合成生物学 ›› 2021, Vol. 2 ›› Issue (5): 778-791.DOI: 10.12211/2096-8280.2021-018
解脂耶氏酵母中光控表达系统的构建及其应用研究
张萍1, 魏文平2, 周英1, 叶邦策1,2
- 1.华东理工大学微分析与生物系统工程实验室,生物反应器工程国家重点实验室,上海 200237
2.浙江工业大学工程生物学与健康研究中心,长三角绿色制药协同创新中心,药学院,浙江 杭州 310014
-
收稿日期:2021-02-05修回日期:2021-08-13出版日期:2021-10-31发布日期:2021-11-19 -
通讯作者:周英,叶邦策 -
作者简介:张萍 (1998—),女,博士研究生。研究方向为合成生物学及天然产物的微生物合成。E-mail:pingzh@mail.ecust.edu.cn周英 (1979—),女,硕士生导师,副教授。研究方向为合成生物学及工程化肠道微生物。E-mail:zhouying@ecust.edu.cn叶邦策 (1967—),男,博士生导师,教授。研究方向为合成生物学及智能生物系统设计,生物分析及生物传感研究。E-mail:bcye@ecust.edu.cn -
基金资助:国家重点研发计划“合成生物学”重点专项(2018YFA0900404);国家自然科学基金(31730004)
Construction of a light-controlled expression system and its application in Yarrowia lipolytica
ZHANG Ping1, WEI Wenping2, ZHOU Ying1, YE Bangce1,2
- 1.Laboratory of Biosystems and Microanalysis,State Key Laboratory of Bioreactor Engineering,East China University of Science and Technology,Shanghai 200237,China
2.Institute of Engineering Biology and Health,Collaborative Innovation Center of Yangtze River Delta Region Green Pharmaceuticals,College of Pharmaceutical Sciences,Zhejiang University of Technology,Hangzhou 310014,Zhejiang,China
-
Received:2021-02-05Revised:2021-08-13Online:2021-10-31Published:2021-11-19 -
Contact:ZHOU Ying, YE Bangce
摘要:
光作为诱导因子具有响应速度快、无损、可逆和独特的时空特异性等优势,已广泛应用于生物底盘细胞的工程化改造中。本研究构建重组解脂耶氏酵母(Yarrowia lipolytica)的光控表达系统,成功实现了对香豆酸和柚皮素合成。该光控系统基于绿光响应调控因子CarH以及转录激活因子VPR-HSF1,构建了光响应复合体CarH-VPRH,而该复合体在绿光照射条件下无法聚合,从而不能调控目标基因的转录与表达。结果表明,基于光控因子CarH-VPRH、诱导型启动子2CarOTEF以及报告基因mCherry构建的光诱导传感器能够显著响应光照,并在黑暗条件下激活mCherry转录,72 h和120 h时产生的荧光信号分别是光照下的43倍和143倍。在此基础上,将该元件应用于对香豆酸及柚皮素的生物合成及合成途径的动态调控中,实现黑暗条件下的对香豆酸产量是绿光诱导下的2.0倍,达到99.1 mg/L;柚皮素产量是绿光诱导下的2.6倍达到117.1 mg/L;在细胞生长方面,绿光照射组的生长量高于黑暗组,特别是在绿光照射24 h,然后切换成黑暗条件,细胞的生长量有较为明显的优势,显示了绿光诱导系统在调控Y. lipolytica中产物合成与细胞生长平衡的潜力。因此该绿光响应基因调控系统可有效应用于Y. lipolytica细胞中目标基因的转录调控,具有廉价易得、无毒害、灵活添加及易去除等优势,因此具有大规模合成和生产目的产物的潜力。
中图分类号:
引用本文
张萍, 魏文平, 周英, 叶邦策. 解脂耶氏酵母中光控表达系统的构建及其应用研究[J]. 合成生物学, 2021, 2(5): 778-791.
ZHANG Ping, WEI Wenping, ZHOU Ying, YE Bangce. Construction of a light-controlled expression system and its application in Yarrowia lipolytica[J]. Synthetic Biology Journal, 2021, 2(5): 778-791.
| Type | Characteristics | Source |
|---|---|---|
| Strains | ||
| Yarrowia lipolytica Po1f | URA3-, Leu2- | This laboratory |
| Yl-CVH | Po1f with plasmid p13-CV-2CarO-Tm | This study |
| Yl-CA | Po1f with plasmid p13-CV-2CarO-TAL | This study |
| Yl-NA | Yl-CA with plasmid p12-4CC | This study |
| Plasmids | ||
| pINA1312 | The plasmid contains a strong constitutive promoter hp4d, ura3 gene expression cassette and kanamycin resistance gene | This laboratory |
| pINA1269 | The plasmid contains a strong constitutive promoter hp4d, leu2 gene expression cassette and ampicillin resistance gene | This laboratory |
| pSET152-Pj23119-2CarO-GFP | The plasmid is a commonly used plasmid pSET152 of Streptomyces which contains CarO site | This laboratory |
| pUC57-CarH | The plasmid contains codon optimized CarH gene | This laboratory |
| pUC57-VPRH | The plasmid contains codon optimized VPR and HSF1 genes | This laboratory |
| p13PxoTAL | The plasmid pINA1312 contains codon optimized tal gene and Ptef promoter | This laboratory |
| p13-Tm | The plasmid pINA1312 contains the mcherry expression module controlled by the TEF promoter | This laboratory |
| p13-CV | The plasmid pINA1312 contains the CarH-VPRH expression module controlled by the hp4d promoter | This study |
| p13-CV-2CarO-Tm | The plasmid is a fluorescent protein expression module 2CarO-TEF-mcherry controlled by an inducible promoter inserted into the plasmid p13-CV | This study |
| p13-CV-2CarO-TAL | The plasmid is a TAL expression module controlled by an inducible promoter inserted into the plasmid p13-CV | This study |
| p12-4CL | The plasmid pINA1269 contains codon optimized 4cl gene with FBAin promoter | This laboratory |
| pCas-AXPCHSCHI | The plasmid contains codon optimized chs gene with Pexp1 promoter and codon optimized chi gene with FBAin promoter, ampicillin resistance gene | This laboratory |
| p12-4CC | The plasmid pINA1269 contains the 4cl gene with FBAin promoter, chs gene with Pexp1 promoter and chi gene with FBAin promoter | This study |
表1 本研究所用的质粒和菌株
Tab. 1 Plasmids and strains used in this study
| Type | Characteristics | Source |
|---|---|---|
| Strains | ||
| Yarrowia lipolytica Po1f | URA3-, Leu2- | This laboratory |
| Yl-CVH | Po1f with plasmid p13-CV-2CarO-Tm | This study |
| Yl-CA | Po1f with plasmid p13-CV-2CarO-TAL | This study |
| Yl-NA | Yl-CA with plasmid p12-4CC | This study |
| Plasmids | ||
| pINA1312 | The plasmid contains a strong constitutive promoter hp4d, ura3 gene expression cassette and kanamycin resistance gene | This laboratory |
| pINA1269 | The plasmid contains a strong constitutive promoter hp4d, leu2 gene expression cassette and ampicillin resistance gene | This laboratory |
| pSET152-Pj23119-2CarO-GFP | The plasmid is a commonly used plasmid pSET152 of Streptomyces which contains CarO site | This laboratory |
| pUC57-CarH | The plasmid contains codon optimized CarH gene | This laboratory |
| pUC57-VPRH | The plasmid contains codon optimized VPR and HSF1 genes | This laboratory |
| p13PxoTAL | The plasmid pINA1312 contains codon optimized tal gene and Ptef promoter | This laboratory |
| p13-Tm | The plasmid pINA1312 contains the mcherry expression module controlled by the TEF promoter | This laboratory |
| p13-CV | The plasmid pINA1312 contains the CarH-VPRH expression module controlled by the hp4d promoter | This study |
| p13-CV-2CarO-Tm | The plasmid is a fluorescent protein expression module 2CarO-TEF-mcherry controlled by an inducible promoter inserted into the plasmid p13-CV | This study |
| p13-CV-2CarO-TAL | The plasmid is a TAL expression module controlled by an inducible promoter inserted into the plasmid p13-CV | This study |
| p12-4CL | The plasmid pINA1269 contains codon optimized 4cl gene with FBAin promoter | This laboratory |
| pCas-AXPCHSCHI | The plasmid contains codon optimized chs gene with Pexp1 promoter and codon optimized chi gene with FBAin promoter, ampicillin resistance gene | This laboratory |
| p12-4CC | The plasmid pINA1269 contains the 4cl gene with FBAin promoter, chs gene with Pexp1 promoter and chi gene with FBAin promoter | This study |
| Primers | Sequence (5′- 3′) |
|---|---|
| 1312CarH-F | acacatacaaccacacacatc |
| CarH-VPRH-R | tcgtccttgtagtcggatccaga |
| CarH-VPRH-F | tgagaaggaggctatc |
| KpnIHSF1-R | tggggacaggccatgga |
| SalI-Pj23119F | aagcactttgctagataga |
| CarO-TEF-R | acggtggtcttttatacccttggctcgtcgtccttgtagtccatcatatgtctagct |
| CarO-TEF-F | agctagacatatgatggactacaaggacgacgagccaagggtataaaagaccaccgt |
| StuI-XPR2-277R | agctctgtacaccgagaaac |
| TEF-TAL-F | agcatttccttctgagtataagaatcattcaaagctcctcgacctacttcgcagtcgc |
| 1269-SpeI-XPR2-F | aacgcgcgaggcagcagatcc |
| FBAin-CYCt-R | gggacgctcgaaggctttaatttgccagtgtacgcagtactatagaggaacaattgcc |
| FBAin-CYCt-F | tgttcctctatagtactgcgtacactggcaaattaaagccttcgagcgtcccaaaacc |
| 1269-SpeI-XPR2-R | agatccgcggccgcataggcc |
表2 本研究中使用的引物
Tab. 2 Primers used in this study
| Primers | Sequence (5′- 3′) |
|---|---|
| 1312CarH-F | acacatacaaccacacacatc |
| CarH-VPRH-R | tcgtccttgtagtcggatccaga |
| CarH-VPRH-F | tgagaaggaggctatc |
| KpnIHSF1-R | tggggacaggccatgga |
| SalI-Pj23119F | aagcactttgctagataga |
| CarO-TEF-R | acggtggtcttttatacccttggctcgtcgtccttgtagtccatcatatgtctagct |
| CarO-TEF-F | agctagacatatgatggactacaaggacgacgagccaagggtataaaagaccaccgt |
| StuI-XPR2-277R | agctctgtacaccgagaaac |
| TEF-TAL-F | agcatttccttctgagtataagaatcattcaaagctcctcgacctacttcgcagtcgc |
| 1269-SpeI-XPR2-F | aacgcgcgaggcagcagatcc |
| FBAin-CYCt-R | gggacgctcgaaggctttaatttgccagtgtacgcagtactatagaggaacaattgcc |
| FBAin-CYCt-F | tgttcctctatagtactgcgtacactggcaaattaaagccttcgagcgtcccaaaacc |
| 1269-SpeI-XPR2-R | agatccgcggccgcataggcc |
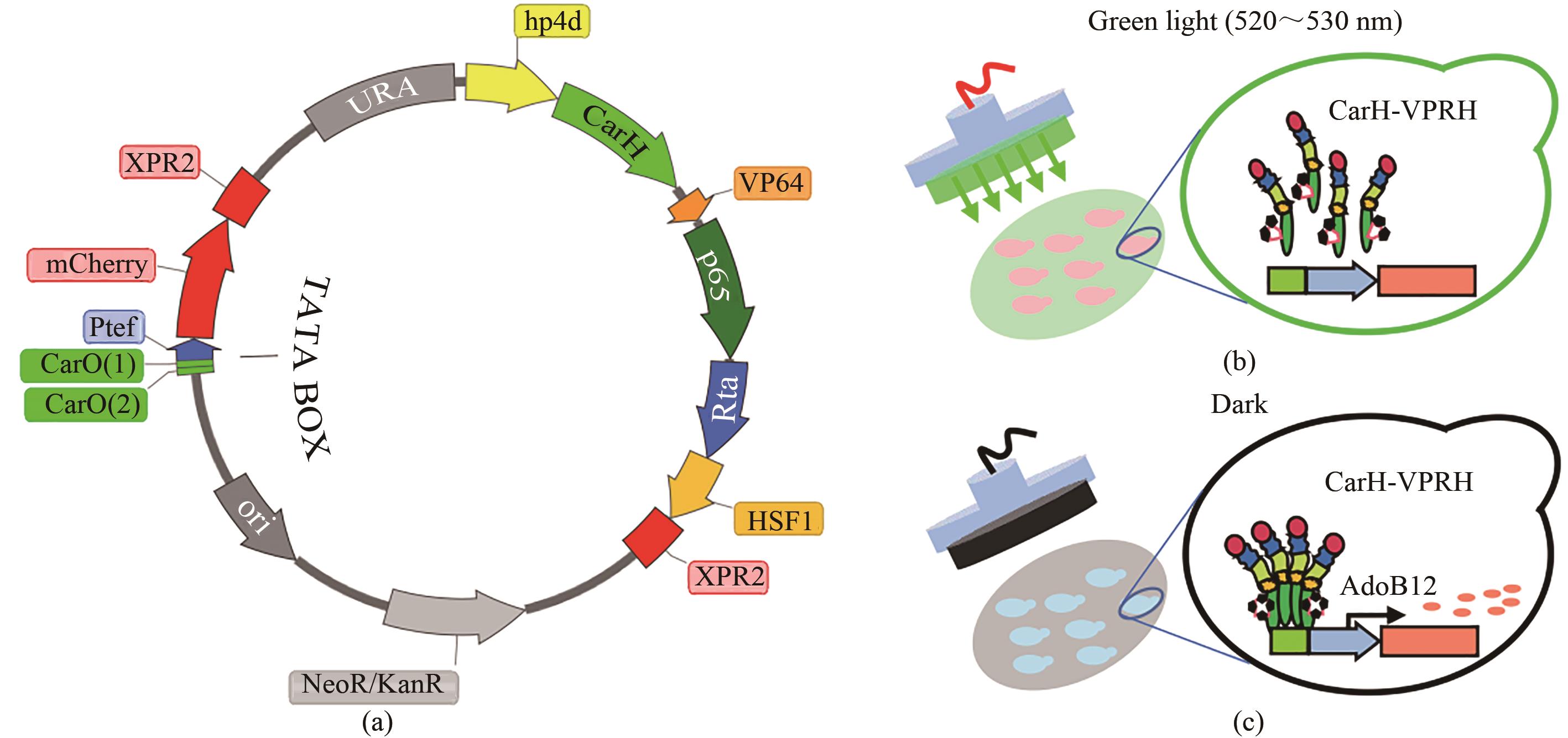
图1 绿光响应传感器的设计((a)绿光响应传感器工具质粒设计,绿光响应转录因子CarH-VPRH由组成型启动子hp4d驱动表达,报告基因mcherry由诱导型启动子2CarO-TEF驱动表达;(b)绿光光照条件下的“Off”模式;(c)黑暗条件下的启动表达“On”模式)
Fig. 1 Design of the green light response sensor((a)Tool plasmid design of green light response sensor; green light response transcription factor CarH-VPRH is driven by constitutive promoter hp4d, expression of reporter gene mcherry is driven by inducible promoter 2CarO-TEF; (b)"Off" mode under green light; (c) "On" mode of expression in dark condition)
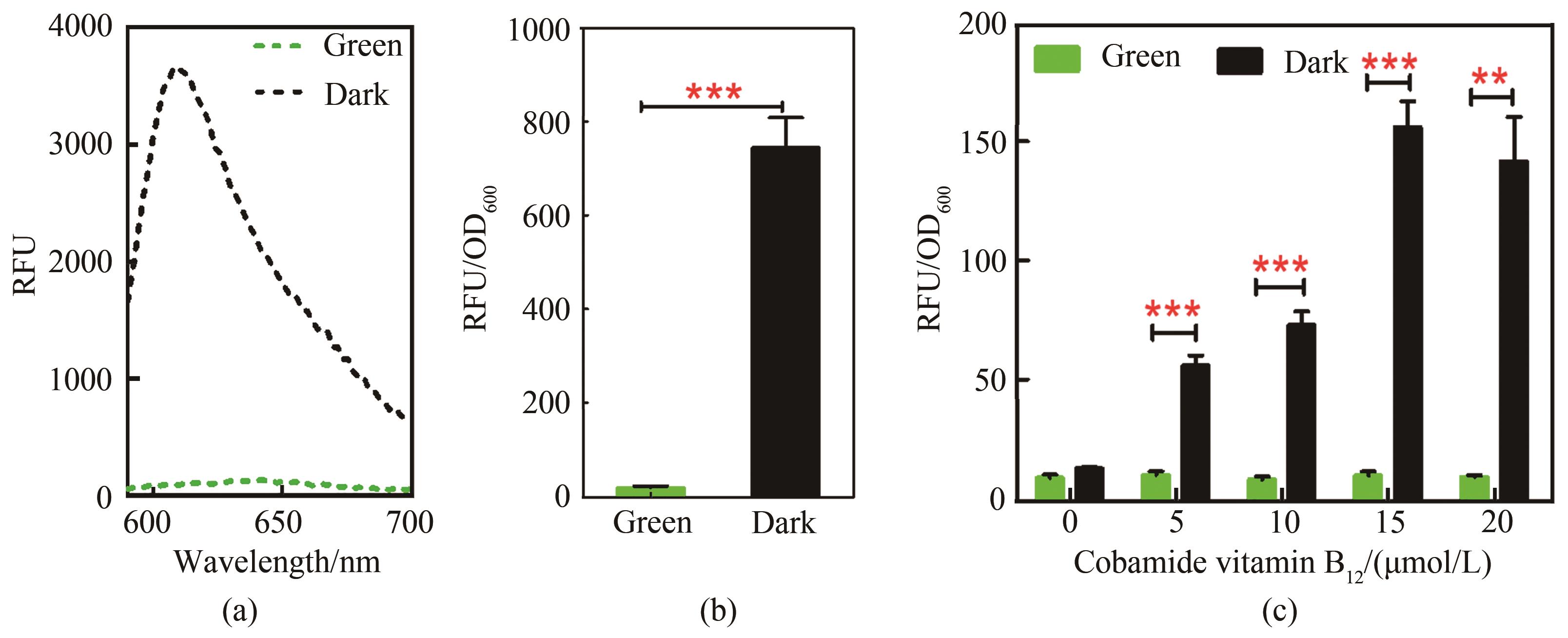
图2 绿光响应传感器的性能表征(a)传感菌株的荧光蛋白检测光谱图,黑色曲线为黑暗组,绿色曲线为绿光照射组,激发光波长570 nm,发射光波长区间590~700 nm;(b)传感菌株绿光响应信号输出差异,测试菌株为Yl-CVH,培养基为YPD,培养时间72 h;(c)腺苷钴胺素浓度对光控诱导系统的影响,培养时间48 h
Fig. 2 Performance of the green light response sensor(a)Fluorescence protein detection spectrogram of sensing strain. Black curve is dark group, and green curve is green light irradiation group, excitation light: 570 nm, emission light: 590~700 nm; (b)Green light response signal output difference of strains; Tested strain was Yl-CVH; Culture medium YPD, 72 h; (c)Effect of cobamide vitamin B12 concentration on light-controlled induction system,48 h
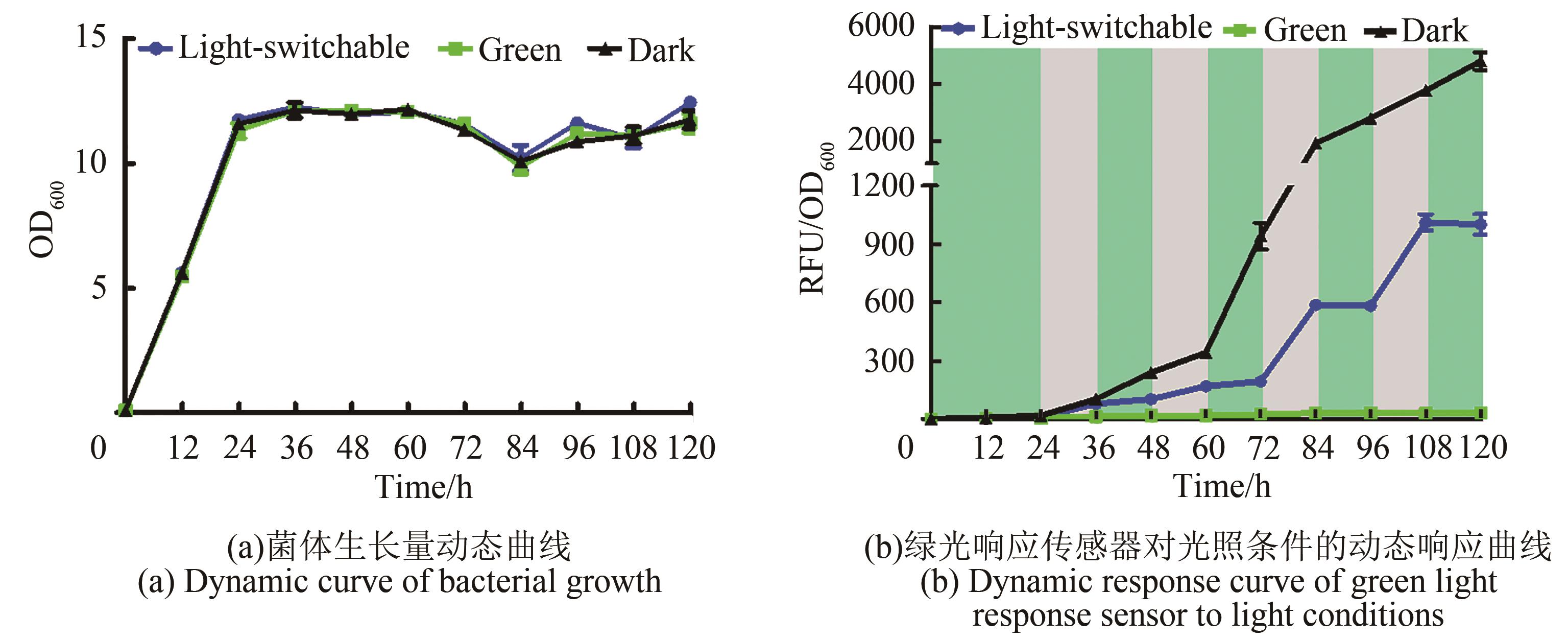
图3 绿光响应传感器对光照条件的动态响应(图中绿色区域代表绿光照射条件,灰色区域代表黑暗条件。培养条件:YPD液体培养基,30 ℃,220 r/min培养,并在0 h、24 h、48 h、72 h和96 h补加15 μmol/L腺苷钴胺素;荧光值测定条件:激发光580 nm,发射光608 nm;取样时间间隔12 h)
Fig. 3 Dynamic response of the green light response sensor to light conditions(Green area in figure represents green light irradiation condition, and gray area represents dark condition. Culture conditions: YPD liquid medium, 30 ℃, 220 r/min, and supplemented with 15 μmol/L cobamide vitamin B12 at 0 h, 24 h, 48 h, 72 h and 96 h; Fluorescence value measurement conditions: excitation light: 580 nm, emission light: 608 nm; Sampling time interval: 12 h)
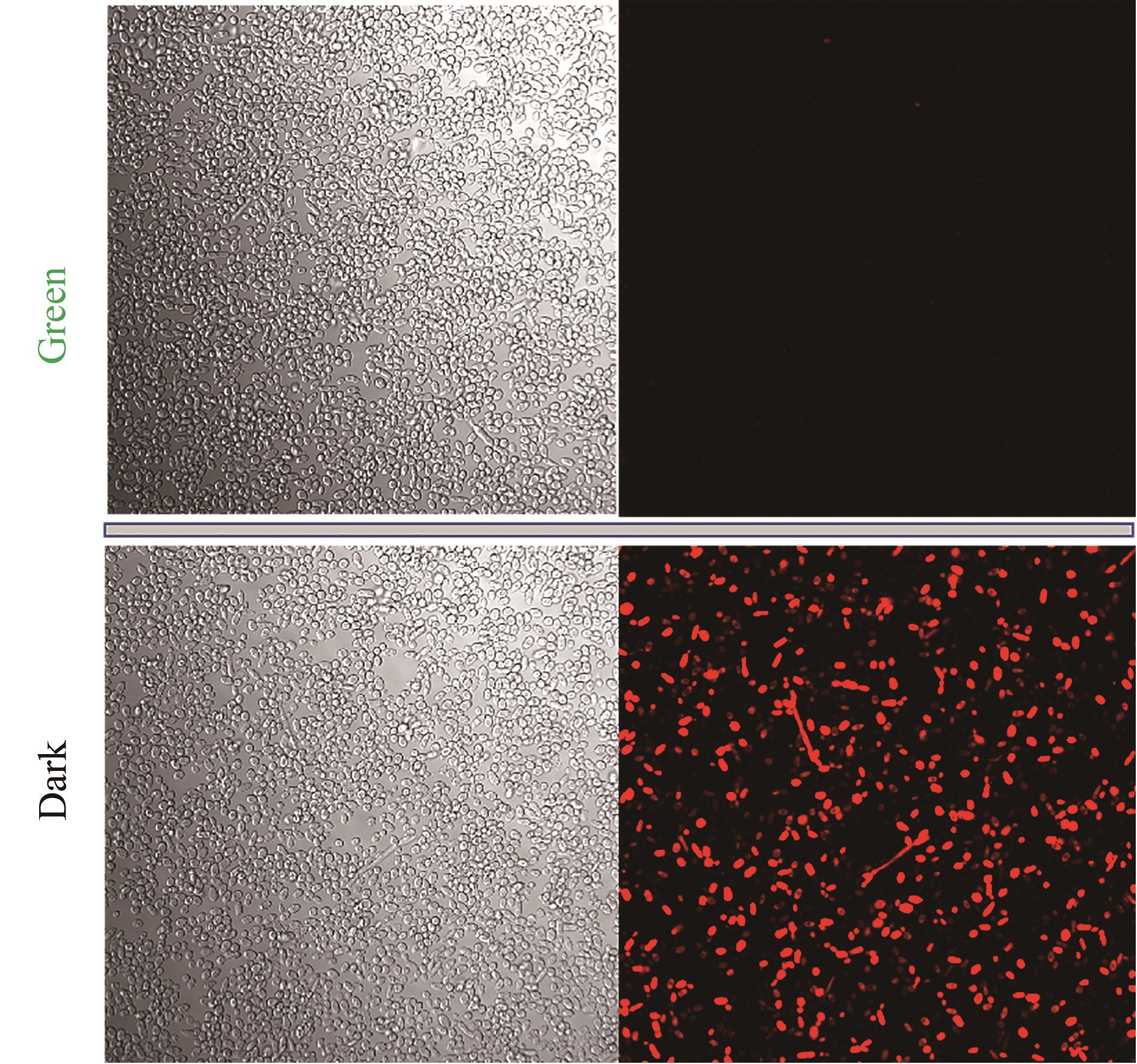

图4 绿光调控细胞成像(成像菌种悬浮保存介质为pH 7.4的PBS缓冲液;实验菌种培养条件为YPD培养基,并在0 h加入15 μmol/L腺苷钴胺素,取样时间72 h)
Fig. 4 Cell imagines regulated by green light irradiation(Imaging strains were suspended and preserved in PBS buffer solution with pH 7.4. Experimental strains were cultured in YPD medium, and added 15 μmol/L cobamide vitamin B12 at 0 h. The strain samples were obtained at 72 h)
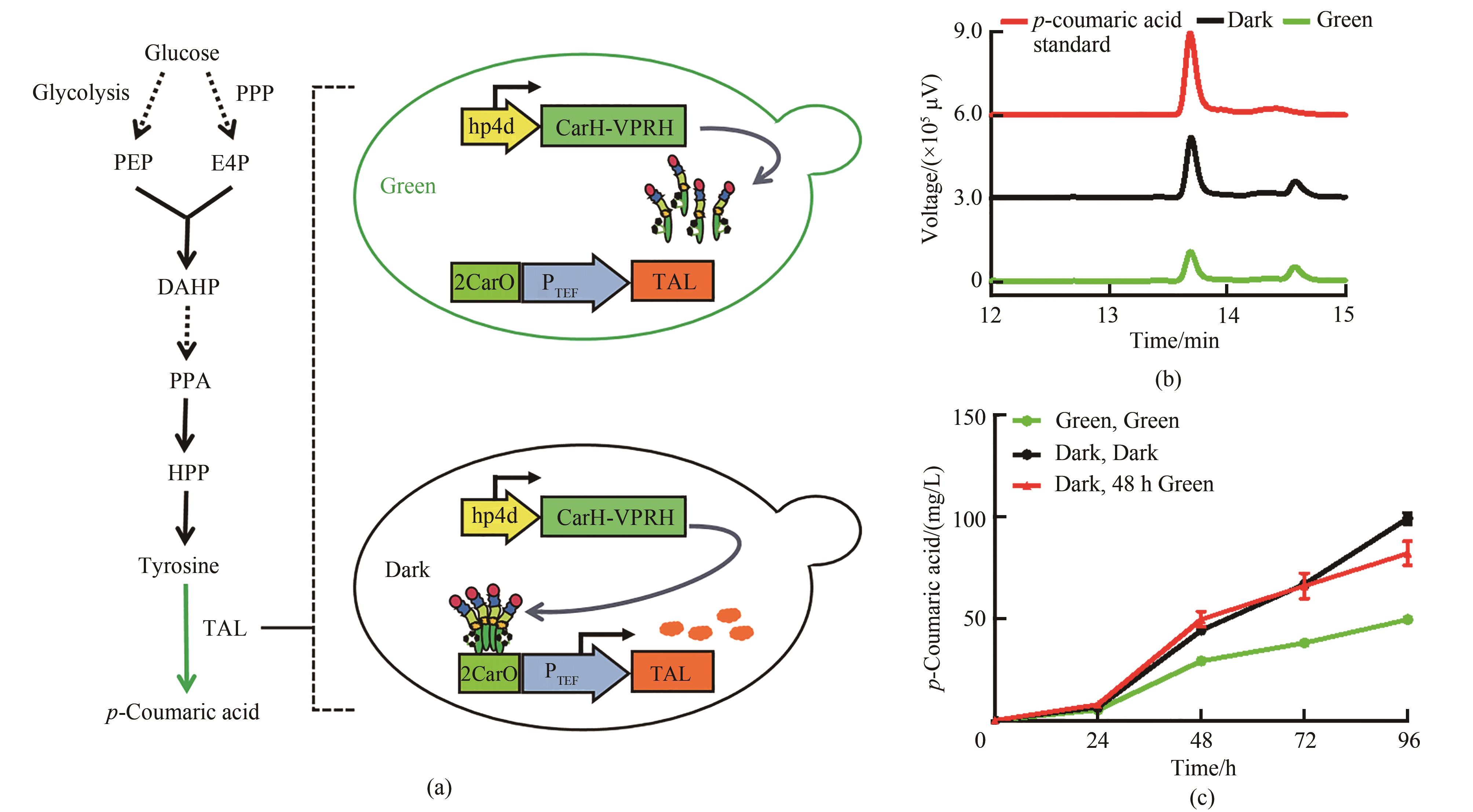
图5 绿光调控对香豆酸合成(a)绿光控制对香豆酸合成途径。PPP—磷酸戊糖途径;PEP—磷酸烯醇式丙酮酸;E4P—赤藓糖-4-磷酸;DAHP—3-脱氧-D-阿拉伯庚酮糖-7-磷酸;PPA—预苯酸;HPP—4-羟基-苯丙酮酸;TAL—酪氨酸解氨酶(b)菌株Yl-CA对香豆酸HPLC检测图谱。其中红色曲线代表50 mg/L的对香豆酸标准品,黑色曲线代表黑暗条件下的菌株,绿色曲线代表绿光照射下的菌株(c)不同光照条件下菌株Yl-CA对香豆酸含量变化。 培养基YPD,均在0 h加入15 μmol/L腺苷钴胺素
Fig. 5 p-Coumaric acid synthesis regulated by green light irradiation(a) p-Coumaric acid synthesis pathway controlled by green light. PPP—pentose phosphate pathway; PEP—phosphoenolpyruvate; E4P— erythrose 4-phosphate; DAHP—3-deoxy-D-arabino-heptulosonic acid-7-phosphate; PPA—prephenate; HPP—4-hydroxy-phenylpyruvate; TAL—tyrosine ammonia lyase(b) HPLC chromatogram of strain Yl-CA. The red curve represents the 50 mg/L p-coumaric acid standard, the black curve represents the strain under dark conditions, and the green curve represents the strain under green light(c) Changes of p-coumaric acid concentration of strain Yl-CA under different light conditions; Medium YPD, and all added 15 μmol/L cobamide vitamin B12 at 0 h
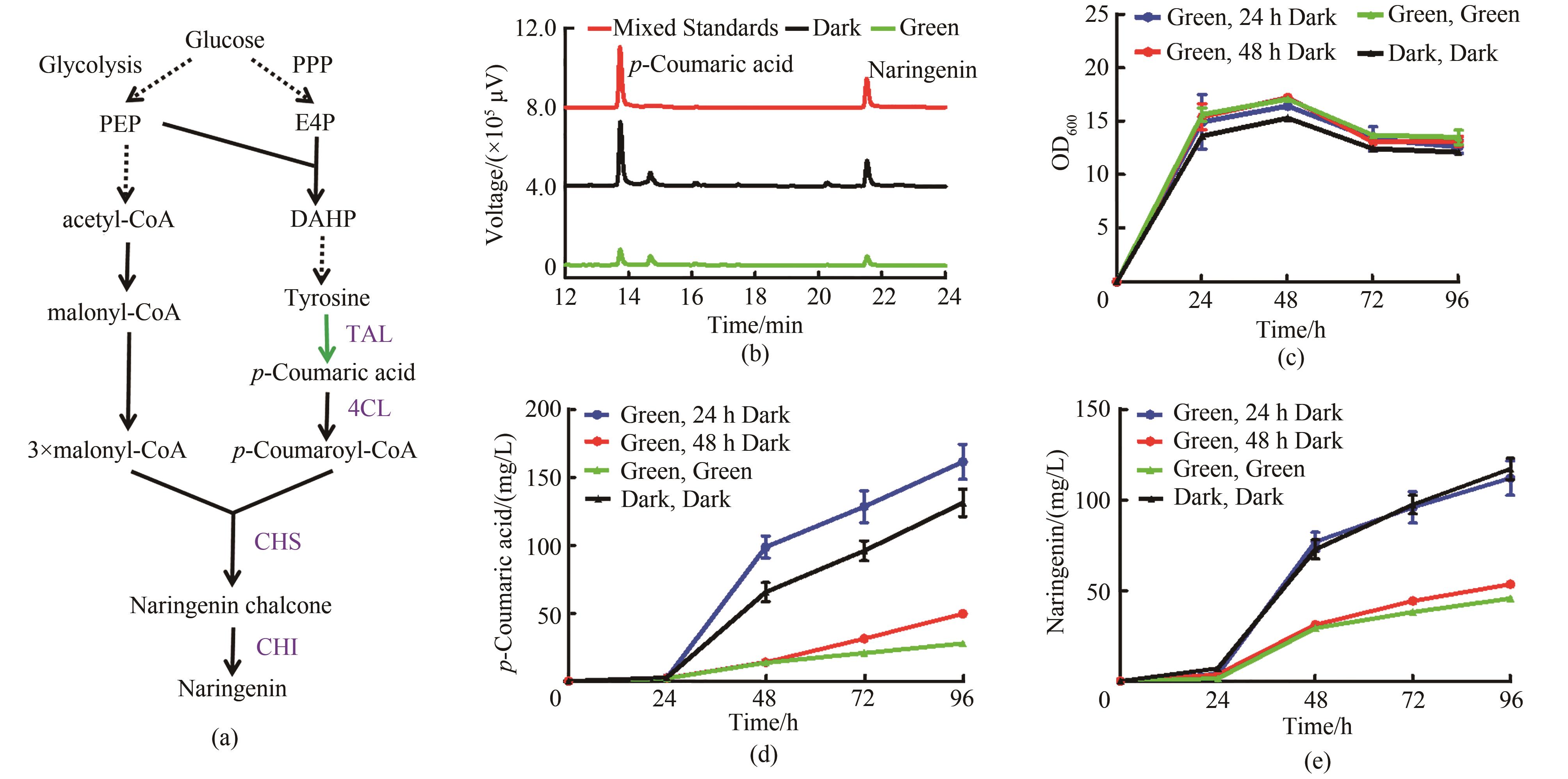
图6 绿光调控柚皮素合成(a)绿光控制柚皮素合成途径。PPP—磷酸戊糖途径;PEP—磷酸烯醇式丙酮酸;E4P—赤藓糖-4-磷酸;DAHP—3-脱氧-D-阿拉伯庚酮糖-7-磷酸;TAL—酪氨酸解氨酶;4CL—4-香豆酸辅酶A连接酶;CHS—查尔酮合酶;CHI—查尔酮异构酶(b)菌株Yl-NA对香豆酸及柚皮素HPLC检测图谱,其中红色曲线代表50 mg/L对香豆酸和50 mg/L柚皮素的混合标准品,黑色曲线代表黑暗条件下的菌株,绿色曲线代表绿光照射下的菌株(c)菌株Yl-NA的生长曲线(d)菌株Yl-NA的对香豆酸发酵曲线(e)菌株Yl-NA的柚皮素发酵曲线;培养基YPD,均在0 h加入15 μmol/L腺苷钴胺素,其中光照条件在24 h和48 h由光照切换为黑暗的两组分别在24 h和48 h再次添加15 μmol/L腺苷钴胺素
Fig. 6 Naringenin synthesis regulated by green light irradiation(a) Naringenin synthesis pathway controlled by green light. PPP—pentose phosphate pathway; PEP—phosphoenolpyruvate; E4P—erythrose 4-phosphate; DAHP—3-deoxy-D-arabino-heptulosonic acid-7-phosphate; TAL—tyrosine ammonia lyase; 4CL—4-coumaroyl-CoA ligase; CHS—chalcone synthase; CHI—chalcone isomerase(b) HPLC chromatogram of strain Yl-NA. The red curve represents the 50 mg/L p-coumaric acid standard and 50 mg/L naringenin standard, the black curve represents the strain under dark conditions, and the green curve represents the strain under green light(c) Growth curve of strain Yl-NA(d) p-Coumaric acid fermentation curve of strain Yl-NA(e) Naringenin fermentation curve of strain Yl-NA; YPD medium, all added 15 μmol/L cobamide vitamin B12 at 0 h, and the two groups whose light conditions were switched from light to dark at 24 h and 48 h were added 15 μv cobamide vitamin B12 again at 24 h and 48 h, respectively
| 1 | LIU W S, STEWART C N. Plant synthetic biology[J]. Trends in Plant Science, 2015, 20(5): 309-317. |
| 2 | 尤迪, 叶邦策. 从翻译后修饰角度解析人工合成途径与底盘细胞的适配性[J]. 合成生物学, 2020, 1(2): 212-225. |
| YOU Di, YE Bangce. Compatibility between synthetic pathway and chassis cells from the viewpoint of post-translational modifications[J]. Synthetic Biology Journal, 2020, 1(2): 212-225. | |
| 3 | 陈显军, 左方婷, 杨弋. 光控基因表达系统[J]. 生命科学, 2019, 31(4): 343-356. |
| CHEN Xianjun, ZUO Fangting, YANG Yi. Light-regulated gene expression systems[J]. Chineses Bulletin of Life Sciences, 2019, 31(4): 343-356. | |
| 4 | CHATELLE C, OCHOA-FERNANDEZ R, ENGESSER R, et al. A green-light-responsive system for the control of transgene expression in mammalian and plant cells[J]. ACS Synthetic Biology, 2018, 7(5): 1349-1358. |
| 5 | JAYARAMAN P, DEVARAJAN K, CHUA T K, et al. Blue light-mediated transcriptional activation and repression of gene expression in bacteria[J]. Nucleic Acids Research, 2016, 44(14): 6994-7005. |
| 6 | ZHAO E M, ZHANG Y F, MEHL J, et al. Optogenetic regulation of engineered cellular metabolism for microbial chemical production[J]. Nature, 2018, 555(7698): 683-687. |
| 7 | ZHAO E M, SUEK N, WILSON M Z, et al. Light-based control of metabolic flux through assembly of synthetic organelles[J]. Nature Chemical Biology, 2019, 15(6): 589-597. |
| 8 | TANDAR S T, SENOO S, TOYA Y, et al. Optogenetic switch for controlling the central metabolic flux of Escherichia coli [J]. Metabolic Engineering, 2019, 55: 68-75. |
| 9 | CRAVENS A, PAYNE J, SMOLKE C D. Synthetic biology strategies for microbial biosynthesis of plant natural products[J]. Nature Communications, 2019, 10(1): 2142. |
| 10 | 饶聪, 云轩, 虞沂, 等. 微生物药物的合成生物学研究进展[J]. 合成生物学, 2020, 1(1): 92-102. |
| RAO Cong, YUN Xuan, YU Yi, et al. Recent progress of synthetic biology applications in microbial pharmaceuticals research[J]. Synthetic Biology Journal, 2020, 1(1): 92-102. | |
| 11 | MA J B, GU Y, MARSAFARI M, et al. Synthetic biology, systems biology, and metabolic engineering of Yarrowia lipolytica toward a sustainable biorefinery platform[J]. Journal of Industrial Microbiology & Biotechnology, 2020, 47(9/10): 845-862. |
| 12 | DARVISHI F, ARIANA M, MARELLA E R, et al. Advances in synthetic biology of oleaginous yeast Yarrowia lipolytica for producing non-native chemicals[J]. Applied Microbiology and Biotechnology, 2018, 102(14): 5925-5938. |
| 13 | LEDESMA-AMARO R, LAZAR Z, RAKICKA M, et al. Metabolic engineering of Yarrowia lipolytica to produce chemicals and fuels from xylose[J]. Metabolic Engineering, 2016, 38: 115-124. |
| 14 | LI H B, ALPER H S. Enabling xylose utilization in Yarrowia lipolytica for lipid production[J]. Biotechnology Journal, 2016, 11(9): 1230-1240. |
| 15 | MARSAFARI M, SAMIZADEH H, RABIEI B, et al. Biotechnological production of flavonoids: an update on plant metabolic engineering, microbial host selection and genetically encoded biosensors[J]. Biotechnology Journal, 2020, 15(8): 1900432. |
| 16 | ABDEL-MAWGOUD A M, MARKHAM K A, PALMER C M, et al. Metabolic engineering in the host Yarrowia lipolytica [J]. Metabolic Engineering, 2018, 50: 192-208. |
| 17 | MARKHAM K A, PALMER C M, CHWATKO M, et al. Rewiring Yarrowia lipolytica toward triacetic acid lactone for materials generation[J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(9): 2096-2101. |
| 18 | PALMER C M, MILLER K K, NGUYEN A, et al. Engineering 4-coumaroyl-CoA derived polyketide production in Yarrowia lipolytica through a β-oxidation mediated strategy[J]. Metabolic Engineering, 2020, 57: 174-181. |
| 19 | SÁEZ-SÁEZ J, WANG G, MARELLA E R, et al. Engineering the oleaginous yeast Yarrowia lipolytica for high-level resveratrol production[J]. Metabolic Engineering, 2020, 62: 51-61. |
| 20 | HE Q, SZCZEPAŃSKA P, YUZBASHEV T, et al. De novo production of resveratrol from glycerol by engineering different metabolic pathways in Yarrowia lipolytica [J]. Metabolic Engineering Communications, 2020, 11: e00146. |
| 21 | CAO X, LV Y B, CHEN J, et al. Metabolic engineering of oleaginous yeast Yarrowia lipolytica for limonene overproduction[J]. Biotechnology for Biofuels, 2016, 9(1): 214. |
| 22 | ARNESEN J A, KILDEGAARD K R, CERNUDA PASTOR M, et al. Yarrowia lipolytica strains engineered for the production of terpenoids[J]. Frontiers in Bioengineering and Biotechnology, 2020, 8: 945. |
| 23 | QIAN Y D, TAN S Y, DONG G R, et al. Increased campesterol synthesis by improving lipid content in engineered Yarrowia lipolytica [J]. Applied Microbiology and Biotechnology, 2020, 104(16): 7165-7175. |
| 24 | LIU H, WANG F, DENG L, et al. Genetic and bioprocess engineering to improve squalene production in Yarrowia lipolytica [J]. Bioresource Technology, 2020, 317: 123991. |
| 25 | LIU X N, DING W T, JIANG H F. Engineering microbial cell factories for the production of plant natural products: from design principles to industrial-scale production[J]. Microbial Cell Factories, 2017, 16(1): 125. |
| 26 | HUSSAIN M S, WHEELDON I, BLENNER M A. A strong hybrid fatty acid inducible transcriptional sensor built from Yarrowia lipolytica upstream activating and regulatory sequences[J]. Biotechnology Journal, 2017, 12(10): 11. |
| 27 | XIONG X C, CHEN S L. Expanding toolbox for genes expression of Yarrowia lipolytica to include novel inducible, repressible, and hybrid promoters[J]. ACS Synthetic Biology, 2020, 9(8): 2208-2213. |
| 28 | WEI W P, SHANG Y Z, ZHANG P, et al. Engineering prokaryotic transcriptional activator XylR as a xylose-inducible biosensor for transcription activation in yeast[J]. ACS Synthetic Biology, 2020, 9(5): 1022-1029. |
| 29 | 牛福星, 杜云平, 黄远斌, 等. 工程微生物合成苯丙酸类化合物及其衍生物的研究进展[J]. 合成生物学, 2020, 1(3): 337-357. |
| NIU Fuxing, DU Yunping, HUANG Yuanbin, et al. Recent advances in the production of phenylpropanoic acids and their derivatives by genetically engineered microorganisms[J]. Synthetic Biology Journal, 2020, 1(3): 337-357. | |
| 30 | LI Y K, LI J, QIAN B B, et al. De novo biosynthesis of p-coumaric acid in E. coli with a trans-cinnamic acid 4-hydroxylase from the amaryllidaceae plant Lycoris aurea [J]. Molecules, 2018, 23(12): 19. |
| 31 | MA J, GU Y, XU P. A roadmap to engineering antiviral natural products synthesis in microbes[J]. Current Opinion in Biotechnology, 2020, 66: 140-149. |
| 32 | MUHAMMAD A, FENG X, RASOOL A, et al. Production of plant natural products through engineered Yarrowia lipolytica [J]. Biotechnology Advances, 2020, 43: 107555. |
| 33 | SANTOS C N S, KOFFAS M, STEPHANOPOULOS G. Optimization of a heterologous pathway for the production of flavonoids from glucose[J]. Metabolic Engineering, 2011, 13(4): 392-400. |
| 34 | WEI W P, ZHANG P, SHANG Y Z, et al. Metabolically engineering of Yarrowia lipolytica for the biosynthesis of naringenin from a mixture of glucose and xylose[J]. Bioresource Technology, 2020, 314: 123726. |
| 35 | KOOPMAN F, BEEKWILDER J, CRIMI B, et al. De novo production of the flavonoid naringenin in engineered Saccharomyces cerevisiae [J]. Microbial Cell Factories, 2012, 11(1): 155. |
| 36 | JOST M, FERNÁNDEZ-ZAPATA J, POLANCO M C, et al. Structural basis for gene regulation by a B12-dependent photoreceptor[J]. Nature, 2015, 526(7574): 536-541. |
| 37 | MADZAK C, TRÉTON B, BLANCHIN-ROLAND S. Strong hybrid promoters and integrative expression/secretion vectors for quasi-constitutive expression of heterologous proteins in the yeast Yarrowia lipolytica [J]. Journal of Molecular Microbiology and Biotechnology, 2000, 2(2): 207-216. |
| 38 | BLANCHIN-ROLAND S, CORDERO OTERO R R, GAILLARDIN C. Two upstream activation sequences control the expression of the XPR2 gene in the yeast Yarrowia lipolytica [J]. Molecular and Cellular Biology, 1994, 14(1): 327-338. |
| 39 | GAO S, LÜ Y B, ZENG W Z, et al. Efficient biosynthesis of (2S)-naringenin from p-coumaric acid in Saccharomyces cerevisiae [J]. Journal of Agricultural and Food Chemistry, 2020, 68: 1015-1021. |
| 40 | LÜ Y K, GU Y, XU J L, et al. Coupling metabolic addiction with negative autoregulation to improve strain stability and pathway yield[J]. Metabolic Engineering, 2020, 61: 79-88. |
| 41 | ZHOU S H, YUAN S F, NAIR P H, et al. Development of a growth coupled and multi-layered dynamic regulation network balancing malonyl-CoA node to enhance (2S)-naringenin biosynthesis in Escherichia coli [J]. Metabolic Engineering, 2021, 67: 41-52. |
| 42 | HOCHREIN L, MACHENS F, MESSERSCHMIDT K, et al. PhiReX: a programmable and red light-regulated protein expression switch for yeast[J]. Nucleic Acids Research, 2017, 45(15): 9193-9205. |
| 43 | MELENDEZ J, PATEL M, OAKES B L, et al. Real-time optogenetic control of intracellular protein concentration in microbial cell cultures[J]. Integrative Biology, 2014, 6(3): 366-372. |
| 44 | BINDER D, FROHWITTER J, MAHR R, et al. Light-controlled cell factories employing photocaged isopropyl-β-d-thiogalactopyranoside for light-mediated optimization of lac promoter-based gene expression and (+)-valencene biosynthesis in Corynebacterium glutamicum [J]. Applied and Environmental Microbiology. 2016, 82(20):6141-6149. |
| [1] | 赵静宇, 张健, 祁庆生, 王倩. 基于细菌双组分系统的生物传感器的研究进展[J]. 合成生物学, 2024, 5(1): 38-52. |
| [2] | 孙美莉, 王凯峰, 陆然, 纪晓俊. 解脂耶氏酵母底盘细胞的工程改造及应用[J]. 合成生物学, 2023, 4(4): 779-807. |
| [3] | 杨璐, 吴楠, 白茸茸, 董维亮, 周杰, 姜岷. 基因回路型全细胞微生物传感器的设计、优化与应用[J]. 合成生物学, 2022, 3(6): 1061-1080. |
| [4] | 张博, 马永硕, 尚轶, 黄三文. 植物合成生物学研究进展[J]. 合成生物学, 2020, 1(2): 121-140. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
