): 1313-1319. |
| 46 | SARBENG E B, LIU Q D, TIAN X L, et al. A functional DnaK dimer is essential for the efficient interaction with Hsp40 heat shock protein[J]. Journal of Biological Chemistry, 2015, 290(14): 8849-8862. |
| 47 | LIU Q D, LI H T, YANG Y, et al. A disulfide-bonded DnaK dimer is maintained in an ATP-bound state[J]. Cell Stress & Chaperones, 2017, 22(2): 201-212. |
| 48 | MAYER M P. Gymnastics of molecular chaperones[J]. Molecular Cell, 2010, 39(3): 321-331. |
| 49 | KAMPINGA H H, CRAIG E A. The HSP70 chaperone machinery: J proteins as drivers of functional specificity[J]. Nature Reviews Molecular Cell Biology, 2010, 11(8): 579-592. |
| 50 | ZHU X, ZHAO X, BURKHOLDER W F, et al. Structural analysis of substrate binding by the molecular chaperone DnaK[J]. Science, 1996, 272(5268): 1606-1614. |
| 51 | FLAHERTY K M, DELUCA-FLAHERTY C, MCKAY D B. Three-dimensional structure of the ATPase fragment of a 70K heat-shock cognate protein[J]. Nature, 1990, 346(6285): 623-628. |
| 52 | KITYK R, KOPP J, MAYER M P. Molecular mechanism of J-domain-triggered ATP hydrolysis by Hsp70 chaperones[J]. Molecular Cell, 2018, 69(2): 227-237. |
| 53 | YU H Y, ZIEGELHOFFER T, OSIPIUK J, et al. Roles of intramolecular and intermolecular interactions in functional regulation of the Hsp70 J-protein co-chaperone Sis1[J]. Journal of Molecular Biology, 2015, 427(7): 1632-1643. |
| 54 | YU H Y, ZIEGELHOFFER T, CRAIG E A. Functionality of class A and class B J-protein co-chaperones with Hsp70[J]. FEBS Letters, 2015, 589(19PartB): 2825-2830. |
| 55 | KITYK R, KOPP J, SINNING I, et al. Structure and dynamics of the ATP-bound open conformation of Hsp70 chaperones[J]. Molecular Cell, 2012, 48(6): 863-874. |
| 56 | RÜDIGER S, BUCHBERGER A, BUKAU B. Interaction of Hsp70 chaperones with substrates[J]. Nature Structural Biology, 1997, 4(5): 342-349. |
| 57 | LACKIE R E, MACIEJEWSKI A, OSTAPCHENKO V G, et al. The Hsp70/Hsp90 chaperone machinery in neurodegenerative diseases[J]. Frontiers in Neuroscience, 2017, 11: 254. |
| 58 | MOGK A, KUMMER E, BUKAU B. Cooperation of Hsp70 and Hsp100 chaperone machines in protein disaggregation[J]. Frontiers in Molecular Biosciences, 2015, 2: 22. |
| 59 | CRAIG E A. Hsp70 at the membrane: Driving protein translocation[J]. BMC Biology, 2018, 16(1): 11. |
| 60 | SU P H, LI H M. Stromal Hsp70 is important for protein translocation into pea and Arabidopsis chloroplasts[J]. The Plant Cell, 2010, 22(5): 1516-1531. |
| 61 | SAIBIL H R, FENTON W A, CLARE D K, et al. Structure and allostery of the chaperonin GroEL[J]. Journal of Molecular Biology, 2013, 425(9): 1476-1487. |
| 62 | HAYER-HARTL M, BRACHER A, HARTL F U. The GroEL-GroES chaperonin machine: A nano-cage for protein folding[J]. Trends in Biochemical Sciences, 2016, 41(1): 62-76. |
| 63 | CHEN D H, MADAN D M, WEAVER J, et al. Visualizing GroEL/ES in the act of encapsulating a folding protein[J]. Cell, 2013, 153(6): 1354-1365. |
| 64 | BRAIG K, OTWINOWSKI Z, HEGDE R, et al. The crystal structure of the bacterial chaperonin GroEL at 2.8 A[J]. Nature, 1994, 371(6498): 578-586. |
| 65 | FUKAMI T A, YOHDA M, TAGUCHI H, et al. Crystal structure of chaperonin-60 from paracoccus denitrificans[J]. Journal of Molecular Biology, 2001, 312(3): 501-509. |
| 66 | FIAUX J, BERTELSEN E B, HORWICH A L, et al. NMR analysis of a 900K GroEL-GroES complex[J]. Nature, 2002, 418(6894): 207-211. |
| 67 | XU Z H, HORWICH A L, SIGLER P B. The crystal structure of the asymmetric GroEL-GroES-(ADP)7 chaperonin complex[J]. Nature, 1997, 388(6644): 741-750. |
| 68 | HORWICH A L, FENTON W A. Chaperonin-mediated protein folding: Using a central cavity to kinetically assist polypeptide chain folding[J]. Quarterly Reviews of Biophysics, 2009, 42(2): 83-116. |
| 69 | XU Z H, HORWICH A L, SIGLER P B. The crystal structure of the asymmetric GroEL-GroES-(ADP)7 chaperonin complex[J]. Nature, 1997, 388(6644): 741-750. |
| 70 | HYEON C, LORIMER G H, THIRUMALAI D. Dynamics of allosteric transitions in GroEL[J]. Proceedings of the National Academy of Sciences of the United States of America, 2006, 103(50): 18939-18944. |
| 71 | CARUSO BAVISOTTO C, ALBERTI G, VITALE A M, et al. Hsp60 post-translational modifications: Functional and pathological consequences[J]. Frontiers in Molecular Biosciences, 2020, 7: 95. |
| 72 | WHITESELL L, LINDQUIST S L. HSP90 and the chaperoning of cancer[J]. Nature Reviews Cancer, 2005, 5(10): 761-772. |
| 73 | LUO W J, SUN W L, TALDONE T, et al. Heat shock protein 90 in neurodegenerative diseases[J]. Molecular Neurodegeneration, 2010, 5: 24. |
| 74 | CSERMELY P, SCHNAIDER T, SŐTI C, et al. The 90-kDa molecular chaperone family: Structure, function, and clinical applications. A comprehensive review[J]. Pharmacology & Therapeutics, 1998, 79(2): 129-168. |
| 75 | DIDENKO T, DUARTE A M S, KARAGÖZ G E, et al. Hsp90 structure and function studied by NMR spectroscopy[J]. Biochimica et Biophysica Acta (BBA) - Molecular Cell Research, 2012, 1823(3): 636-647. |
| 76 | ALI M M U, ROE S M, VAUGHAN C K, et al. Crystal structure of an Hsp90-nucleotide-p23/Sba1 closed chaperone complex[J]. Nature, 2006, 440(7087): 1013-1017. |
| 77 | PRODROMOU C. Mechanisms of Hsp90 regulation[J]. The Biochemical Journal, 2016, 473(16): 2439-2452. |
| 78 | KARAGÖZ G E, RÜDIGER S G D. Hsp90 interaction with clients[J]. Trends in Biochemical Sciences, 2015, 40(2): 117-125. |
| 79 | DOLLINS D E, WARREN J J, IMMORMINO R M, et al. Structures of GRP94-nucleotide complexes reveal mechanistic differences between the hsp90 chaperones[J]. Molecular Cell, 2007, 28(1): 41-56. |
| 80 | LI J, RICHTER K, REINSTEIN J, et al. Integration of the accelerator Aha1 in the Hsp90 co-chaperone cycle[J]. Nature Structural & Molecular Biology, 2013, 20(3): 326-331. |
| 81 | VERBA K A, WANG R Y R, ARAKAWA A, et al. Atomic structure of Hsp90-Cdc37-Cdk4 reveals that Hsp90 traps and stabilizes an unfolded kinase[J]. Science, 2016, 352(6293): 1542-1547. |
| 82 | WU F H, YUAN Y, LI D, et al. Extracellular HSPA1A promotes the growth of hepatocarcinoma by augmenting tumor cell proliferation and apoptosis-resistance[J]. Cancer Letters, 2012, 317(2): 157-164. |
| 83 | JAGADISH N, PARASHAR D, GUPTA N, et al. Heat shock protein 70-2 (HSP70-2) is a novel therapeutic target for colorectal cancer and is associated with tumor growth[J]. BMC Cancer, 2016, 16: 561. |
| 84 | KAMPINGA H H, HAGEMAN J, VOS M J, et al. Guidelines for the nomenclature of the human heat shock proteins[J]. Cell Stress & Chaperones, 2009, 14(1): 105-111. |
| 85 | CHEETHAM M E, CAPLAN A J. Structure, function and evolution of DnaJ: conservation and adaptation of chaperone function[J]. Cell Stress & Chaperones, 1998, 3(1): 28-36. |
| 86 | VILASI S, et al. Chaperonin of group I: oligomeric spectrum and biochemical and biological implications.Frontiers in Mole cular Biosciences, 2017, 4: 99. |
| 87 | KUBOTA H, HYNES G, WILLISON K. The chaperonin containing t-complex polypeptide 1 (TCP-1). Multisubunit machinery assisting in protein folding and assembly in the eukaryotic cytosol[J]. European Journal of Biochemistry, 1995, 230(1): 3-16. |
| 88 | HOTER A, EL-SABBAN M E, NAIM H Y. The HSP90 family: structure, regulation, function, and implications in health and disease[J]. International Journal of Molecular Sciences, 2018, 19(9): 2560. |
| 89 | LEE S C, CHOI Y C, YU M H. Effect of the N-terminal hydrophobic sequence of hepatitis B virus surface antigen on the folding and assembly of hybrid beta-galactosidase in Escherichia coli [J]. European Journal of Biochemistry, 1990, 187(2): 417-424. |
| 90 | ZHANG X, LIU Y, GENEREUX J C, et al. Heat-shock response transcriptional program enables high-yield and high-quality recombinant protein production in Escherichia coli [J]. ACS Chemical Biology, 2014, 9(9): 1945-1949. |
| 91 | ZHAO K, LIU M Z, BURGESS R R. The global transcriptional response of Escherichia coli to induced sigma 32 protein involves sigma 32 regulon activation followed by inactivation and degradation of sigma 32 in vivo [J]. The Journal of Biological Chemistry, 2005, 280(18): 17758-17768. |
| 92 | GUISBERT E, HERMAN C, LU C Z, et al. A chaperone network controls the heat shock response in E. coli [J]. Genes & Development, 2004, 18(22): 2812-2821. |
| 93 | YURA T, GUISBERT E, PORITZ M, et al. Analysis of a 32 mutants defective in chaperone-mediated feedback control reveals unexpected complexity of the heat shock response[J]. Proceedings of the National Academy of Sciences of the United States of America, 2007, 104(45): 17638-17643. |
| 94 | SUBBARAO SREEDHAR A, KALMÁR É, CSERMELY P, et al. Hsp90 isoforms: Functions, expression and clinical importance[J]. FEBS Letters, 2004, 562(1/2/3): 11-15. |
| 95 | CHEN B, PIEL W H, GUI L M, et al. The HSP90 family of genes in the human genome: insights into their divergence and evolution[J]. Genomics, 2005, 86(6): 627-637. |
| 96 | YUNO A, LEE M J, LEE S M, et al. Clinical evaluation and biomarker profiling of Hsp90 inhibitors[J]. Methods in Molecular Biology (Clifton, N J), 2018, 1709: 423-441. |
| 97 | NECKERS L, BLAGG B, HAYSTEAD T, et al. Methods to validate Hsp90 inhibitor specificity, to identify off-target effects, and to rethink approaches for further clinical development[J]. Cell Stress & Chaperones, 2018, 23(4): 467-482. |
| 98 | XIAO Y, LIU Y J. Recent advances in the discovery of novel HSP90 inhibitors: An update from 2014[J]. Current Drug Targets, 2020, 21(3): 302-317. |
| 99 | DAI C K, SAMPSON S B. HSF1: guardian of proteostasis in cancer[J]. Trends in Cell Biology, 2016, 26(1): 17-28. |
| 100 | YOON Y J, KIM J A, SHIN K D, et al. KRIBB11 inhibits HSP70 synthesis through inhibition of heat shock factor 1 function by impairing the recruitment of positive transcription elongation factor b to the hsp70 promoter[J]. Journal of Biological Chemistry, 2011, 286(3): 1737-1747. |
| 101 | SALAMANCA H H, ANTONYAK M A, CERIONE R A, et al. Inhibiting heat shock factor 1 in human cancer cells with a potent RNA aptamer[J]. PLoS One, 2014, 9(5): e96330. |
| 102 | VILABOA N, BORÉ A, MARTIN-SAAVEDRA F, et al. New inhibitor targeting human transcription factor HSF1: effects on the heat shock response and tumor cell survival[J]. Nucleic Acids Research, 2017, 45(10): 5797-5817. |
| 103 | HUBEL P, URBAN C, BERGANT V, et al. A protein-interaction network of interferon-stimulated genes extends the innate immune system landscape[J]. Nature Immunology, 2019, 20(4): 493-502. |
| 104 | MA J, ZHANG X F, FENG Y B, et al. Structural and functional study of apoptosis-linked gene-2·Heme-binding protein 2 interactions in HIV-1 production[J]. Journal of Biological Chemistry, 2016, 291(52): 26670-26685. |
| 105 | MCKAY T B, HJORTDAL J, SEJERSEN H, et al. Differential effects of hormones on cellular metabolism in keratoconus in vitro [J]. Scientific Reports, 2017, 7: 42896. |
| 106 | CICATIELLO A G, DI GIROLAMO D, DENTICE M. Metabolic effects of the intracellular regulation of thyroid hormone: Old players, new concepts[J]. Frontiers in Endocrinology, 2018, 9: 474. |
| 107 | BRUICE T C, BENKOVIC S J. Chemical basis for enzyme catalysis[J]. Biochemistry, 2000, 39(21): 6267-6274. |
| 108 | AQVIST J, et al. Entropy and Enzyme Catalysis.Accounts of Chemical Reserc, 2017, 50(2): 199-207. |
| 109 | DRUMMOND D A, BLOOM J D, ADAMI C, et al. Why highly expressed proteins evolve slowly[J]. Proceedings of the National Academy of Sciences of the United States of America, 2005, 102(40): 14338-14343. |
| 110 | BLOOM J D, WILKE C O, ARNOLD F H, et al. Stability and the evolvability of function in a model protein[J]. Biophysical Journal, 2004, 86(5): 2758-2764. |
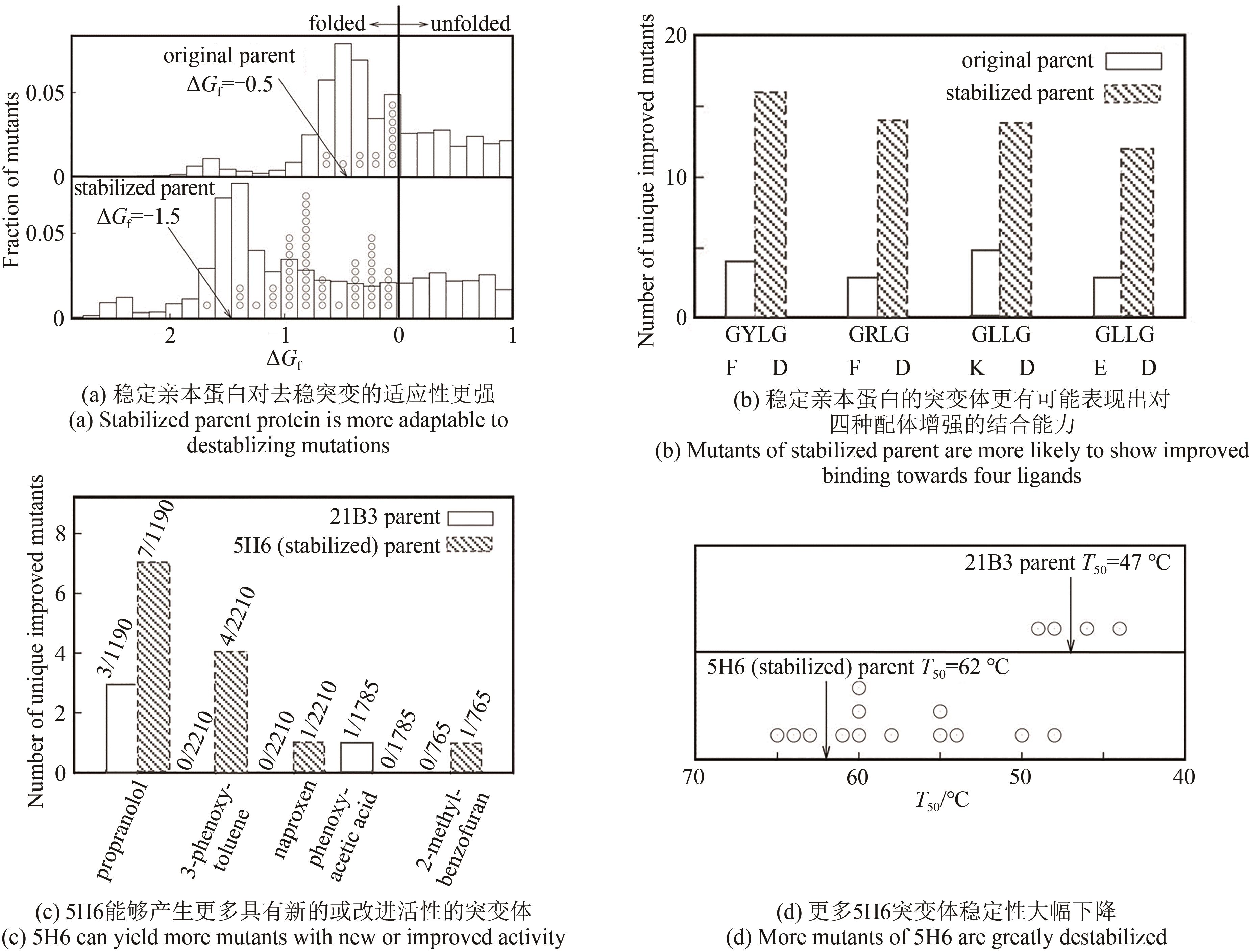
| 111 | BLOOM J D, LABTHAVIKUL S T, OTEY C R, et al. Protein stability promotes evolvability[J]. Proceedings of the National Academy of Sciences of the United States of America, 2006, 103(15): 5869-5874. |
| 112 | ZHENG J, GUO N, WAGNER A. Selection enhances protein evolvability by increasing mutational robustness and foldability[J]. Science, 2020, 370(6521): eabb5962. |
| 113 | JAMES L C, TAWFIK D S. Conformational diversity and protein evolution-a 60-year-old hypothesis revisited[J]. Trends in Biochemical Sciences, 2003, 28(7): 361-368. |
| 114 | TOKURIKI N, TAWFIK D S. Protein dynamism and evolvability[J]. Science, 2009, 324(5924): 203-207. |
| 115 | CREAN R M, GARDNER J M, KAMERLIN S C L. Harnessing conformational plasticity to generate designer enzymes[J]. Journal of the American Chemical Society, 2020, 142(26): 11324-11342. |
| 116 | QU, G, et al. Unlocking the stereoselectivity and substrate acceptance of enzymes: proline-induced loop engineering test[J]. Angewandte Chemie Internation Edition, 2022, 61(1): e202110793. |
| 117 | TOKURIKI N, TAWFIK D S. Chaperonin overexpression promotes genetic variation and enzyme evolution[J]. Nature, 2009, 459(7247): 668-673. |
| 118 | PHILLIPS A M, PONOMARENKO A I, CHEN K, et al. Destabilized adaptive influenza variants critical for innate immune system escape are potentiated by host chaperones[J]. PLoS Biology, 2018, 16(9): e3000008. |
| 119 | PHILLIPS A M, DOUD M B, GONZALEZ L O, et al. Enhanced ER proteostasis and temperature differentially impact the mutational tolerance of influenza hemagglutinin[J]. eLife, 2018, 7: e38795. |
| 120 | BANDYOPADHYAY A, SAXENA K, KASTURIA N, et al. Chemical chaperones assist intracellular folding to buffer mutational variations[J]. Nature Chemical Biology, 2012, 8(3): 238-245. |
| 121 | VELASQUEZ M T, RAMEZANI A, MANAL A, et al. Trimethylamine N-oxide: The good, the bad and the unknown[J]. Toxins, 2016, 8(11): 326. |
| 122 | QIAN Y Q, PATEL D, HARTL F U, et al. Nuclear magnetic resonance solution structure of the human Hsp40 (HDJ-1) J-domain[J]. Journal of Molecular Biology, 1996, 260(2): 224-235. |
| 123 | CAPLAN A J, CYR D M, DOUGLAS M G. Eukaryotic homologues of Escherichia coli dnaJ: A diverse protein family that functions with hsp70 stress proteins[J]. Molecular Biology of the Cell, 1993, 4(6): 555-563. |
| 124 | ACHARYA P, KUMAR R, TATU U. Chaperoning a cellular upheaval in malaria: heat shock proteins in Plasmodium falciparum [J]. Molecular and Biochemical Parasitology, 2007, 153(2): 85-94. |
| 125 | FELDHEIM D, ROTHBLATT J, SCHEKMAN R. Topology and functional domains of Sec63p, an endoplasmic reticulum membrane protein required for secretory protein translocation[J]. Molecular and Cellular Biology, 1992, 12(7): 3288-3296. |
| 126 | SCHLENSTEDT G, HARRIS S, RISSE B, et al. A yeast DnaJ homologue, Scj1p, can function in the endoplasmic reticulum with BiP/Kar2p via a conserved domain that specifies interactions with Hsp70s[J]. The Journal of Cell Biology, 1995, 129(4): 979-988. |
| 127 | NISHIKAWA S, ENDO T. The yeast JEM1p is a DnaJ-like protein of the endoplasmic reticulum membrane required for nuclear fusion[J]. The Journal of Biological Chemistry, 1997, 272(20): 12889-12892. |
| 128 | WANG J D, HERMAN C, TIPTON K A, et al. Directed evolution of substrate-optimized GroEL/S chaperonins[J]. Cell, 2002, 111(7): 1027-1039. |
| 129 | JACKREL M E, SHORTER J. Engineering enhanced protein disaggregases for neurodegenerative disease[J]. Prion, 2015, 9(2): 90-109. |
| [1] | FU Yu, ZHONG Fangrui. Recent advances in chemically driven enantioselective photobiocatalysis [J]. Synthetic Biology Journal, 2024, 5(5): 1021-1049. |
| [2] | TANG Zhijun, HU Youcai, LIU Wen. Enzymatic (4+2)- and (2+2)-cycloaddition reactions: fundamentals and applications of regio- and stereoselectivity [J]. Synthetic Biology Journal, 2024, 5(3): 401-407. |
| [3] | SUN Mengchu, LU Liangyu, SHEN Xiaolin, SUN Xinxiao, WANG Jia, YUAN Qipeng. Fluorescence detection-based high-throughput screening systems and devices facilitate cell factories construction [J]. Synthetic Biology Journal, 2023, 4(5): 947-965. |
| [4] | MING Yang, CHEN Bin, HUANG Xiaoqiang. Recent advances in photoenzymatic synthesis [J]. Synthetic Biology Journal, 2023, 4(4): 651-675. |
| [5] | KANG Liqi, TAN Pan, HONG Liang. Enzyme engineering in the age of artificial intelligence [J]. Synthetic Biology Journal, 2023, 4(3): 524-534. |
| [6] | RUAN Qingyun, HUANG Xin, MENG Zijun, QUAN Shu. Computational design and directed evolution strategies for optimizing protein stability [J]. Synthetic Biology Journal, 2023, 4(1): 5-29. |
| [7] | QI Yanping, ZHU Jin, ZHANG Kai, LIU Tong, WANG Yajie. Recent development of directed evolution in protein engineering [J]. Synthetic Biology Journal, 2022, 3(6): 1081-1108. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||