合成生物学 ›› 2020, Vol. 1 ›› Issue (4): 440-453.DOI: 10.12211/2096-8280.2020-029
动态调控策略在代谢工程中的应用研究进展
于政, 申晓林, 孙新晓, 王佳, 袁其朋
- 北京化工大学化工资源有效利用国家重点实验室,北京 100029
-
收稿日期:2020-03-19修回日期:2020-04-29出版日期:2020-08-31发布日期:2020-10-09 -
通讯作者:王佳,袁其朋 -
作者简介:于政(1998—),男,硕士研究生。研究方向为代谢工程及合成生物学。E-mail:2016018277@mail.buct.edu.cn
王佳(1989—),女,博士,副教授。研究方向为代谢工程及合成生物学。E-mail:wangjia@mail.buct.edu.cn
袁其朋(1969—),男,博士生导师,教授。研究方向为生物化工。E-mail:yuanqp@mail.buct.edu.cn -
基金资助:国家自然科学基金(21908003);国家重点研发计划(2018YFA0903000)
Application of dynamic regulation strategies in metabolic engineering
Yu Zheng, SHEN Xiaolin, Sun Xinxiao, Wang Jia, Yuan Qipeng
- State Key Laboratory of Chemical Resource Engineering,Beijing University of Chemical Technology,Beijing 100029,China
-
Received:2020-03-19Revised:2020-04-29Online:2020-08-31Published:2020-10-09 -
Contact:Wang Jia, Yuan Qipeng
摘要:
微生物细胞工厂作为一种可持续的生化反应器,被广泛应用于天然产物、药品、营养保健品等高附加值产物的生产中。为了使细胞工厂在生产过程中以最大的产量、产率和生产能力生产目标化合物,往往需要利用代谢工程方法对细胞工厂进行合理的改造和调控。以基因敲除和过表达为主要策略的静态调控不可避免地带来了细胞代谢流与能量流失衡、生长阻滞和毒性中间体积累等问题,限制了细胞工厂的生产能力、碳收率和产物产量。为了解决这一问题,构建调控元件并设计基因线路以精确调节物质流及能量流的动态调控策略被普遍应用于代谢工程领域,成为调控微生物细胞工厂的常用方法之一。本文依据不同动态调控策略的特点,将动态调控策略分为代谢物响应、群体感应响应、环境响应和蛋白质水平调控四种类型,重点介绍了各种调控元件的构建方法及其在代谢工程中的应用,分析了不同调控策略在工业化应用中面临的挑战。同时,指出了高通量筛选和蛋白质工程方法、计算机模拟和数学模型分析、耦合基因控制元件等方面的策略在解决动态调控工具响应阈值窄、调控范围有限等问题中的应用潜力。
中图分类号:
引用本文
于政, 申晓林, 孙新晓, 王佳, 袁其朋. 动态调控策略在代谢工程中的应用研究进展[J]. 合成生物学, 2020, 1(4): 440-453.
Yu Zheng, SHEN Xiaolin, Sun Xinxiao, Wang Jia, Yuan Qipeng. Application of dynamic regulation strategies in metabolic engineering[J]. Synthetic Biology Journal, 2020, 1(4): 440-453.
| 调控类型 | 输入信号 | 调控元件 | 菌株 | 产物 | 调控效果 |
|---|---|---|---|---|---|
| 转录因子 | 乙酰磷酸 | NRI | 大肠杆菌 | 番茄红素 | 50%[ |
| 丙二酰辅酶A | FapR | 大肠杆菌 | 脂肪酸 | 2.1倍[ | |
| 丙二酰辅酶A | FapR | 大肠杆菌 | 脂肪酸 | 34%[ | |
| 衣康酸 | ItcR | 大肠杆菌 | 衣康酸 | 0.78mmol/L[ | |
| 阿魏酸、香草醛 | HucR | 大肠杆菌 | 香草醛 | 11mmol/L[ | |
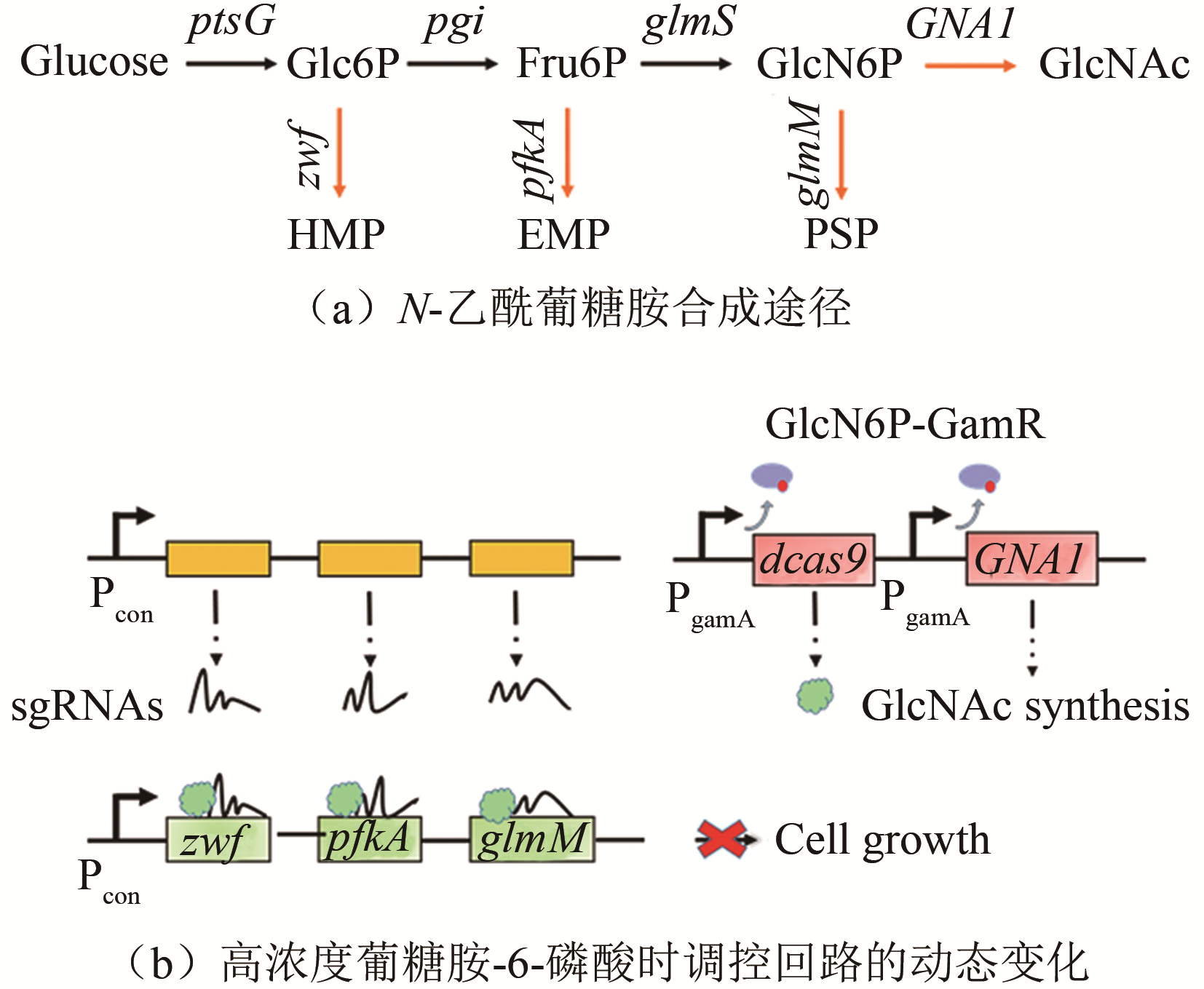
| 葡糖胺-6-磷酸 | GamR | 枯草芽孢杆菌 | N-乙酰葡糖胺 | 131.6 g/L[ | |
| 黏糠酸 | CatR | 大肠杆菌 | 黏糠酸 | 16.3倍[ | |
| 核糖开关 | 茶碱 | 茶碱核糖开关 | 大肠杆菌 | — | 3倍[ |
| 硫胺素焦磷酸 | 硫胺素焦磷酸核糖开关 | 米曲霉 | — | 4.7倍[ | |
| 赖氨酸 | 赖氨酸核糖开关 | 谷氨酸棒状杆菌 | 赖氨酸 | 63%[ | |
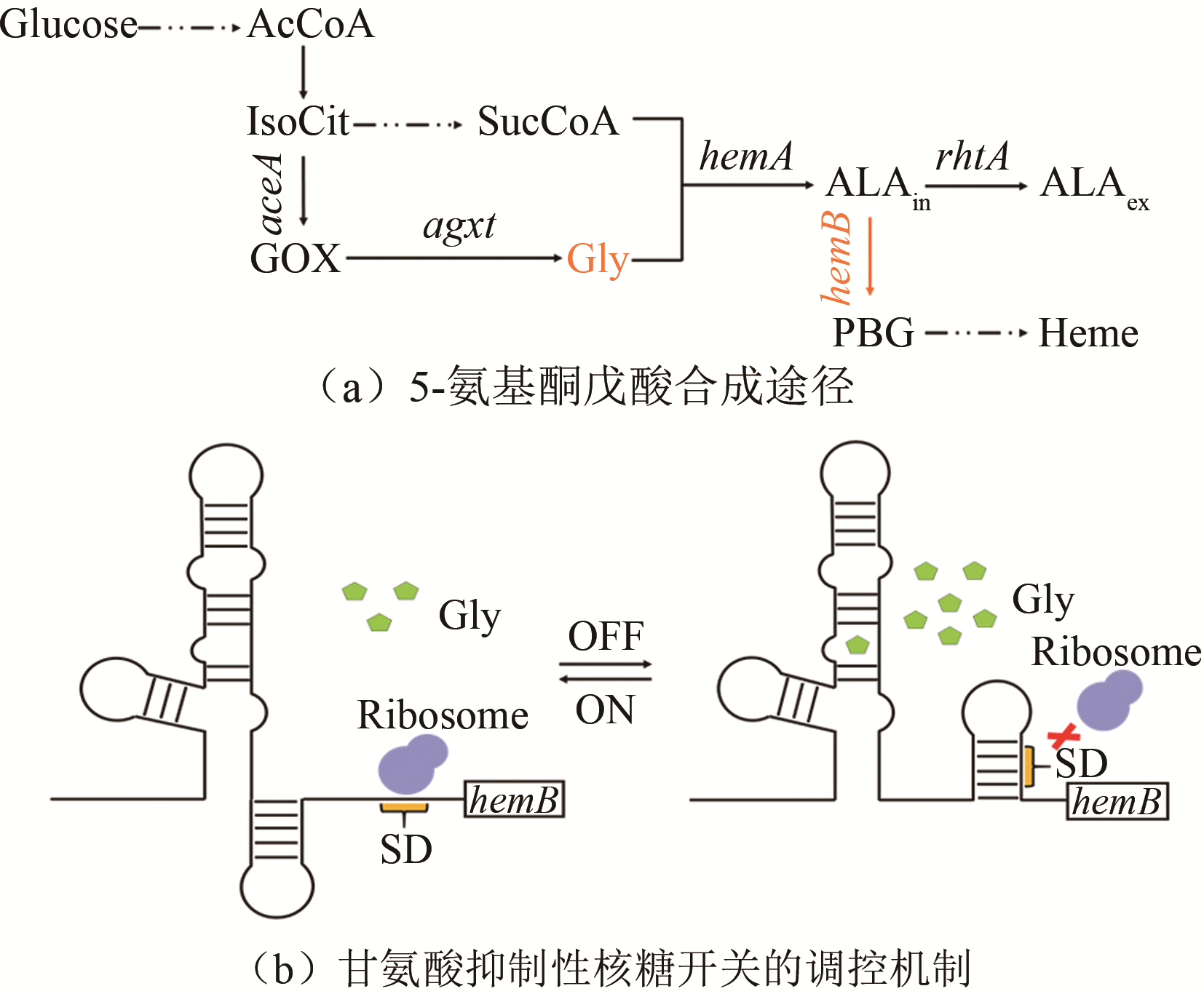
| 甘氨酸 | 甘氨酸核糖开关 | 大肠杆菌 | 5-氨基酮戊酸 | 11%[ | |
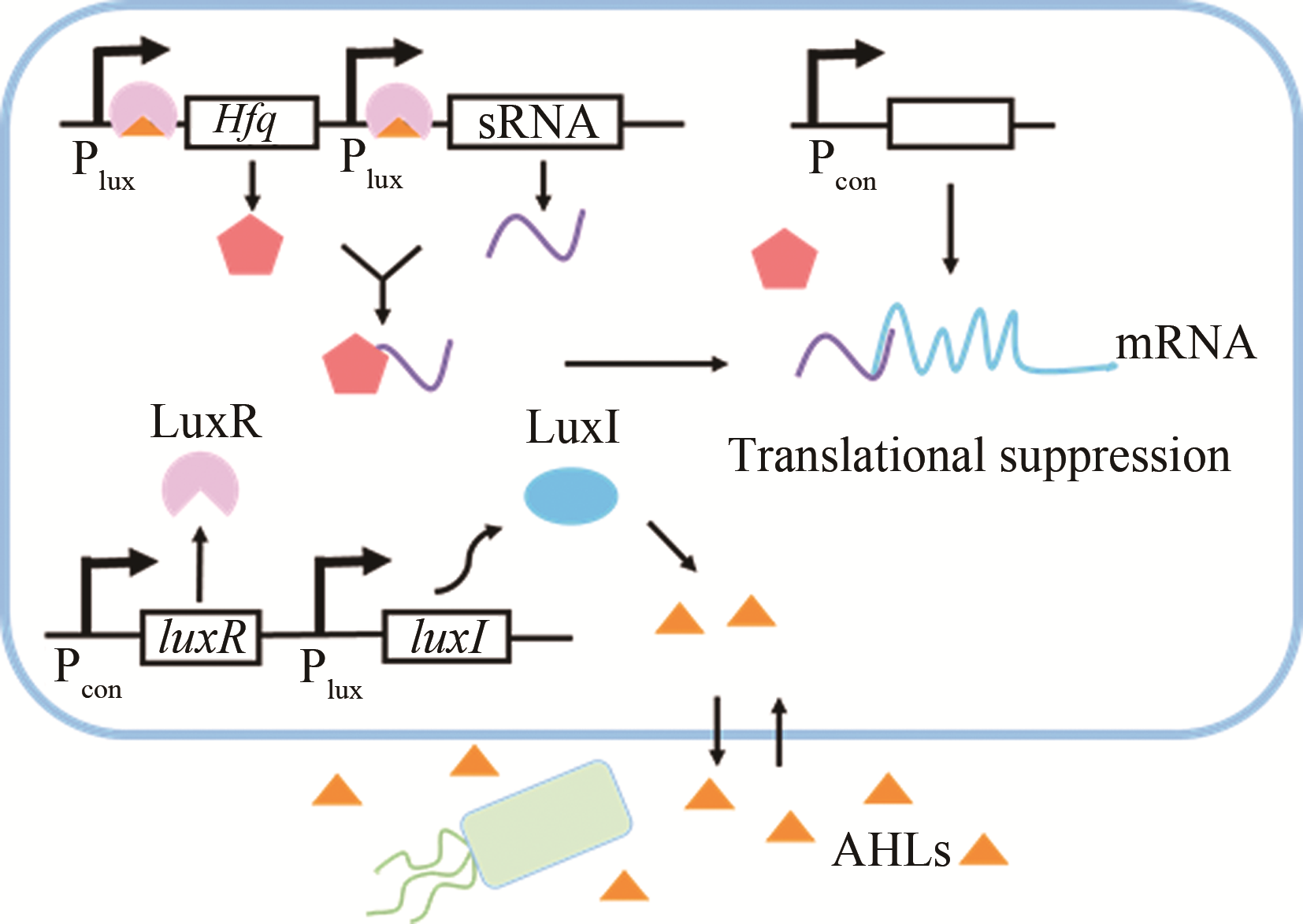
| 群体感应 | AHLs | luxI/luxR | 大肠杆菌 | 红没药烯 | 44%[ |
| AHLs | luxI/luxR | 大肠杆菌 | — | 6%~76%[ | |
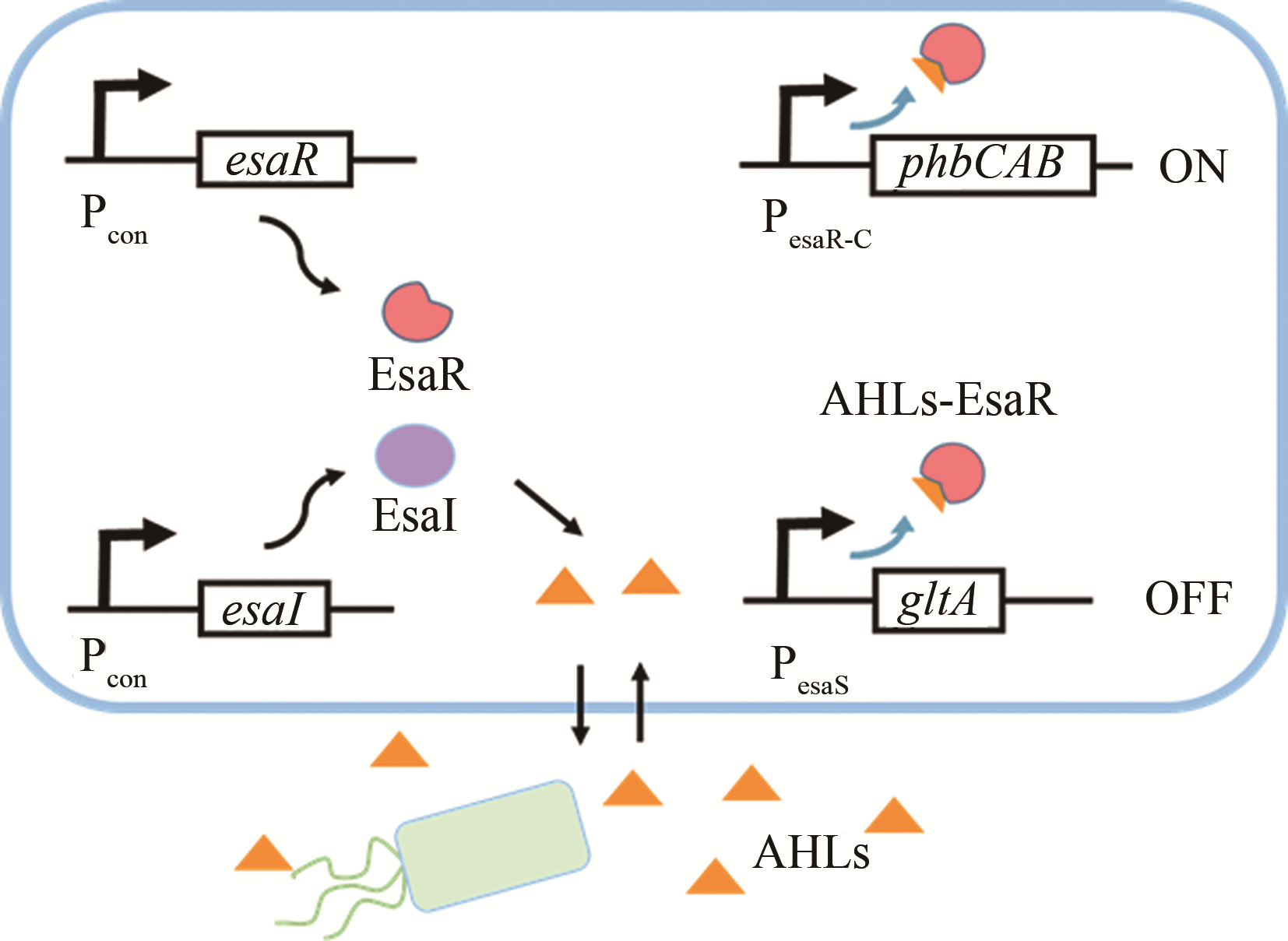
| AHLs | esaI/esaR | 大肠杆菌 | 4-羟基苯乙酸 | 46.4%[ | |
| AHLs | esaI/esaR | 大肠杆菌 | 聚-β-羟丁酸 | 6倍[ | |
| AHLs | esaI/esaR | 大肠杆菌 | 葡萄糖二酸 | 2g/L[ | |
| AHLs | lux、esa | 大肠杆菌 | 水杨酸、柚皮素 | 1.8、6倍[ | |
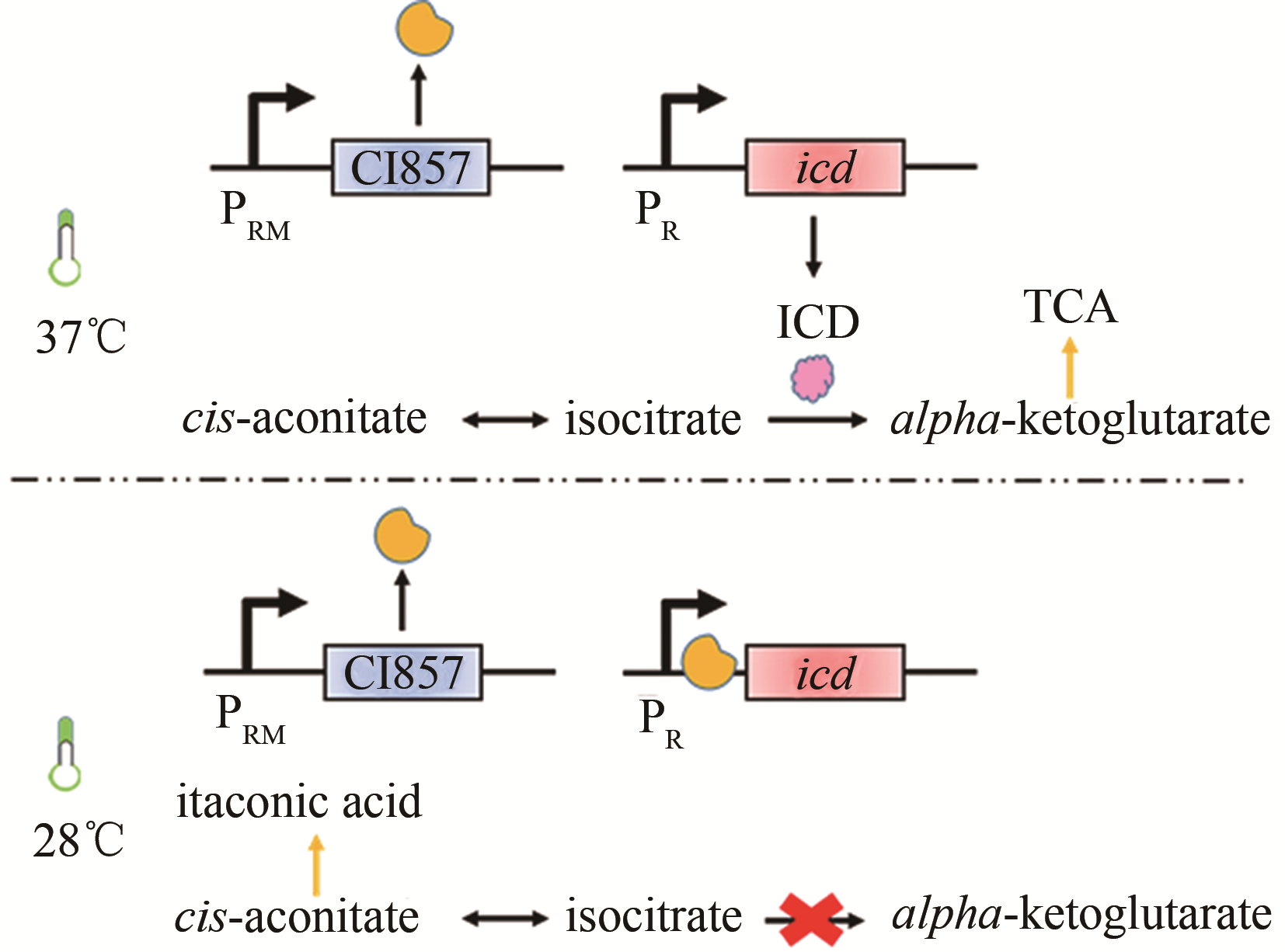
| 发酵条件 | 温度 | CI857 | 大肠杆菌 | 衣康酸 | 47g/L[ |
| 温度 | Gal4M9 | 酵母 | 番茄红素 | 1.12g/L[ | |
| 溶氧 | Pnar | 大肠杆菌 | D-乳酸 | 113.12g/L[ | |
| pH | Pgas | 黑曲霉 | 衣康酸 | 4.92g/L[ | |
| pH | CadCΔ | 大肠杆菌 | 乙二醇 | 170%[ | |
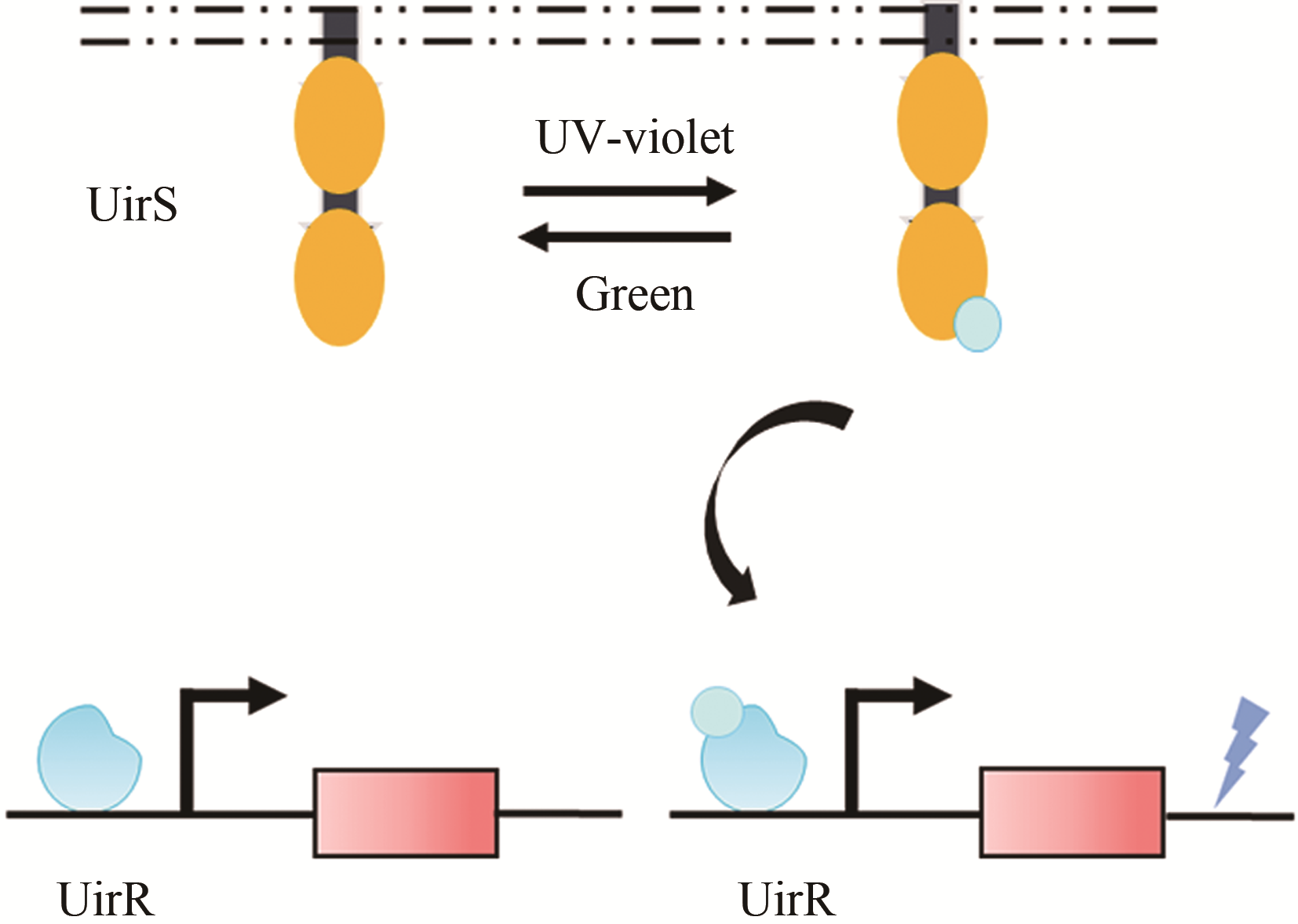
| 光 | UirS/UirR | 大肠杆菌 | — | 6.24倍[ | |
| 光 | Magnets | 大肠杆菌 | — | 300倍[ | |
| 光 | OptoEXP | 酵母 | 异丁醇 | 4倍[ | |
| 葡萄糖 | PHXT1 | 酵母 | 3-羟基丙酸 | 10倍[ | |
| 蛋白水平 | 赖氨酸 | 高丝氨酸脱氢酶 | 大肠杆菌 | 赖氨酸 | —[ |
| — | 振荡器 | 大肠杆菌 | D-木糖酸 | 199.44g/L[ | |
| — | SsrA | 大肠杆菌 | 肌醇 | 2倍[ |
表1 动态调控元件在代谢工程中的应用
Tab. 1 Applications of dynamic regulation elements in metabolic engineering
| 调控类型 | 输入信号 | 调控元件 | 菌株 | 产物 | 调控效果 |
|---|---|---|---|---|---|
| 转录因子 | 乙酰磷酸 | NRI | 大肠杆菌 | 番茄红素 | 50%[ |
| 丙二酰辅酶A | FapR | 大肠杆菌 | 脂肪酸 | 2.1倍[ | |
| 丙二酰辅酶A | FapR | 大肠杆菌 | 脂肪酸 | 34%[ | |
| 衣康酸 | ItcR | 大肠杆菌 | 衣康酸 | 0.78mmol/L[ | |
| 阿魏酸、香草醛 | HucR | 大肠杆菌 | 香草醛 | 11mmol/L[ | |
| 葡糖胺-6-磷酸 | GamR | 枯草芽孢杆菌 | N-乙酰葡糖胺 | 131.6 g/L[ | |
| 黏糠酸 | CatR | 大肠杆菌 | 黏糠酸 | 16.3倍[ | |
| 核糖开关 | 茶碱 | 茶碱核糖开关 | 大肠杆菌 | — | 3倍[ |
| 硫胺素焦磷酸 | 硫胺素焦磷酸核糖开关 | 米曲霉 | — | 4.7倍[ | |
| 赖氨酸 | 赖氨酸核糖开关 | 谷氨酸棒状杆菌 | 赖氨酸 | 63%[ | |
| 甘氨酸 | 甘氨酸核糖开关 | 大肠杆菌 | 5-氨基酮戊酸 | 11%[ | |
| 群体感应 | AHLs | luxI/luxR | 大肠杆菌 | 红没药烯 | 44%[ |
| AHLs | luxI/luxR | 大肠杆菌 | — | 6%~76%[ | |
| AHLs | esaI/esaR | 大肠杆菌 | 4-羟基苯乙酸 | 46.4%[ | |
| AHLs | esaI/esaR | 大肠杆菌 | 聚-β-羟丁酸 | 6倍[ | |
| AHLs | esaI/esaR | 大肠杆菌 | 葡萄糖二酸 | 2g/L[ | |
| AHLs | lux、esa | 大肠杆菌 | 水杨酸、柚皮素 | 1.8、6倍[ | |
| 发酵条件 | 温度 | CI857 | 大肠杆菌 | 衣康酸 | 47g/L[ |
| 温度 | Gal4M9 | 酵母 | 番茄红素 | 1.12g/L[ | |
| 溶氧 | Pnar | 大肠杆菌 | D-乳酸 | 113.12g/L[ | |
| pH | Pgas | 黑曲霉 | 衣康酸 | 4.92g/L[ | |
| pH | CadCΔ | 大肠杆菌 | 乙二醇 | 170%[ | |
| 光 | UirS/UirR | 大肠杆菌 | — | 6.24倍[ | |
| 光 | Magnets | 大肠杆菌 | — | 300倍[ | |
| 光 | OptoEXP | 酵母 | 异丁醇 | 4倍[ | |
| 葡萄糖 | PHXT1 | 酵母 | 3-羟基丙酸 | 10倍[ | |
| 蛋白水平 | 赖氨酸 | 高丝氨酸脱氢酶 | 大肠杆菌 | 赖氨酸 | —[ |
| — | 振荡器 | 大肠杆菌 | D-木糖酸 | 199.44g/L[ | |
| — | SsrA | 大肠杆菌 | 肌醇 | 2倍[ |
| 1 | SUN X, SHEN X, JAIN R, et al. Synthesis of chemicals by metabolic engineering of microbes[J]. Chemical Society Reviews, 2015, 44(11): 3760-3785. |
| 2 | SHEN X, WANG J, WANG J, et al. High-level de novo biosynthesis of arbutin in engineered Escherichia coli [J]. Metabolic Engineering, 2017, 42: 52-58. |
| 3 | WANG J, SHEN X, YUAN Q, et al. Microbial synthesis of pyrogallol using genetically engineered Escherichia coli [J]. Metabolic Engineering, 2018, 45: 134-141. |
| 4 | ATSUMI S, HANAI T, LIAO J C. Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels[J]. Nature, 2008, 451(7174): 86-89. |
| 5 | CHEN X, GAO C, GUO L, et al. DCEO biotechnology: tools to design, construct, evaluate, and optimize the metabolic pathway for biosynthesis of chemicals[J]. Chemical Reviews, 2018, 118(1): 4-72. |
| 6 | SHEN X, WANG J, LI C, et al. Dynamic gene expression engineering as a tool in pathway engineering[J]. Current Opinion in Biotechnology, 2019, 59: 122-129. |
| 7 | TAN S Z, PRATHER K L. Dynamic pathway regulation: recent advances and methods of construction[J]. Current Opinion in Chemical Biology, 2017, 41: 28-35. |
| 8 | XU P, GU Q, WANG W, et al. Modular optimization of multi-gene pathways for fatty acids production in E. coli [J]. Nature Communications, 2013, 4: 1409. |
| 9 | FARMER W R, LIAO J C. Improving lycopene production in Escherichia coli by engineering metabolic control[J]. Nature Biotechnology, 2000, 18(5): 533-537. |
| 10 | DOONG S J, GUPTA A, PRATHER K L J. Layered dynamic regulation for improving metabolic pathway productivity in Escherichia coli [J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(12): 2964-2969. |
| 11 | WU J, LIU Y, ZHAO S, et al. Application of dynamic regulation to increase L-phenylalanine production in Escherichia coli [J]. Journal of Microbiology and Biotechnology, 2019, 29(6): 923-932. |
| 12 | SPITZ F, FURLONG E E. Transcription factors: from enhancer binding to developmental control[J]. Nature Reviews Genetics, 2012, 13(9): 613-626. |
| 13 | ZHANG F, KEASLING J. Biosensors and their applications in microbial metabolic engineering[J]. Trends in Microbiology, 2011, 19(7): 323-329. |
| 14 | XU P. Production of chemicals using dynamic control of metabolic fluxes[J]. Current Opinion in Biotechnology, 2018, 53: 12-19. |
| 15 | WU J, YU O, DU G, et al. Fine-tuning of the fatty acid pathway by synthetic antisense RNA for enhanced (2S)-naringenin production from L-tyrosine in Escherichia coli [J]. Applied and Environmental Microbiology, 2014, 80(23): 7283-7292. |
| 16 | CHEN Z, HUANG J, WU Y, et al. Metabolic engineering of Corynebacterium glutamicum for the production of 3-hydroxypropionic acid from glucose and xylose[J]. Metabolic Engineering, 2017, 39: 151-158. |
| 17 | SHEN X, MAHAJANI M, WANG J, et al. Elevating 4-hydroxycoumarin production through alleviating thioesterase-mediated salicoyl-CoA degradation[J]. Metabolic Engineering, 2017, 42: 59-65. |
| 18 | XU P, LI L, ZHANG F, et al. Improving fatty acids production by engineering dynamic pathway regulation and metabolic control[J]. Proceedings of the National Academy of Sciences of the United States of America, 2014, 111(31): 11299-11304. |
| 19 | LIU D, XIAO Y, EVANS B S, et al. Negative feedback regulation of fatty acid production based on a malonyl-CoA sensor-actuator[J]. ACS Synthetic Biology, 2015, 4(2): 132-140. |
| 20 | HANKO E K R, MINTON N P, MALYS N. A transcription factor-based biosensor for detection of itaconic acid[J]. ACS Synthetic Biology, 2018, 7(5): 1436-1446. |
| 21 | KOCH M, PANDI A, BORKOWSKI O, et al. Custom-made transcriptional biosensors for metabolic engineering[J]. Current Opinion in Biotechnology, 2019, 59: 78-84. |
| 22 | LIANG C, ZHANG X, WU J, et al. Dynamic control of toxic natural product biosynthesis by an artificial regulatory circuit[J]. Metabolic Engineering, 2020, 57: 239-246. |
| 23 | DABIRIAN Y, LI X, CHEN Y, et al. Expanding the dynamic range of a transcription factor-based biosensor in Saccharomyces cerevisiae [J]. ACS Synthetic Biology, 2019, 8(9): 1968-1975. |
| 24 | CHEN Y, HO J M L, SHIS D L, et al. Tuning the dynamic range of bacterial promoters regulated by ligand-inducible transcription factors[J]. Nature Communications, 2018, 9(1): 64. |
| 25 | MANNAN A A, LIU D, ZHANG F, et al. Fundamental design principles for transcription-factor-based metabolite biosensors[J]. ACS Synthetic Biology, 2017, 6(10): 1851-1859. |
| 26 | TIAN J, YANG G, GU Y, et al. Developing an endogenous quorum-sensing based CRISPRi circuit for autonomous and tunable dynamic regulation of multiple targets in industrial Streptomyces [J]. Nucleic Acids Research, 2020. DOI: 10.1093/nar/gkaa602 . |
| 27 | CHO S, SHIN J, CHO B K. Applications of CRISPR/Cas system to bacterial metabolic engineering[J]. International Journal of Molecular Sciences, 2018, 19(4): 1089. |
| 28 | WU Y, CHEN T, LIU Y, et al. Design of a programmable biosensor-CRISPRi genetic circuits for dynamic and autonomous dual-control of metabolic flux in Bacillus subtilis [J]. Nucleic Acids Research, 2020, 48(2): 996-1009. |
| 29 | YANG Y, LIN Y, WANG J, et al. Sensor-regulator and RNAi based bifunctional dynamic control network for engineered microbial synthesis[J]. Nature Communications, 2018, 9(1): 3043. |
| 30 | GALIZI R, JARAMILLO A. Engineering CRISPR guide RNA riboswitches for in vivo applications[J]. Current Opinion in Biotechnology, 2019, 55: 103-113. |
| 31 | LIU D, EVANS T, ZHANG F. Applications and advances of metabolite biosensors for metabolic engineering[J]. Metabolic Engineering, 2015, 31: 35-43. |
| 32 | JANG S, JANG S, XIU Y, et al. Development of artificial riboswitches for monitoring of naringenin in vivo [J]. ACS Synthetic Biology, 2017, 6(11): 2077-2085. |
| 33 | BREAKER R R. Riboswitches and translation control[J]. Cold Spring Harbor Perspectives in Biology, 2018, 10(11): a032797. |
| 34 | LOTZ T S, SUESS B. Small-molecule-binding riboswitches[J]. Microbiology Spectrum, 2018, 6(4): 75-88. |
| 35 | WACHSMUTH M, FINDEISS S, WEISSHEIMER N, et al. De novo design of a synthetic riboswitch that regulates transcription termination[J]. Nucleic Acids Research, 2013, 41(4): 2541-2551. |
| 36 | MASUDA S, LZAWA S. Applied RNA bioscience[M]. Singapore: Springer, 2018: 33-46. |
| 37 | ZHOU L B, ZENG A P. Exploring lysine riboswitch for metabolic flux control and improvement of L-lysine synthesis in Corynebacterium glutamicum [J]. ACS Synthetic Biology, 2015, 4(6): 729-734. |
| 38 | ZHOU L, REN J, LI Z, et al. Characterization and engineering of a Clostridium glycine riboswitch and its use to control a novel metabolic pathway for 5-aminolevulinic acid production in Escherichia coli [J]. ACS Synthetic Biology, 2019, 8(10): 2327-2335. |
| 39 | PANG Q, HAN H, LIU X, et al. In vivo evolutionary engineering of riboswitch with high-threshold for N-acetylneuraminic acid production[J]. Metabolic Engineering, 2020, 59: 36-43. |
| 40 | HAN L, HAN D, LI L, et al. Discovery and identification of medium‐chain fatty acid responsive promoters in Saccharomyces cerevisiae [J]. Engineering in Life Sciences, 2020, 5: 186-196. |
| 41 | MAURY J, KANNAN S, JENSEN N B, et al. Glucose-dependent promoters for dynamic regulation of metabolic pathways[J]. Frontiers in Bioengineering and Biotechnology, 2018, 6: 63. |
| 42 | ABISADO R G, BENOMAR S, KLAUS J R, et al. Bacterial quorum sensing and microbial community interactions[J]. mBio, 2018, 9(3): e02331-17. |
| 43 | DANG H T, KOMATSU S, MASUDA H, et al. Characterization of LuxI and LuxR protein homologs of N-Acylhomoserine lactone-dependent quorum sensing system in Pseudoalteromonas sp. 520P1[J]. Marine Biotechnology, 2017, 19(1): 1-10. |
| 44 | WHITELEY M, DIGGLE S P, GREENBERG E P. Progress in and promise of bacterial quorum sensing research[J]. Nature, 2017, 551(7680): 313-320. |
| 45 | KIM E M, WOO H M, TIAN T, et al. Autonomous control of metabolic state by a quorum sensing (QS)-mediated regulator for bisabolene production in engineered E. coli [J]. Metabolic Engineering, 2017, 44: 325-336. |
| 46 | BAO S H, LI W Y, LIU C J, et al. Quorum-sensing based small RNA regulation for dynamic and tuneable gene expression[J]. Biotechnology Letters, 2019, 41(10): 1147-1154. |
| 47 | KIMATU J N. Advances in plant pathology[M]. London: IntechOpen, 2018: 68-89. |
| 48 | PAPENFORT K, BASSLER B L. Quorum sensing signal-response systems in Gram-negative bacteria[J]. Nature Reviews Microbiology, 2016, 14(9): 576-588. |
| 49 | SHEN Y P, FONG L S, YAN Z B, et al. Combining directed evolution of pathway enzymes and dynamic pathway regulation using a quorum-sensing circuit to improve the production of 4-hydroxyphenylacetic acid in Escherichia coli [J]. Biotechnology for Biofuels, 2019, 12: 94. |
| 50 | GU F, JIANG W, MU Y, et al. Quorum sensing-based dual-function switch and its application in solving two key metabolic engineering problems[J]. ACS Synthetic Biology, 2020, 9(2): 209-217. |
| 51 | GUPTA A, REIZMAN I M, REISCH C R, et al. Dynamic regulation of metabolic flux in engineered bacteria using a pathway-independent quorum-sensing circuit[J]. Nature Biotechnology, 2017, 35(3): 273-279. |
| 52 | DOONG S J, GUPTA A, PRATHER K L J. Layered dynamic regulation for improving metabolic pathway productivity in Escherichia coli [J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(12): 2964-2969. |
| 53 | DINH C V, PRATHER K L J. Development of an autonomous and bifunctional quorum-sensing circuit for metabolic flux control in engineered Escherichia coli [J]. Proceedings of the National Academy of Sciences of the United States of America, 2019, 116(51): 25562-25568. |
| 54 | ZENG W, DU P, LOU Q, et al. Rational design of an ultrasensitive quorum-sensing switch[J]. ACS Synthetic Biology, 2017, 6(8): 1445-1452. |
| 55 | DAER R, BARRETT C M, MELENDEZ E L, et al. Characterization of diverse homoserine lactone synthases in Escherichia coli [J]. Plos One, 2018, 13(8): e0202294. |
| 56 | TEKEL S J, SMITH C L, LOPEZ B, et al. Engineered orthogonal quorum sensing systems for synthetic gene regulation in Escherichia coli [J]. Frontiers in Bioengineering and Biotechnology, 2019, 7: 80. |
| 57 | SCOTT S R, HASTY J. Quorum sensing communication modules for microbial consortia[J]. ACS Synthetic Biology, 2016, 5(9): 969-977. |
| 58 | HARDER B J, BETTENBROCK K, KLAMT S. Temperature-dependent dynamic control of the TCA cycle increases volumetric productivity of itaconic acid production by Escherichia coli [J]. Biotechnology and Bioengineering, 2018, 115(1): 156-164. |
| 59 | ZHOU P, XIE W, YAO Z, et al. Development of a temperature-responsive yeast cell factory using engineered Gal4 as a protein switch[J]. Biotechnology and Bioengineering, 2018, 115(5): 1321-1330. |
| 60 | HWANG H J, KIM J W, JU S Y, et al. Application of an oxygen-inducible nar promoter system in metabolic engineering for production of biochemicals in Escherichia coli [J]. Biotechnology and Bioengineering, 2017, 114(2): 468-473. |
| 61 | YIN X, SHIN H D, LI J, et al. Pgas, a low-pH-induced promoter, as a tool for dynamic control of gene expression for metabolic engineering of Aspergillus niger [J]. Applied and Environmental Microbiology, 2017, 83(6): e03222-16. |
| 62 | BANARES A B, VALDEHUESA K N G, RAMOS K R M, et al. A pH-responsive genetic sensor for the dynamic regulation of D-xylonic acid accumulation in Escherichia coli [J]. Applied Microbiology and Biotechnology, 2020, 104(5): 2097-2108. |
| 63 | LIU Z, ZHANG J, JIN J, et al. Programming bacteria with light-sensors and applications in synthetic biology[J]. Frontiers in Microbiology, 2018, 9: 2692. |
| 64 | TANDAR S T, SENOO S, TOYA Y, et al. Optogenetic switch for controlling the central metabolic flux of Escherichia coli [J]. Metabolic Engineering, 2019, 55: 68-75. |
| 65 | RAMAKRISHNAN P, TABOR J J. Repurposing Synechocystis PCC6803 UirS-UirR as a UV-violet/Green photoreversible transcriptional regulatory tool in E. coli [J]. ACS Synthetic Biology, 2016, 5(7): 733-740. |
| 66 | BAUMSCHLAGER A, AOKI S K, KHAMMASH M. Dynamic blue light-inducible T7 RNA polymerases (Opto-T7RNAPs) for precise spatiotemporal gene expression control[J]. ACS Synthetic Biology, 2017, 6(11): 2157-2167. |
| 67 | ZHAO E M, ZHANG Y, MEHL J, et al. Optogenetic regulation of engineered cellular metabolism for microbial chemical production[J]. Nature, 2018, 555(7698): 683-687. |
| 68 | SHEETS M B, WONG W W, DUNLOP M J. Light-inducible recombinases for bacterial optogenetics[J]. ACS Synthetic Biology, 2020, 9(2): 227-235. |
| 69 | BOTHFELD W, KAPOV G, TYO K E J. A glucose-sensing toggle switch for autonomous, high productivity genetic control[J]. ACS Synthetic Biology, 2017, 6(7): 1296-1304. |
| 70 | DAVID F, NIELSEN J, SIEWERS V. Flux control at the malonyl-CoA node through hierarchical dynamic pathway regulation in Saccharomyces cerevisiae [J]. ACS Synthetic Biology, 2016, 5(3): 224-233. |
| 71 | MOTLAGH H N, WRABL J O, LI J, et al. The ensemble nature of allostery[J]. Nature, 2014, 508(7496): 331-339. |
| 72 | DOKHOLYAN N V. Controlling allosteric networks in proteins[J]. Chemical Reviews, 2016, 116(11): 6463-6487. |
| 73 | CHEN Z, RAPPERT S, ZENG A P. Rational design of allosteric regulation of homoserine dehydrogenase by a nonnatural inhibitor L-lysine[J]. ACS Synthetic Biology, 2015, 4(2): 126-131. |
| 74 | GAO C, HOU J, XU P, et al. Programmable biomolecular switches for rewiring flux in Escherichia coli [J]. Nature Communications, 2019, 10(1): 3751. |
| 75 | BROCKMAN I M, PRATHER K L J. Dynamic knockdown of E. coli central metabolism for redirecting fluxes of primary metabolites[J]. Metabolic Engineering, 2015, 28: 104-113. |
| 76 | HE F, STUMPF M P H. Quantifying dynamic regulation in metabolic pathways with nonparametric flux inference[J]. Biophysical Journal, 2019, 116(10): 2035-2046. |
| 77 | JABARIVELISDEH B, WALDHERR S. Optimization of bioprocess productivity based on metabolic-genetic network models with bilevel dynamic programming[J]. Biotechnology and Bioengineering, 2018, 115(7): 1829-1841. |
| [1] | 郭姝媛, 张倩楠, 姑丽克孜·买买提热夏提, 杨一群, 于涛. 液体生物燃料合成与炼制的研究进展[J]. 合成生物学, 2025, 6(1): 18-44. |
| [2] | 邵明威, 孙思勉, 杨时茂, 陈国强. 基于极端微生物的生物制造[J]. 合成生物学, 2024, 5(6): 1419-1436. |
| [3] | 赵亮, 李振帅, 付丽平, 吕明, 王士安, 张全, 刘立成, 李福利, 刘自勇. 生物转化一碳化合物原料产油脂与单细胞蛋白研究进展[J]. 合成生物学, 2024, 5(6): 1300-1318. |
| [4] | 竺方欢, 岑雪聪, 陈振. 微生物合成二元醇研究进展[J]. 合成生物学, 2024, 5(6): 1367-1385. |
| [5] | 郑皓天, 李朝风, 刘良叙, 王嘉伟, 李恒润, 倪俊. 负碳人工光合群落的设计、优化与应用[J]. 合成生物学, 2024, 5(5): 1189-1210. |
| [6] | 禹伟, 高教琪, 周雍进. 一碳生物转化合成有机酸的研究进展[J]. 合成生物学, 2024, 5(5): 1169-1188. |
| [7] | 陈锡玮, 张华然, 邹懿. 真菌源非核糖体肽类药物生物合成及代谢工程[J]. 合成生物学, 2024, 5(3): 571-592. |
| [8] | 谢皇, 郑义蕾, 苏依婷, 阮静怡, 李永泉. 放线菌聚酮类化合物生物合成体系重构研究进展[J]. 合成生物学, 2024, 5(3): 612-630. |
| [9] | 惠真, 唐啸宇. CRISPR/Cas9编辑系统在微生物天然产物研究中的应用[J]. 合成生物学, 2024, 5(3): 658-671. |
| [10] | 赵静宇, 张健, 祁庆生, 王倩. 基于细菌双组分系统的生物传感器的研究进展[J]. 合成生物学, 2024, 5(1): 38-52. |
| [11] | 孙绘梨, 崔金玉, 栾国栋, 吕雪峰. 面向高效光驱固碳产醇的蓝细菌合成生物技术研究进展[J]. 合成生物学, 2023, 4(6): 1161-1177. |
| [12] | 晏雄鹰, 王振, 娄吉芸, 张皓瑜, 黄星宇, 王霞, 杨世辉. 生物燃料高效生产微生物细胞工厂构建研究进展[J]. 合成生物学, 2023, 4(6): 1082-1121. |
| [13] | 程真真, 张健, 高聪, 刘立明, 陈修来. 代谢工程改造微生物利用甲酸研究进展[J]. 合成生物学, 2023, 4(4): 756-778. |
| [14] | 刘家宇, 杨智晗, 杨蕾, 朱丽英, 朱政明, 江凌. 合成生物技术驱动酪丁酸梭菌细胞工厂开发的研究进展[J]. 合成生物学, 2022, 3(6): 1174-1200. |
| [15] | 郭姝媛, 吴良焕, 刘香健, 王博, 于涛. 微生物中一碳代谢网络构建的进展与挑战[J]. 合成生物学, 2022, 3(1): 116-137. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||