合成生物学 ›› 2024, Vol. 5 ›› Issue (6): 1367-1385.DOI: 10.12211/2096-8280.2024-014
微生物合成二元醇研究进展
竺方欢1, 岑雪聪1, 陈振1,2
- 1.清华大学化学工程系,工业生物催化教育部重点实验室,北京 100084
2.清华大学合成与系统生物学中心,北京 100084
-
收稿日期:2024-02-04修回日期:2024-05-08出版日期:2024-12-31发布日期:2025-01-10 -
通讯作者:陈振 -
作者简介:竺方欢 (1996—),女,博士研究生。研究方向为二元醇的绿色生物制造。E-mail:zfh21@mails.tsinghua.edu.cn岑雪聪 (1996—),女,博士研究生。研究方向为二元醇的绿色生物制造。 E-mail:cxc18@mails.tsinghua.edu.cn陈振 (1983—),男,副教授,博士生导师。研究方向为材料、化学品及生物医药的绿色生物制造。E-mail:zhenchen2013@tsinghua.edu.cn -
基金资助:国家重点研发计划(2021YFC2100900);国家自然科学基金(22078172)
Research progress of diols production by microbes
ZHU Fanghuan1, CEN Xuecong1, CHEN Zhen1,2
- 1.Key Laboratory of Industrial Biocatalysis (Ministry of Education),Department of Chemical Engineering,Tsinghua University,Beijing 100084,China
2.Center for Synthetic and Systems Biology,Tsinghua University,Beijing 100084,China
-
Received:2024-02-04Revised:2024-05-08Online:2024-12-31Published:2025-01-10 -
Contact:CHEN Zhen
摘要:
二元醇是一类重要的大宗化学品,在高分子材料、化妆品、燃料、食品和制药行业有着广泛应用。开发可利用生物质及碳一原料等可再生原料生产二元醇的生物合成路线对于降低化石资源依赖、减少二氧化碳的排放具有重要意义,近年来受到了国内外广泛关注。虽然通过生物法生产1,3-丙二醇、1,3-丁二醇和1,4-丁二醇已实现商业化,但大多数其他二元醇的高效生物合成仍面临挑战,主要原因包括缺乏有效的天然生物合成途径、基因工程菌的产率低等。本综述全面探讨了微生物合成二元醇的最新研究进展,特别是在开发新代谢途径和代谢工程策略方面,以实现C2至C5二元醇的高效生物合成。例如通过对非天然合成途径的设计和构建以实现系列非天然二元醇的生物合成,以及利用非传统的碳源(如木质纤维素等)通过特定的代谢途径和优化策略合成二元醇,为生物合成领域开辟新的道路。此外,本文还讨论了这些生物合成过程向工业应用转化的主要挑战和未来的发展前景,包括廉价和可持续原料的获取、大规模放大过程的复杂性、满足下游特定需求的后提取工艺开发等。
中图分类号:
引用本文
竺方欢, 岑雪聪, 陈振. 微生物合成二元醇研究进展[J]. 合成生物学, 2024, 5(6): 1367-1385.
ZHU Fanghuan, CEN Xuecong, CHEN Zhen. Research progress of diols production by microbes[J]. Synthetic Biology Journal, 2024, 5(6): 1367-1385.
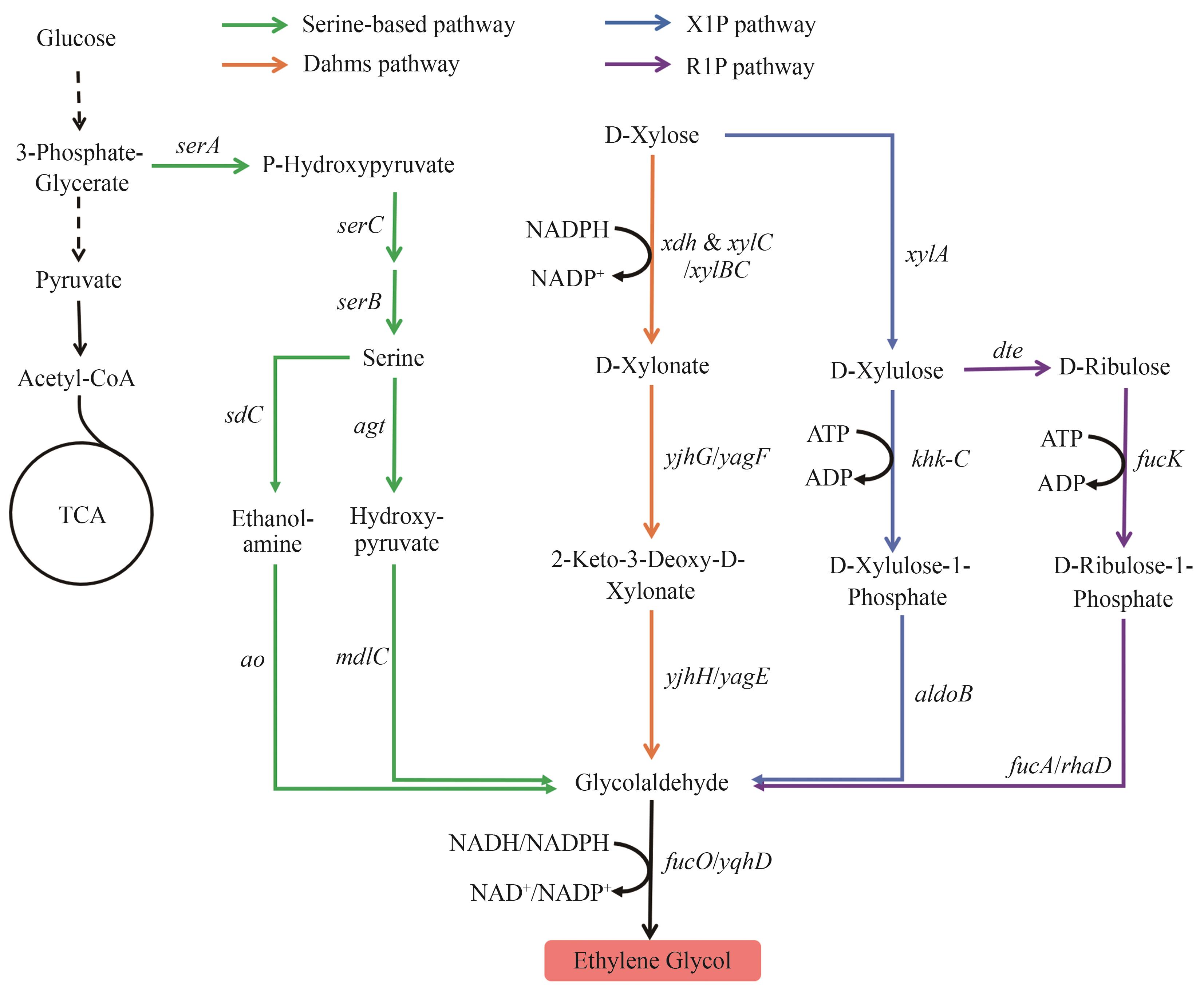
图1 乙二醇的生物合成路径X1P—木糖醛酸-1-磷酸; R1P—核糖醛酸-1-磷酸图中基因所编码的酶:serA—磷酸甘油酸脱氢酶;serC—磷酸丝氨酸氨基转移酶;serB—磷酸丝氨酸磷酸酶;sdC—丝氨酸脱羧酶;ao—胺氧化酶;agt—丝氨酸-乙醛酸氨基转移酶;mdlC—苯甲酰甲酸脱羧酶;fucO/yqhD—醇脱氢酶;xdh—D-木糖脱氢酶;xylB—木糖激酶;xylC—木糖醛酸酯酶;yjhG/yagF—D-木酮酸脱氢酶;yjhH/yagE—2-酮-3-脱氧-D-戊糖醛缩酶;xylA—D-木糖异构酶;khk-C—D-木糖醛激酶;aldoB—D-木糖醛糖-1-磷酸醛缩酶;dte—D-塔格糖异构酶;fucK—岩藻糖激酶;fucA/rhaD—D-核糖醛糖-1-磷酸醛缩酶
Fig. 1 Biosynthetic pathways of EGX1P—Xylulose-1-phosphate; R1P—Ribulose-1-phosphate The enzymes encoded by the genes: serA—phosphoglycerate dehydrogenase; serC—phosphoserine aminotransferase; serB—phosphoserine phosphatase; sdC—L-serine decarboxylase; ao—amine oxidase; agt—serine-glyoxylate aminotransferase; mdlC—benzoylformate decarboxylase; fucO/yqhD—alcohol dehydrogenase; xdh—D-xylose dehydrogenase; xylB—xylulokinase; xylC—xylonolactonase; yjhG/yagF—D-xylonate dehydrogenase; yjhH/yagE—2-keto-3-deoxy-D-pentose aldolase; xylA—D-xylose isomerase; khk-C—D-xylulose kinase; aldoB—D-xylulose-1-phosphate aldolase; dte—D-tagatose epimerase; fucK—fuculokinase; fucA/rhaD—D-ribulose 1-phosphate aldolase
| 产物 | 合成路径 | 菌株 | 底物 | 基因改造策略 | 产量 /(g/L) | 得率 /(g/g) | 生产效率 /[g/(L·h)] | 理论转化率 /(g/g) | 参考文献 |
|---|---|---|---|---|---|---|---|---|---|
| 乙二醇(EG) | Dahms路径 | Escherichia coli | D-木糖 | (1)下调xylB表达 (2)过表达yqhD | 108.2 | 0.36 | 2.25 | 0.41 | [ |
| Dahms路径 | Escherichia coli | D-木糖 | (1)敲除arcA和aldA (2)过表达yjhH, xdh, xylC, fucO和yjhG | 72.0 | 0.40 | 1.38 | [ | ||
| R1P路径 | Escherichia coli | D-木糖 | (1)过表达fucK, fucO和fucA (2)敲除ald和xylB | 40 | 0.35 | 0.58 | 0.41 | [ | |
| X1P路径 | Escherichia coli | D-木糖 | (1)过表达khk-C, aldoB和fucO (2)敲除xylB和aldA | 20 | 0.38 | 0.37 | 0.41 | [ | |
| X1P路径 | Saccharomyces cerevisiae F251 | D-木糖 | (1)过表达pfk1和pfk2 (2)敲除xks1 | 4.05 | 0.12 | 0.06 | [ | ||
| 丝氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达aao, sdc, serABC和fucO (2)敲除aldA | 4.1 | 0.14 | — | 0.7 | [ | |
| 丝氨酸路径 | Corynebacterium glutamicum | 葡萄糖 | (1)过表达sgt, mdlC, sdc, AO和yqhD (2)插入serACB (3)敲除pabABC和sdaA | 3.5 | 0.09 | — | [ | ||
1,2-丙二醇 (1,2-PDO) | 丙酮醛途径 | K. pneumoniae | 甘油 | (1)过表达mgsA和yqhD (2)敲除tpiA | 9.3 | 0.20 | 0.06 | 0.56 | [ |
| 丙酮醛途径 | Escherichia coli | 葡萄糖 | (1)过表达mgsA, gldA, fdh1和fucO (2)敲除zwf, tpiA, adhE, gloA和ldhA | 5.13 | 0.48 | — | [ | ||
| 乳酸途径 | Escherichia coli | 葡萄糖 | (1)敲除adhE, dld, lldD, frdA, pflB, mgsA, aldA和arcA | 17.3 | 0.18 | 0.72 | 0.56 | [ | |
1,3-丙二醇 (1,3-PDO) | 甘油途径 | Escherichia coli | 葡萄糖 | — | 135 | 3.50 | — | 0.59 | [ |
| 甘油途径 | Corynebacterium glutamicum | 葡萄糖 | (1)敲除ald, pyk, adh, poxB, ldhA, ppc和zwf (2)过表达hdpA-gldA, gpd1, gpp2, yqhD, 和pduCEDGH (3)下调表达gapA | 110.4 | 0.42 | 2.30 | 0.59 | [ | |
| 甘油途径 | Vibrio natriegens | 甘油 | (1)敲除pta-ackA, arcA, adhE, aldB, ldh, pfl, sthA, glpR, aldA和frdABCD (2)过表达pntAB和phaP | 69.5 | 0.51 | 2.90 | 0.67 | [ | |
| 高丝氨酸途径 | Escherichia coli | 葡萄糖 | (1)敲除thrB (2)过表达yqhD, lysC, serCR42W/R77W, metL和pdc | 3.03 | — | 0.05 | 0.55 | [ | |
| 丙二酰辅酶A途径 | Escherichia coli | 葡萄糖 | (1)过表达mcrC, pduP, mcrN, yqhD和prpE | 7.98 | 0.15 | 0.22 | 0.54 | [ | |
| β-丙氨酸途径 | Escherichia coli | 葡萄糖 | (1)敲除lysC (2)过表达ppc (3)下调表达gltA | 11.21 | — | 0.10 | 0.61 | [ | |
1,3-丁二醇 (1,3-BDO) | 3-羟基丁酰辅酶A路径 | Escherichia coli | 葡萄糖 | (1)过表达phaAB和bld | 15.75 | 0.19 | 0.16 | 0.53 | [ |
| 3-羟基丁酰辅酶A路径 | Escherichia coli | 葡萄糖 | (1)过表达 bldL273T, yqhD, phaAB和 pntAB (2)敲除ldh, pta, ackA, adhE | 13.40 | 0.30 | 0.42 | [ | ||
| 3-羟基丁酸还原路径 | Escherichia coli | 葡萄糖 | (1)过表达phaAB, yqhD, pntAB, car和sfp (2)敲除ldh, pta, ackA, adhE | 0.40 | 0.02 | — | [ | ||
| 3-羟基丁酰辅酶A路径 | Escherichia coli | 葡萄糖 | (1)过表达 pk, glpX, thl, hbd, tesB和car (2)敲除 zwf, edd, pfkA和pfkB | 22.66 | 0.40 | 0.32 | 0.56 | [ | |
| 3-羟基丁酰辅酶A路径 | Escherichia coli | 葡萄糖 | (1)过表达phaAB, bld和yqhD (2)敲除adhE, poxB, ldhA, pta-ackA, atoB, tesB和yciA | 23.10 | 0.26 | 0.64 | [ | ||
| 3-羟基丁酰辅酶A路径 | Escherichia coli | 葡萄糖 | (1)过表达phaAB, bld, yjgB和zwf (2)敲除adhE, poxB, ldhA, yciA, pdhR, pgi和gntR | 71.10 | 0.34 | 1.55 | [ | ||
| DERA-AKR 路径 | Escherichia coli | 葡萄糖 | (1)过表达 AKR, DERA和 PDC (2)敲除pta, yjgB, adhE, ldhA, pflB, adhP, yqhD, eutG, ilvB, and poxB; | 2.40 | 0.06 | — | 0.55 | [ | |
1,4-丁二醇 (1,4-BDO) | 琥珀酰辅酶A路径 | Escherichia coli | 葡萄糖 | — | >125 | 0.40 | >3.5 | 0.5 | [ |
| 谷氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达gadB, gabT, yqhD, car, ppc, gltAR163L | 1.41 | 0.07 | 0.03 | 0.58 | [ | |
| 非磷酸化路径 | Escherichia coli | 葡萄糖、木糖 | (1)敲除yagE, xylA和yjhH (2)过表达KvidV461I, xylBCDX和yqhD | 12 | 0.26 (木糖) | 0.40 | 0.41 | [ | |
1,2-丁二醇 (1,2-BDO) | 苏氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达pyc, thrABC, ilvA, L-ldh, car, yqhD (2)敲除lldd, dld | 0.15 | — | — | 0.48 | [ |
2,3-丁二醇 (2,3-BDO) | 丙酮酸路径 | Saccharomyces cerevisiae | 葡萄糖 | (1)过表达BDH1 (2)下调PDC1, PDC6, 和AHD1 | 178 | 0.34 | 2.64 | 0.5 | [ |
| Corynebacterium glutamicum | 葡萄糖 | (1)过表达budABC和acs (2)敲除ldhA, adhE, frdA和pta | 144.9 | 0.43 | 1.10 | [ | |||
1,5-戊二醇 (1,5-PDO) | 赖氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达 lysCQ298G, asd,ddh, dapA,patA, patD, cadA, gabT, yahk, car, sfp, yqhD (2)敲除gabD, gdhA | 9.25 | 0.16 | 0.05 | 0.40 | [ |
| 赖氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达 lysC, dapA, davB, davA, gabT, yqhD, car和sfp (2)敲除iclR | 0.97 | 0.05 | — | 0.38 | [ | |
| 赖氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达 lysCQ298G, asd,ddh, dapA, davB, davA, gabT, yqhD, abfT, bldL273T 和pntAB | 0.12 | 0.006 | — | 0.3 | [ |
表1 生物合成二元醇的典型路径与代谢工程改造策略
Table 1 Typical pathways and metabolic engineering modification strategies for biosynthesis of diols
| 产物 | 合成路径 | 菌株 | 底物 | 基因改造策略 | 产量 /(g/L) | 得率 /(g/g) | 生产效率 /[g/(L·h)] | 理论转化率 /(g/g) | 参考文献 |
|---|---|---|---|---|---|---|---|---|---|
| 乙二醇(EG) | Dahms路径 | Escherichia coli | D-木糖 | (1)下调xylB表达 (2)过表达yqhD | 108.2 | 0.36 | 2.25 | 0.41 | [ |
| Dahms路径 | Escherichia coli | D-木糖 | (1)敲除arcA和aldA (2)过表达yjhH, xdh, xylC, fucO和yjhG | 72.0 | 0.40 | 1.38 | [ | ||
| R1P路径 | Escherichia coli | D-木糖 | (1)过表达fucK, fucO和fucA (2)敲除ald和xylB | 40 | 0.35 | 0.58 | 0.41 | [ | |
| X1P路径 | Escherichia coli | D-木糖 | (1)过表达khk-C, aldoB和fucO (2)敲除xylB和aldA | 20 | 0.38 | 0.37 | 0.41 | [ | |
| X1P路径 | Saccharomyces cerevisiae F251 | D-木糖 | (1)过表达pfk1和pfk2 (2)敲除xks1 | 4.05 | 0.12 | 0.06 | [ | ||
| 丝氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达aao, sdc, serABC和fucO (2)敲除aldA | 4.1 | 0.14 | — | 0.7 | [ | |
| 丝氨酸路径 | Corynebacterium glutamicum | 葡萄糖 | (1)过表达sgt, mdlC, sdc, AO和yqhD (2)插入serACB (3)敲除pabABC和sdaA | 3.5 | 0.09 | — | [ | ||
1,2-丙二醇 (1,2-PDO) | 丙酮醛途径 | K. pneumoniae | 甘油 | (1)过表达mgsA和yqhD (2)敲除tpiA | 9.3 | 0.20 | 0.06 | 0.56 | [ |
| 丙酮醛途径 | Escherichia coli | 葡萄糖 | (1)过表达mgsA, gldA, fdh1和fucO (2)敲除zwf, tpiA, adhE, gloA和ldhA | 5.13 | 0.48 | — | [ | ||
| 乳酸途径 | Escherichia coli | 葡萄糖 | (1)敲除adhE, dld, lldD, frdA, pflB, mgsA, aldA和arcA | 17.3 | 0.18 | 0.72 | 0.56 | [ | |
1,3-丙二醇 (1,3-PDO) | 甘油途径 | Escherichia coli | 葡萄糖 | — | 135 | 3.50 | — | 0.59 | [ |
| 甘油途径 | Corynebacterium glutamicum | 葡萄糖 | (1)敲除ald, pyk, adh, poxB, ldhA, ppc和zwf (2)过表达hdpA-gldA, gpd1, gpp2, yqhD, 和pduCEDGH (3)下调表达gapA | 110.4 | 0.42 | 2.30 | 0.59 | [ | |
| 甘油途径 | Vibrio natriegens | 甘油 | (1)敲除pta-ackA, arcA, adhE, aldB, ldh, pfl, sthA, glpR, aldA和frdABCD (2)过表达pntAB和phaP | 69.5 | 0.51 | 2.90 | 0.67 | [ | |
| 高丝氨酸途径 | Escherichia coli | 葡萄糖 | (1)敲除thrB (2)过表达yqhD, lysC, serCR42W/R77W, metL和pdc | 3.03 | — | 0.05 | 0.55 | [ | |
| 丙二酰辅酶A途径 | Escherichia coli | 葡萄糖 | (1)过表达mcrC, pduP, mcrN, yqhD和prpE | 7.98 | 0.15 | 0.22 | 0.54 | [ | |
| β-丙氨酸途径 | Escherichia coli | 葡萄糖 | (1)敲除lysC (2)过表达ppc (3)下调表达gltA | 11.21 | — | 0.10 | 0.61 | [ | |
1,3-丁二醇 (1,3-BDO) | 3-羟基丁酰辅酶A路径 | Escherichia coli | 葡萄糖 | (1)过表达phaAB和bld | 15.75 | 0.19 | 0.16 | 0.53 | [ |
| 3-羟基丁酰辅酶A路径 | Escherichia coli | 葡萄糖 | (1)过表达 bldL273T, yqhD, phaAB和 pntAB (2)敲除ldh, pta, ackA, adhE | 13.40 | 0.30 | 0.42 | [ | ||
| 3-羟基丁酸还原路径 | Escherichia coli | 葡萄糖 | (1)过表达phaAB, yqhD, pntAB, car和sfp (2)敲除ldh, pta, ackA, adhE | 0.40 | 0.02 | — | [ | ||
| 3-羟基丁酰辅酶A路径 | Escherichia coli | 葡萄糖 | (1)过表达 pk, glpX, thl, hbd, tesB和car (2)敲除 zwf, edd, pfkA和pfkB | 22.66 | 0.40 | 0.32 | 0.56 | [ | |
| 3-羟基丁酰辅酶A路径 | Escherichia coli | 葡萄糖 | (1)过表达phaAB, bld和yqhD (2)敲除adhE, poxB, ldhA, pta-ackA, atoB, tesB和yciA | 23.10 | 0.26 | 0.64 | [ | ||
| 3-羟基丁酰辅酶A路径 | Escherichia coli | 葡萄糖 | (1)过表达phaAB, bld, yjgB和zwf (2)敲除adhE, poxB, ldhA, yciA, pdhR, pgi和gntR | 71.10 | 0.34 | 1.55 | [ | ||
| DERA-AKR 路径 | Escherichia coli | 葡萄糖 | (1)过表达 AKR, DERA和 PDC (2)敲除pta, yjgB, adhE, ldhA, pflB, adhP, yqhD, eutG, ilvB, and poxB; | 2.40 | 0.06 | — | 0.55 | [ | |
1,4-丁二醇 (1,4-BDO) | 琥珀酰辅酶A路径 | Escherichia coli | 葡萄糖 | — | >125 | 0.40 | >3.5 | 0.5 | [ |
| 谷氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达gadB, gabT, yqhD, car, ppc, gltAR163L | 1.41 | 0.07 | 0.03 | 0.58 | [ | |
| 非磷酸化路径 | Escherichia coli | 葡萄糖、木糖 | (1)敲除yagE, xylA和yjhH (2)过表达KvidV461I, xylBCDX和yqhD | 12 | 0.26 (木糖) | 0.40 | 0.41 | [ | |
1,2-丁二醇 (1,2-BDO) | 苏氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达pyc, thrABC, ilvA, L-ldh, car, yqhD (2)敲除lldd, dld | 0.15 | — | — | 0.48 | [ |
2,3-丁二醇 (2,3-BDO) | 丙酮酸路径 | Saccharomyces cerevisiae | 葡萄糖 | (1)过表达BDH1 (2)下调PDC1, PDC6, 和AHD1 | 178 | 0.34 | 2.64 | 0.5 | [ |
| Corynebacterium glutamicum | 葡萄糖 | (1)过表达budABC和acs (2)敲除ldhA, adhE, frdA和pta | 144.9 | 0.43 | 1.10 | [ | |||
1,5-戊二醇 (1,5-PDO) | 赖氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达 lysCQ298G, asd,ddh, dapA,patA, patD, cadA, gabT, yahk, car, sfp, yqhD (2)敲除gabD, gdhA | 9.25 | 0.16 | 0.05 | 0.40 | [ |
| 赖氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达 lysC, dapA, davB, davA, gabT, yqhD, car和sfp (2)敲除iclR | 0.97 | 0.05 | — | 0.38 | [ | |
| 赖氨酸路径 | Escherichia coli | 葡萄糖 | (1)过表达 lysCQ298G, asd,ddh, dapA, davB, davA, gabT, yqhD, abfT, bldL273T 和pntAB | 0.12 | 0.006 | — | 0.3 | [ |
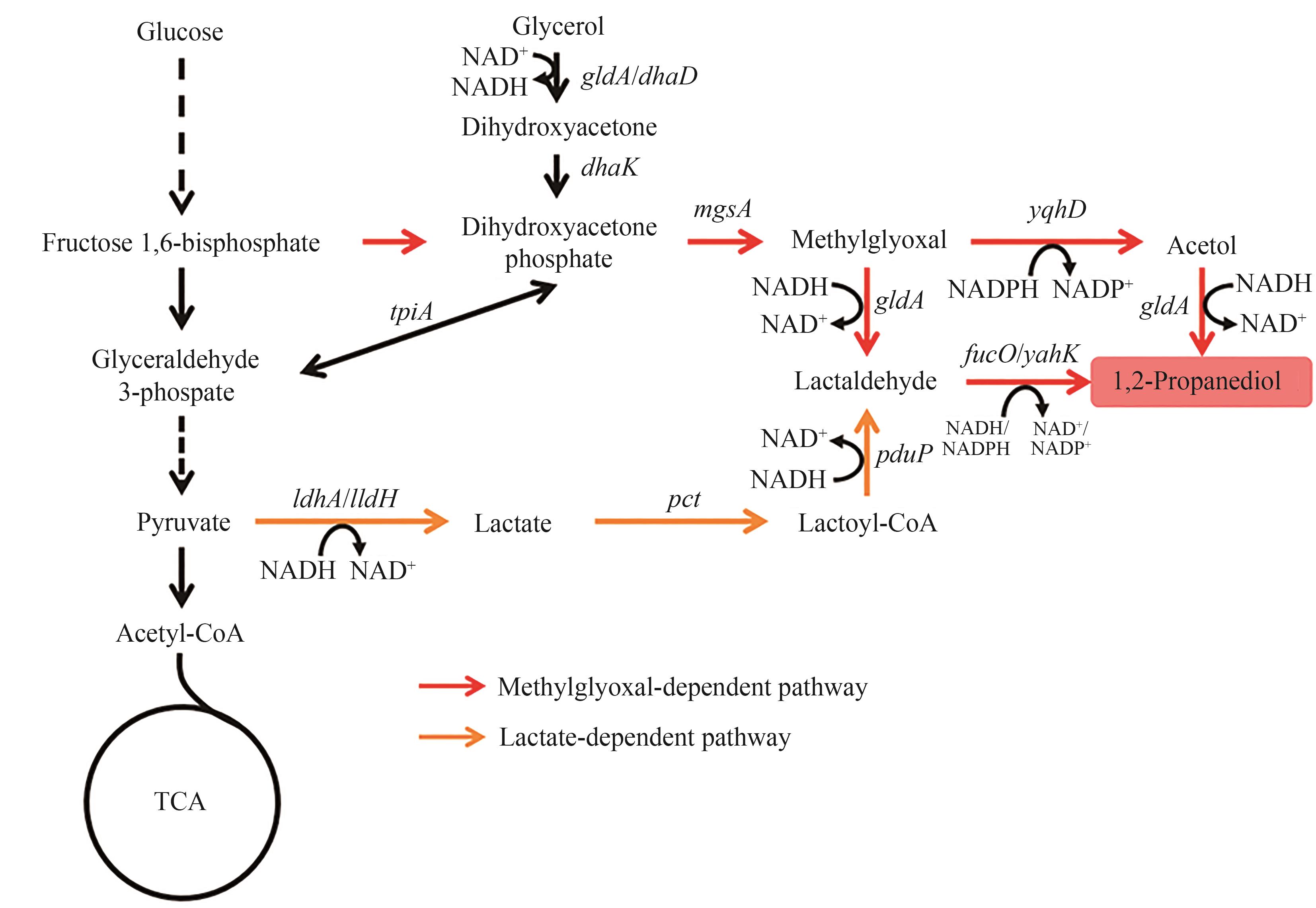
图2 1,2-丙二醇的生物合成路径图中基因所编码的酶:gldA/dhaD—甘油脱氢酶;dhaK—PEP依赖型二羟基丙酮激酶;mgsA—甲基乙二醛合酶;yqhD—醇脱氢酶;yahK—醇脱氢酶;fucO—醇脱氢酶;ldhA/lldH—乳酸脱氢酶;pct—丙酸辅酶A转移酶;pduP—醛脱氢酶
Fig. 2 Biosynthetic pathways of 1,2-PDOThe enzymes encoded by the genes: gldA/dhaD—glycerol dehydrogenase; dhaK—PEP-dependent dihydroxyacetone kinase; mgsA—methylglyoxal synthase; yqhD—alcohol dehydrogenase; yahK—alcohol dehydrogenase; fucO—alcohol dehydrogenase; ldhA/lldH—lactate dehydrogenase; pct—propionate CoA-transferase; pduP, aldehyde dehydrogenase
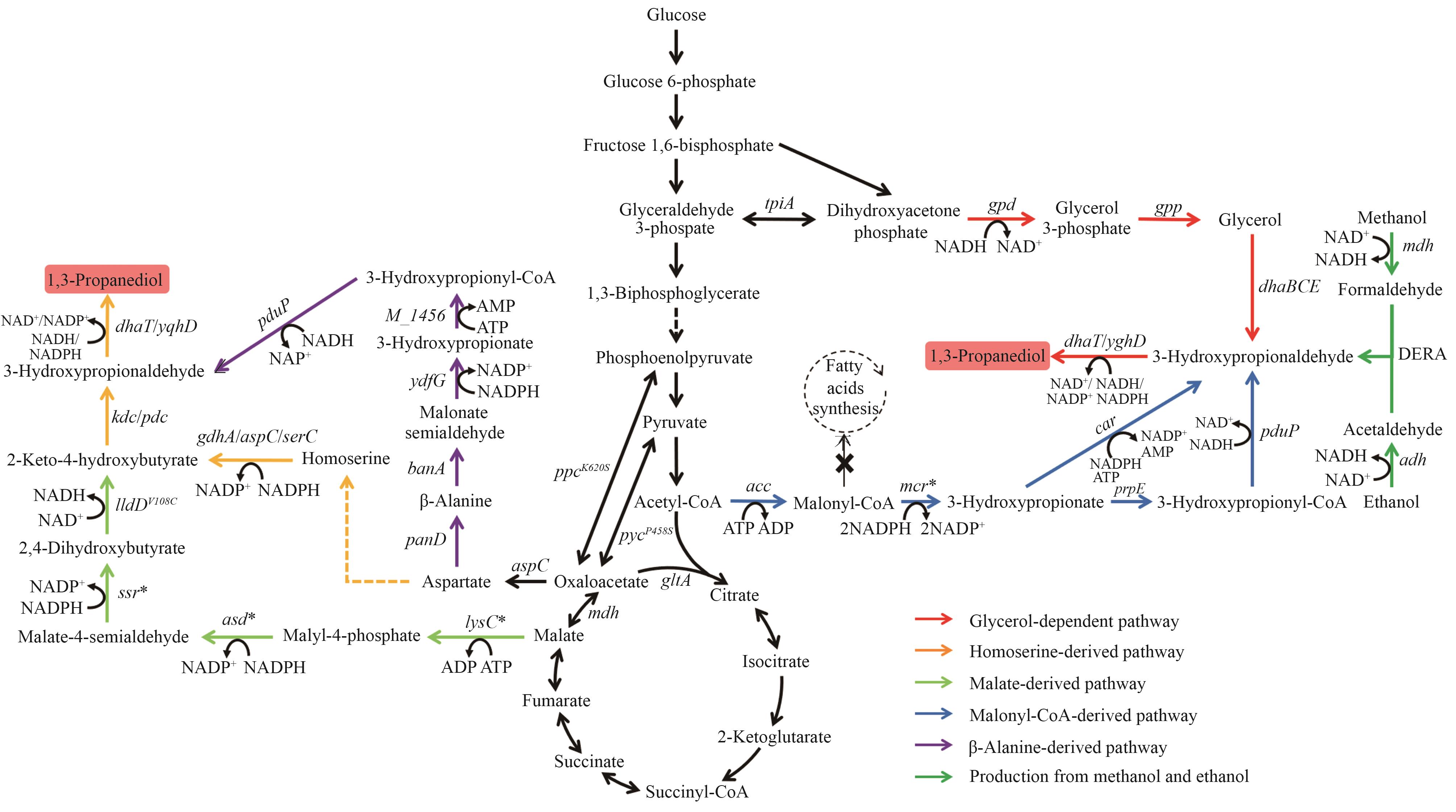
图3 1,3-丙二醇的生物合成路径图中基因所编码的酶:tpiA—丙糖磷酸异构酶;gpd—甘油-3-磷酸脱氢酶;gpp—甘油-3-磷酸磷酸酶;dhaBCE—维生素B12依赖的甘油脱水酶;dhaT/yqhD—醇脱氢酶;acc—乙酰辅酶A羧化酶;mcr—丙二酰辅酶A还原酶;prpE—3-羟基丙酰辅酶A合成酶;pduP—醛脱氢酶;car—羧酸还原酶;mdh—甲醇脱氢酶;adh—醇脱氢酶;DERA—脱氧核糖-5-磷酸醛缩酶;aspC—天冬氨酸转氨酶;panD—天冬氨酸脱羧酶;bauA—β-丙氨酸-丙酮酸氨基转移酶;ydfG—3-羟基酸脱氢酶;M_1456—3-羟基丙酰辅酶A合酶;lysC—苹果酸激酶;asd—苹果酸半醛脱氢酶;ssr—苹果酸半醛还原酶;lldD—乳酸脱氢酶;kdc—酮酸脱羧酶;pdc—丙酮酸脱羧酶;gdhA—谷氨酸脱氢酶;serC—磷酸丝氨酸氨基转移酶
Fig. 3 Biosynthetic pathways of 1,3-PDOThe enzymes encoded by the genes: tpiA—triose-phosphate isomerase; gpd—glycerol-3-phosphate dehydrogenase; gpp—glycerol-3-phosphate phosphatase; dhaBCE—vitamin B12-dependent glycerol dehydratase; dhaT/yqhD—alcohol dehydrogenase; acc—acetyl-CoA carboxylase; mcr—malonyl-CoA reductase; prpE—3-hydroxypropionyl-CoA synthetase; pduP—aldehyde dehydrogenase; car—carboxylic acid reductase; mdh—methanol dehydrogenase; adh—alcohol dehydrogenase; DERA—deoxyribose-5-phosphate aldolase; aspC—aspartate transaminase; panD—aspartate decarboxylase; bauA—β-alanine-pyruvate aminotransferase; ydfG—3-hydroxy acid dehydrogenase; M_1456—3-hydroxypropionyl-coenzyme A synthetase; lysC—malate kinase; asd—malate semialdehyde dehydrogenase; ssr—malate semialdehyde reductase; lldD—lactate dehydrogenase; kdc—ketoacid decarboxylase; pdc—pyruvate decarboxylase; gdhA—glutamate dehydrogenase; serC—phosphoserine aminotransferase
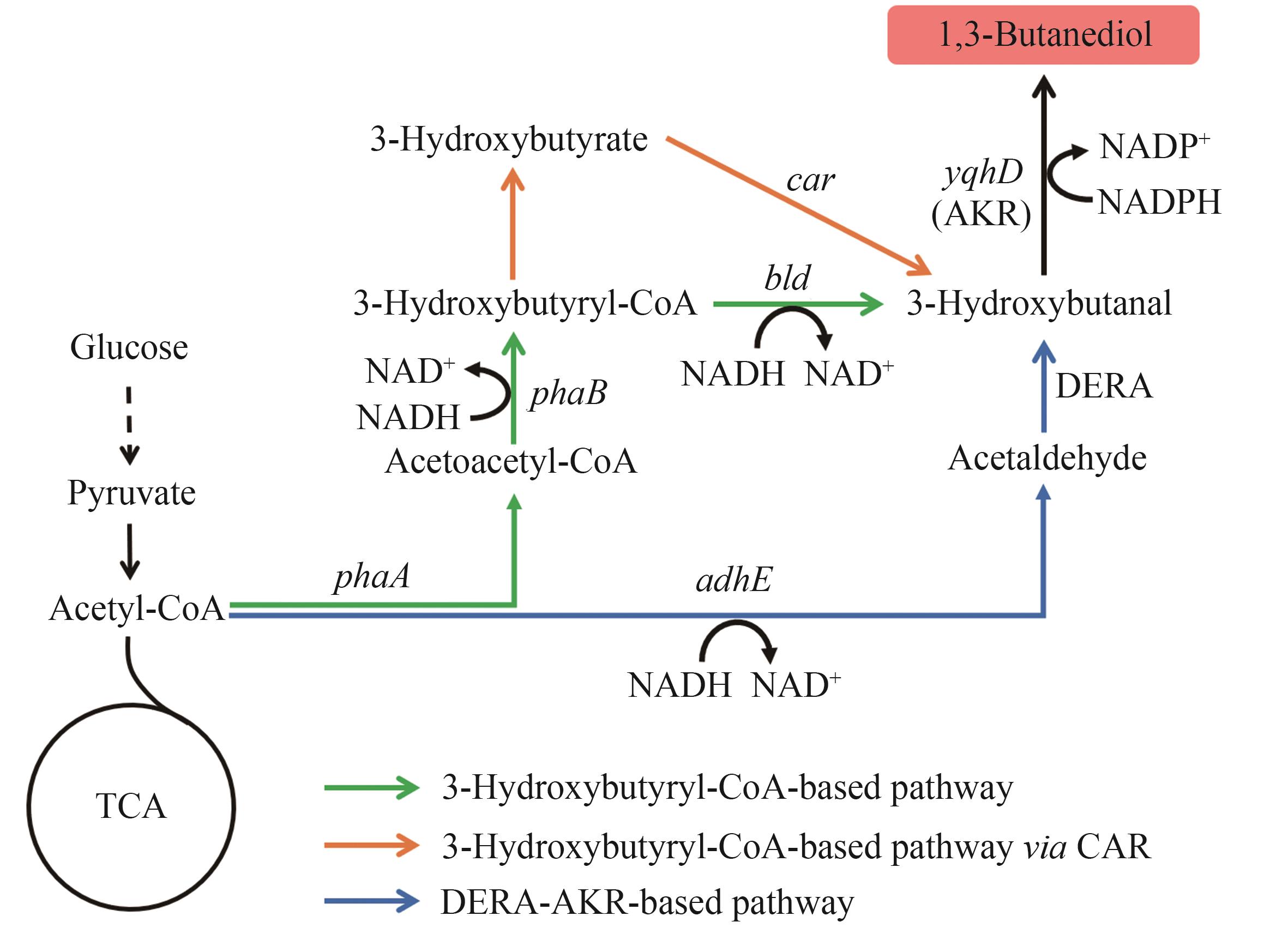
图4 1,3-丁二醇的生物合成路径图中基因所编码的酶:adhE—醇脱氢酶;DERA—脱氧核糖-5-磷酸醛缩酶;yqhD—醇脱氢酶;phaA—乙酰辅酶A乙酰转移酶;phaB—乙酰乙酰辅酶A还原酶;bld—3-羟基丁酰辅酶A脱氢酶;car—羧酸还原酶
Fig. 4 Biosynthetic pathways of 1,3-BDOThe enzymes encoded by the genes: adhE—alcohol dehydrogenase; DERA—deoxyribose-5-phosphate aldolase; yqhD—alcohol dehydrogenase; phaA—acetyl-CoA acetyltransferase; phaB—acetoacetyl-CoA reductase; bld—3-hydroxybutyryl-CoA dehydrogenase; car—carboxylic acid reductase
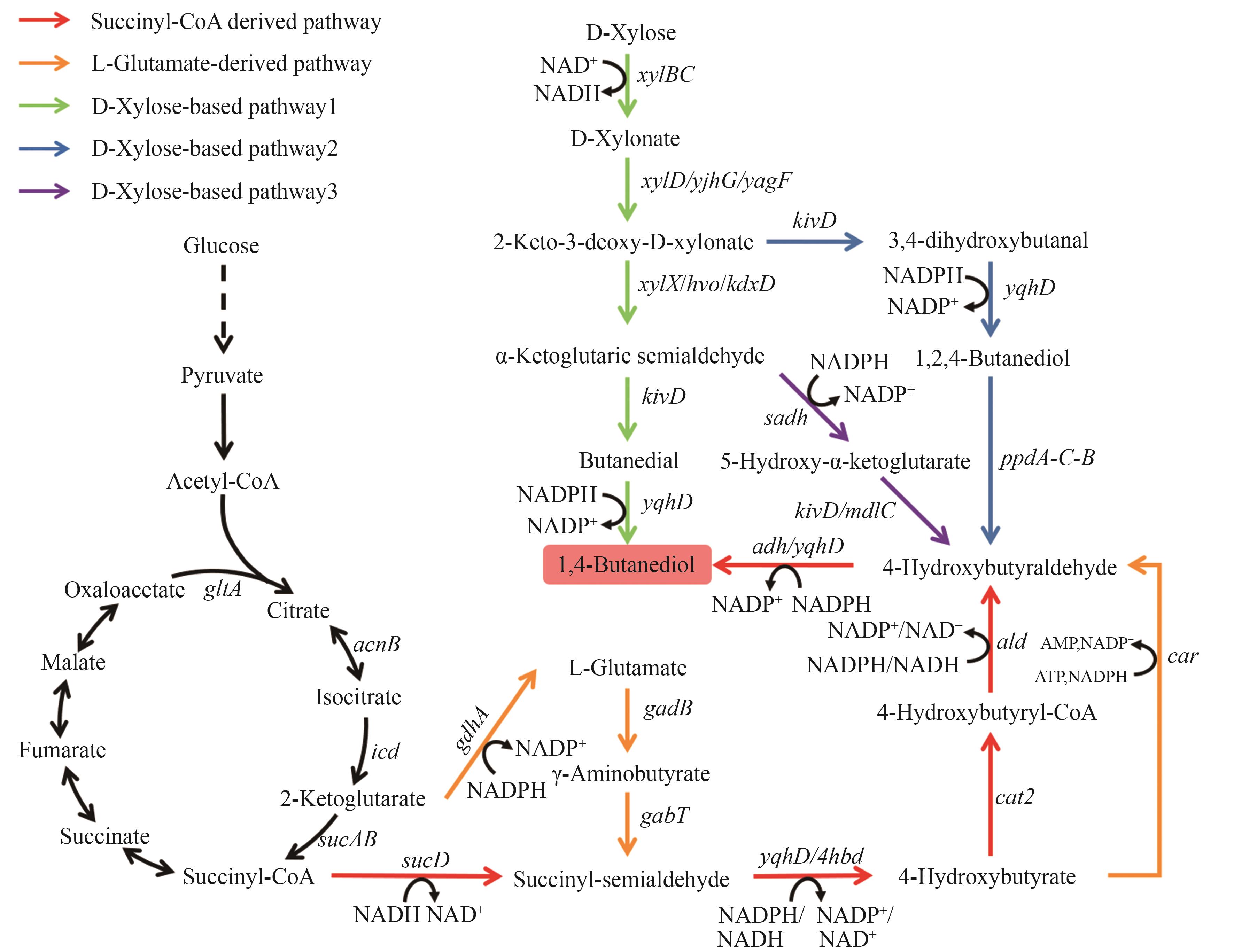
图5 1,4-丁二醇的生物合成路径图中基因所编码的酶:gdhA—谷氨酸脱氢酶;gadB—谷氨酸脱羧酶;gabT—转氨酶;sucD—琥珀酸半醛脱氢酶;yqhD/4hbd/adh—醇脱氢酶;cat2—4-羟基丁酸辅酶A转移酶;ald—醛脱氢酶;car—羧酸还原酶;xylBC—D-木糖脱氢酶;xylD/yjhG/yagF—D-木酮酸脱水酶;xylX/hvo/kdxD—2-酮-3-脱氧-D-木酮酸脱水酶;sadh—醇脱氢酶;kivD/mdlC—脱羧酶
Fig. 5 Biosynthetic pathways of 1,4-BDOThe enzymes encoded by the genes: gdhA—glutamate dehydrogenase; gadB—glutamate decarboxylase; gabT—aminotransferase; sucD—succinate semialdehyde dehydrogenase; yqhD/4hbd/adh—alcohol dehydrogenase; cat2—4-hydroxybutyrate-CoA transferase; ald—aldehyde dehydrogenase; car—carboxylic acid reductase; xylBC—D-xylose dehydrogenase; xylD/yjhG/yagF—D-xylonate dehydratase; xylX/hvo/kdxD—2-keto-3-deoxy-D-xylonate dehydratase; sadh—alcohol dehydrogenase; kivD/mdlC—decarboxylase
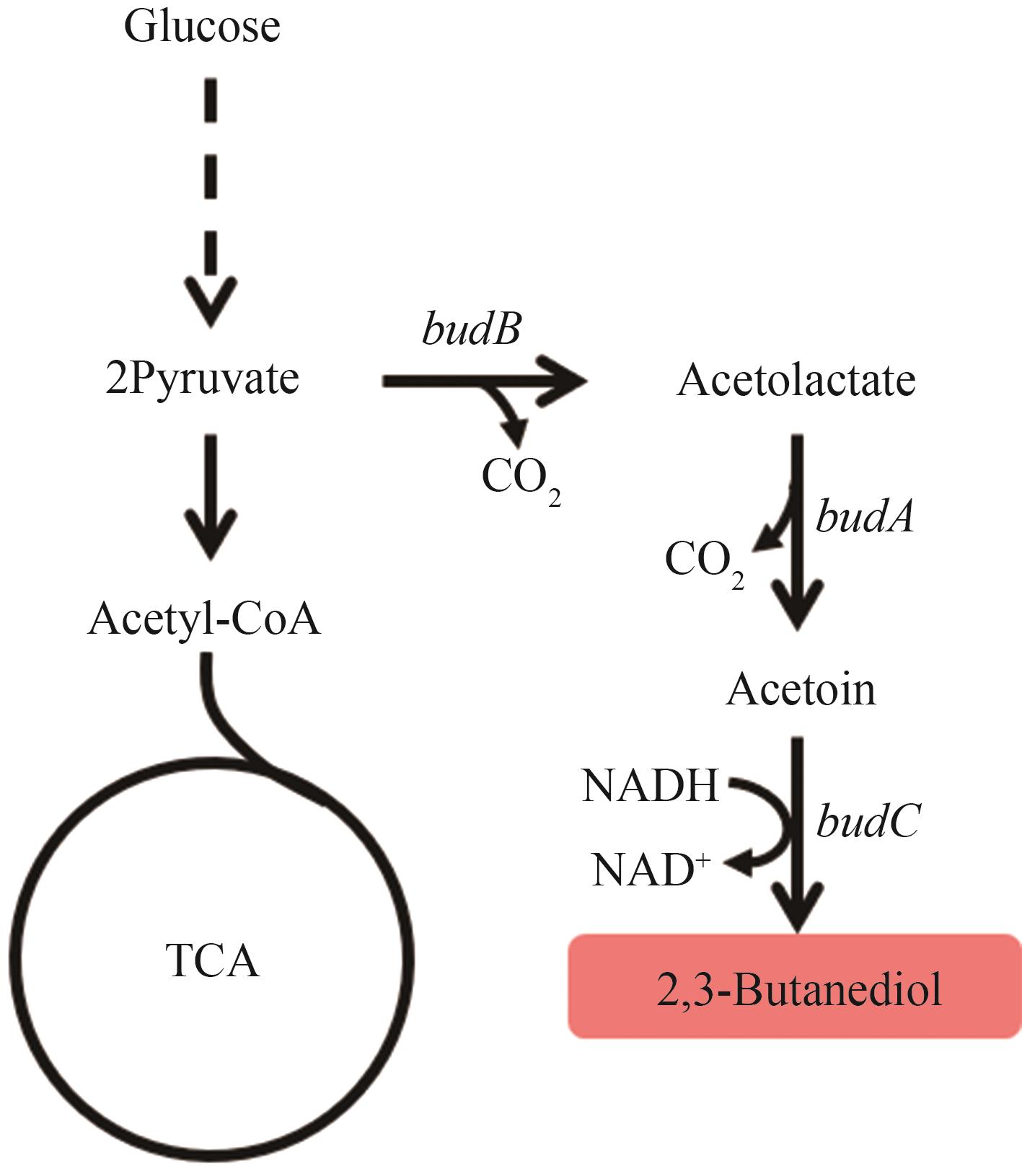
图6 2,3-丁二醇的生物合成路径图中基因所编码的酶:budB—乙酰乳酸合酶;budA—乙酰乳酸脱羧酶;budC—2,3-丁二醇脱氢酶
Fig. 6 Biosynthetic pathways of 2,3-BDOThe enzymes encoded by the genes: budB—acetolactate synthase; budA—acetolactate decarboxylase; budC—2,3-butanediol dehydrogenase
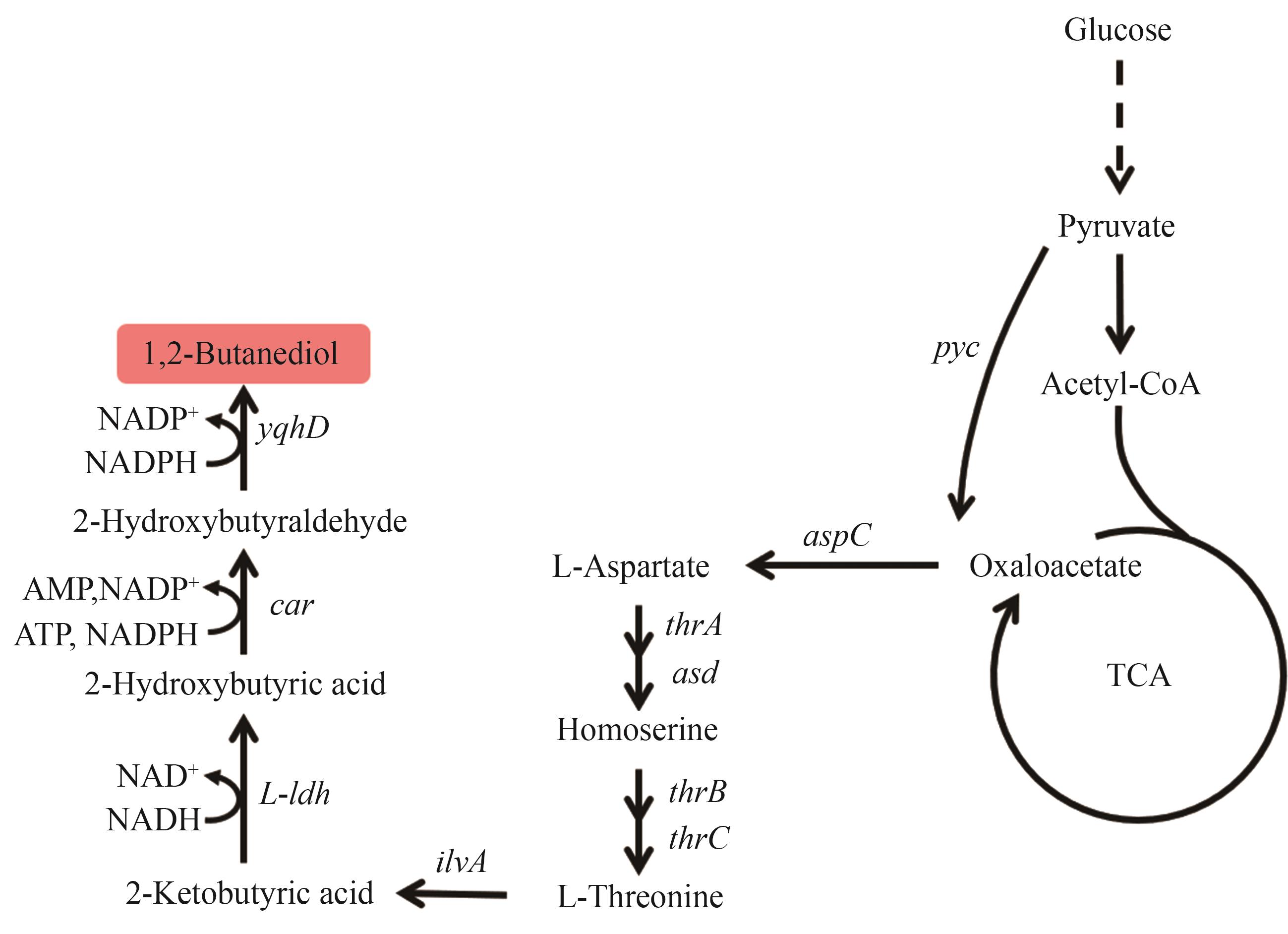
图7 1,2-丁二醇的生物合成路径图中基因所编码的酶:pyc—丙酮酸羧化酶;aspC—天冬氨酸转氨酶;thrA—高丝氨酸脱氢酶;asd—天冬氨酸半醛脱氢酶;thrB—高丝氨酸激酶;thrC—苏氨酸合酶;ilvA—L-苏氨酸脱水酶;L-ldh—L-乳酸脱氢酶;car—羧酸还原酶;yqhD—醇脱氢酶
Fig. 7 Biosynthetic pathway of 1,2-BDOThe enzymes encoded by the genes: pyc—pyruvate carboxylase; aspC—aspartate transaminase; thrA—homoserine dehydrogenase; asd—aspartate semialdehyde dehydrogenase; thrB—homoserine kinase; thrC—threonine synthase; ilvA—L-threonine dehydratase; L-ldh—L-lactate dehydrogenase; car—carboxylic acid reductase; yqhD—alcohol dehydrogenase
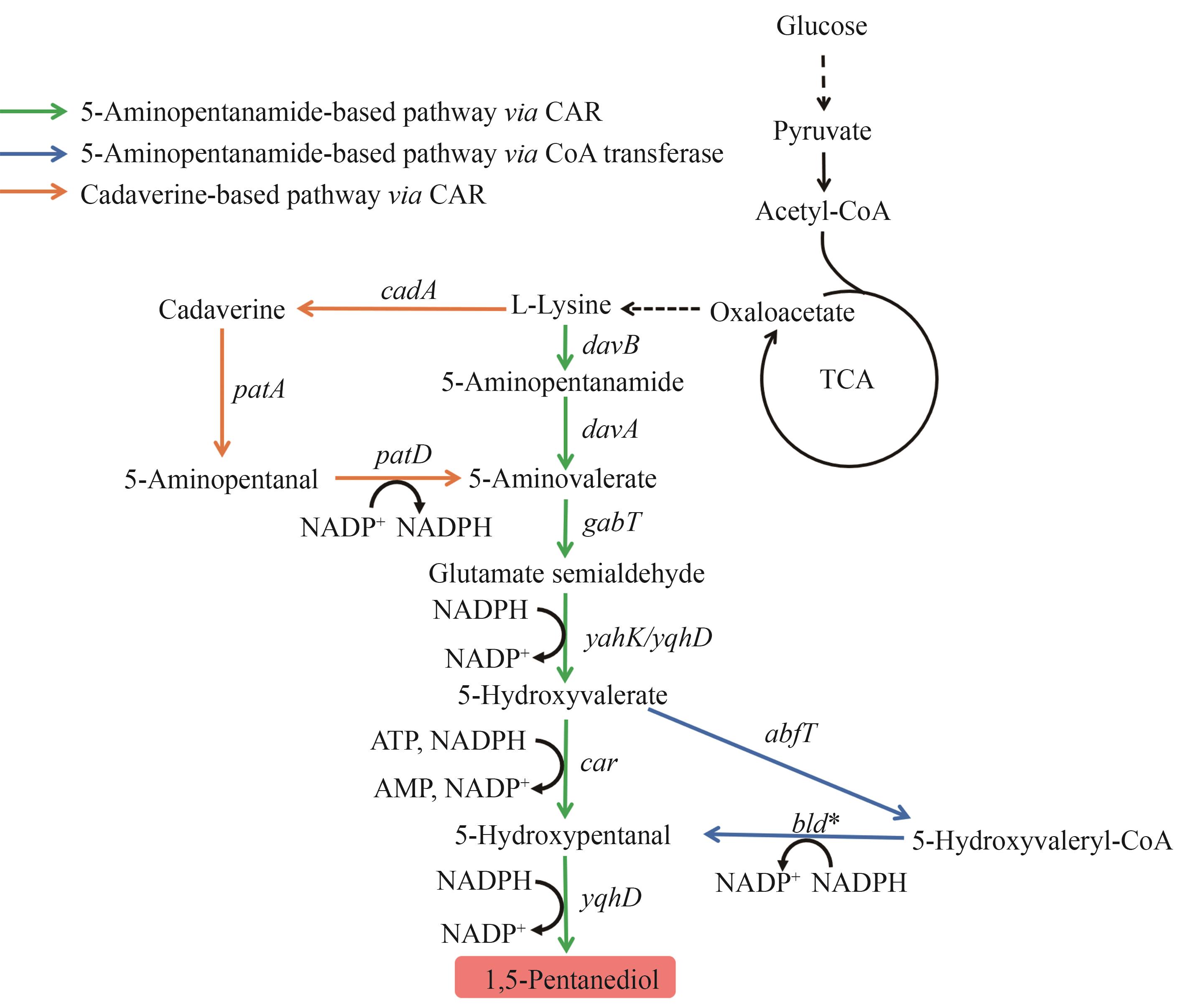
图8 1,5-戊二醇的生物合成路径图中基因所编码的酶:davB—赖氨酸单加氧酶;davA—5-氨基己酰胺酶;gabT—4-氨基丁酸转氨酶;yahK/yqhD—醇脱氢酶;car—羧酸还原酶;abfT—5-羟基戊酸辅酶A转移酶;bld—醛脱氢酶;cadA—赖氨酸脱羧酶;patA—丁二胺转氨酶;patD—醇脱氢酶
Fig. 8 Biosynthetic pathways of 1,5-PDOThe enzymes encoded by the genes: davB—lysine monooxygenase; davA—5-aminovaleramidase; gabT—4-aminobutyrate aminotransferase; yahK/yqhD—alcohol dehydrogenase; car—carboxylic acid reductase; abfT—5-hydroxyvalerate-CoA transferase; bld—aldehyde dehydrogenase; cadA—lysine decarboxylase; patA—putrescine aminotransferase; patD—alcohol dehydrogenase
| 1 | CHOI S, SONG C W, SHIN J H, et al. Biorefineries for the production of top building block chemicals and their derivatives[J]. Metabolic Engineering, 2015, 28: 223-239. |
| 2 | LIU Y F, WANG W, ZENG A P. Biosynthesizing structurally diverse diols via a general route combining oxidative and reductive formations of OH-groups[J]. Nature Communications, 2022, 13(1): 1595. |
| 3 | LIU Y, CEN X C, LIU D H, et al. Metabolic engineering of Escherichia coli for high-yield production of (R)-1,3-butanediol[J]. ACS Synthetic Biology, 2021, 10(8): 1946-1955. |
| 4 | ZHANG Y, LIU D H, CHEN Z. Production of C2-C4 diols from renewable bioresources: new metabolic pathways and metabolic engineering strategies[J]. Biotechnology for Biofuels, 2017, 10: 299. |
| 5 | CEN X C, LIU Y J, ZHU F H, et al. Metabolic engineering of Escherichia coli for high production of 1,5-pentanediol via a cadaverine-derived pathway[J]. Metabolic Engineering, 2022, 74: 168-177. |
| 6 | CEN X C, DONG Y, LIU D H, et al. New pathways and metabolic engineering strategies for microbial synthesis of diols[J]. Current Opinion in Biotechnology, 2022, 78: 102845. |
| 7 | VIVEK N, HAZEENA S H, ALPHY M P, et al. Recent advances in microbial biosynthesis of C3-C5 diols: genetics and process engineering approaches[J]. Bioresource Technology, 2021, 322: 124527. |
| 8 | WU T, LIU Y M, LIU J S, et al. Metabolic engineering and regulation of diol biosynthesis from renewable biomass in Escherichia coli [J]. Biomolecules, 2022, 12(5): 715. |
| 9 | 陈勇,李凤梅,韩荣伟, 等. 酵母不对称催化制备R-1,2丙二醇[J].生物加工过程, 2009, 7(4): 61-64. |
| CHEN Y, LI F M, HAN R W, et al. Preparation of (R)-1,2-propanediol through asymmetric reduction with bakers yeast[J]. Chinese Journal of Bioprocess Engineering, 2009, 7(4): 61-64. | |
| 10 | MATSUYAMA A, YAMAMOTO H, KAWADA N, et al. Industrial production of (R)-1,3-butanediol by new biocatalysts[J]. Journal of Molecular Catalysis B: Enzymatic, 2001, 11(4/5/6): 513-521. |
| 11 | ZHU F H, LIU D H, CHEN Z. Recent advances in biological production of 1,3-propanediol: new routes and engineering strategies[J]. Green Chemistry, 2022, 24(4): 1390-1403. |
| 12 | BURGARD A, BURK M J, OSTERHOUT R, et al. Development of a commercial scale process for production of 1,4-butanediol from sugar[J]. Current Opinion in Biotechnology, 2016, 42: 118-125. |
| 13 | SALUSJÄRVI L, HAVUKAINEN S, KOIVISTOINEN O, et al. Biotechnological production of glycolic acid and ethylene glycol: current state and perspectives[J]. Applied Microbiology and Biotechnology, 2019, 103(6): 2525-2535. |
| 14 | YUE H R, ZHAO Y J, MA X B, et al. Ethylene glycol: properties, synthesis, and applications[J]. Chemical Society Reviews, 2012, 41(11): 4218-4244. |
| 15 | CHAE T U, CHOI S Y, RYU J Y, et al. Production of ethylene glycol from xylose by metabolically engineered Escherichia coli [J]. AIChE Journal, 2018, 64(12): 4193-4200. |
| 16 | WANG Y H, XIAN M, FENG X J, et al. Biosynthesis of ethylene glycol from D-xylose in recombinant Escherichia coli [J]. Bioengineered, 2018, 9(1): 233-241. |
| 17 | PEREIRA B, LI Z J, DE MEY M, et al. Efficient utilization of pentoses for bioproduction of the renewable two-carbon compounds ethylene glycol and glycolate[J]. Metabolic Engineering, 2016, 34: 80-87. |
| 18 | URANUKUL B, WOOLSTON B M, FINK G R, et al. Biosynthesis of monoethylene glycol in Saccharomyces cerevisiae utilizing native glycolytic enzymes[J]. Metabolic Engineering, 2019, 51: 20-31. |
| 19 | ALKIM C, CAM Y, TRICHEZ D, et al. Optimization of ethylene glycol production from (D)-xylose via a synthetic pathway implemented in Escherichia coli [J]. Microbial Cell Factories, 2015, 14: 127. |
| 20 | PEREIRA B, ZHANG H R, DE MEY M, et al. Engineering a novel biosynthetic pathway in Escherichia coli for production of renewable ethylene glycol[J]. Biotechnology and Bioengineering, 2016, 113(2): 376-383. |
| 21 | CHEN Z, HUANG J H, WU Y, et al. Metabolic engineering of Corynebacterium glutamicum for the de novo production of ethylene glycol from glucose[J]. Metabolic Engineering, 2016, 33: 12-18. |
| 22 | SUN S Q, SHU L, LU X Y, et al. 1,2-Propanediol production from glycerol via an endogenous pathway of Klebsiella pneumoniae [J]. Applied Microbiology and Biotechnology, 2021, 105(23): 9003-9016. |
| 23 | JAIN R, SUN X X, YUAN Q P, et al. Systematically engineering Escherichia coli for enhanced production of 1,2-propanediol and 1-propanol[J]. ACS Synthetic Biology, 2015, 4(6): 746-756. |
| 24 | NIU W, KRAMER L, MUELLER J, et al. Metabolic engineering of Escherichia coli for the de novo stereospecific biosynthesis of 1,2-propanediol through lactic acid[J]. Metabolic Engineering Communications, 2019, 8: e00082. |
| 25 | KURIAN J V. A new polymer platform for the future—sorona® from corn derived 1,3-propanediol[J]. Journal of Polymers and the Environment, 2005, 13(2): 159-167. |
| 26 | LI Z H, DONG Y F, LIU Y, et al. Systems metabolic engineering of Corynebacterium glutamicum for high-level production of 1,3-propanediol from glucose and xylose[J]. Metabolic Engineering, 2022, 70: 79-88. |
| 27 | ZHANG Y, SUN Q, LIU Y, et al. Development of a plasmid stabilization system in Vibrio natriegens for the high production of 1,3-propanediol and 3-hydroxypropionate[J]. Bioresources and Bioprocessing, 2021, 8(1): 125. |
| 28 | ZHANG Y J, MA C W, DISCHERT W, et al. Engineering of phosphoserine aminotransferase increases the conversion of L-homoserine to 4-hydroxy-2-ketobutyrate in a glycerol-independent pathway of 1,3-propanediol production from glucose[J]. Biotechnology Journal, 2019, 14(9): e1900003. |
| 29 | LI Z H, WU Z Y, CEN X C, et al. Efficient production of 1,3-propanediol from diverse carbohydrates via a non-natural pathway using 3-hydroxypropionic acid as an intermediate[J]. ACS Synthetic Biology, 2021, 10(3): 478-486. |
| 30 | LI M D, ZHANG Y, LI J C, et al. Biosynthesis of 1,3-propanediol via a new pathway from glucose in Escherichia coli [J]. ACS Synthetic Biology, 2023, 12(7): 2083-2093. |
| 31 | KATAOKA N, VANGNAI A S, UEDA H, et al. Enhancement of (R)-1,3-butanediol production by engineered Escherichia coli using a bioreactor system with strict regulation of overall oxygen transfer coefficient and pH[J]. Bioscience, Biotechnology, and Biochemistry, 2014, 78(4): 695-700. |
| 32 | WANG J, ZHANG R H, ZHANG J L, et al. Tunable hybrid carbon metabolism coordination for the carbon-efficient biosynthesis of 1,3-butanediol in Escherichia coli [J]. Green Chemistry, 2021, 23(21): 8694-8706. |
| 33 | ISLAM T, NGUYEN-VO T P, GAUR V K, et al. Metabolic engineering of Escherichia coli for biological production of 1,3-butanediol[J]. Bioresource Technology, 2023, 376: 128911. |
| 34 | ISLAM T, NGUYEN-VO T P, CHO S, et al. Metabolic engineering of Escherichia coli for enhanced production of 1,3-butanediol from glucose[J]. Bioresource Technology, 2023, 389: 129814. |
| 35 | KIM T, FLICK R, BRUNZELLE J, et al. Novel Aldo-Keto reductases for the biocatalytic conversion of 3-hydroxybutanal to 1,3-butanediol: structural and biochemical studies[J]. Applied and Environmental Microbiology, 2017, 83(7): e03172-16. |
| 36 | WANG J, LI C Y, ZOU Y S, et al. Bacterial synthesis of C3-C5 diols via extending amino acid catabolism[J]. Proceedings of the National Academy of Sciences of the United States of America, 2020, 117(32): 19159-19167. |
| 37 | TAI Y S, XIONG M Y, JAMBUNATHAN P, et al. Engineering nonphosphorylative metabolism to generate lignocellulose-derived products[J]. Nature Chemical Biology, 2016, 12(4): 247-253. |
| 38 | QIN N, ZHU F H, LIU Y Y, et al. Metabolic engineering of Escherichia coli for de novo production of 1,2-butanediol[J]. ACS Synthetic Biology, 2024, 13(1): 351-357. |
| 39 | LEE Y G, SEO J H. Production of 2,3-butanediol from glucose and cassava hydrolysates by metabolically engineered industrial polyploid Saccharomyces cerevisiae [J]. Biotechnology for Biofuels, 2019, 12: 204. |
| 40 | KOU M Y, CUI Z Z, FU J, et al. Metabolic engineering of Corynebacterium glutamicum for efficient production of optically pure (2R,3R)-2,3-butanediol[J]. Microbial Cell Factories, 2022, 21(1): 150. |
| 41 | CEN X C, LIU Y, CHEN B, et al. Metabolic engineering of Escherichia coli for de novo production of 1,5-pentanediol from glucose[J]. ACS Synthetic Biology, 2021, 10(1): 192-203. |
| 42 | TAO Y M, BU C Y, ZOU L H, et al. A comprehensive review on microbial production of 1,2-propanediol: micro-organisms, metabolic pathways, and metabolic engineering[J]. Biotechnology for Biofuels, 2021, 14(1): 216. |
| 43 | MARINAS A, BRUIJNINCX P, FTOUNI J, et al. Sustainability metrics for a fossil- and renewable-based route for 1,2-propanediol production: a comparison[J]. Catalysis Today, 2015, 239: 31-37. |
| 44 | INGVADOTTIR E M, SCULLY S M, ORLYGSSON J. Production of (S)-1,2-propanediol from L-rhamnose using the moderately thermophilic Clostridium strain AK1[J]. Anaerobe, 2018, 54: 26-30. |
| 45 | BADÍA J, ROS J, AGUILAR J. Fermentation mechanism of fucose and rhamnose in Salmonella typhimurium and Klebsiella pneumoniae [J]. Journal of Bacteriology, 1985, 161(1): 435-437. |
| 46 | JUN S A, MOON C, KANG C H, et al. Microbial fed-batch production of 1,3-propanediol using raw glycerol with suspended and immobilized Klebsiella pneumoniae [J]. Applied Biochemistry and Biotechnology, 2010, 161(1-8): 491-501. |
| 47 | LIU H J, XU Y Z, ZHENG Z M, et al. 1, 3-Propanediol and its copolymers: research, development and industrialization[J]. Biotechnology Journal, 2010, 5(11): 1137-1148. |
| 48 | YANG M M, YUN J H, ZHANG H H, et al. Genetically engineered strains: application and advances for 1,3-propanediol production from glycerol[J]. Food Technology and Biotechnology, 2018, 56(1): 3-15. |
| 49 | OH B R, LEE S M, HEO S Y, et al. Efficient production of 1,3-propanediol from crude glycerol by repeated fed-batch fermentation strategy of a lactate and 2,3-butanediol deficient mutant of Klebsiella pneumoniae [J]. Microbial Cell Factories, 2018, 17(1): 92. |
| 50 | ZHOU S, HUANG Y H, MAO X L, et al. Impact of acetolactate synthase inactivation on 1,3-propanediol fermentation by Klebsiella pneumoniae [J]. PLoS One, 2019, 14(4): e0200978. |
| 51 | CHEN Z, GENG F, ZENG A P. Protein design and engineering of a de novo pathway for microbial production of 1,3-propanediol from glucose[J]. Biotechnology Journal, 2015, 10(2): 284-289. |
| 52 | ZHONG W Q, ZHANG Y, WU W J, et al. Metabolic engineering of a homoserine-derived non-natural pathway for the de novo production of 1,3-propanediol from glucose[J]. ACS Synthetic Biology, 2019, 8(3): 587-595. |
| 53 | WALTHER T, TOPHAM C M, IRAGUE R, et al. Construction of a synthetic metabolic pathway for biosynthesis of the non-natural methionine precursor 2,4-dihydroxybutyric acid[J]. Nature Communications, 2017, 8: 15828. |
| 54 | FRAZÃO C J R, TRICHEZ D, SERRANO-BATAILLE H, et al. Construction of a synthetic pathway for the production of 1,3-propanediol from glucose[J]. Scientific Reports, 2019, 9(1): 11576. |
| 55 | ZHANG C J, SHARMA S, MA C W, et al. Strain evolution and novel downstream processing with integrated catalysis enable highly efficient coproduction of 1,3-propanediol and organic acid esters from crude glycerol[J]. Biotechnology and Bioengineering, 2022, 119(6): 1450-1466. |
| 56 | MENG H, WANG C, YUAN Q P, et al. An aldolase-based new pathway for bioconversion of formaldehyde and ethanol into 1,3-propanediol in Escherichia coli [J]. ACS Synthetic Biology, 2021, 10(4): 799-809. |
| 57 | TROTTER C L, BABU G S, WALLACE S. Engineering biology for sustainable 1,4-butanediol synthesis[J]. Trends in Biotechnology, 2023, 41(3): 286-288. |
| 58 | ZHU Y, YANG J M, MEI F, et al. Bio-based 1,4-butanediol and tetrahydrofuran synthesis: perspective[J]. Green Chemistry, 2022, 24(17): 6450-6466. |
| 59 | YIM H, HASELBECK R, NIU W, et al. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol[J]. Nature Chemical Biology, 2011, 7(7): 445-452. |
| 60 | BARTON N R, BURGARD A P, BURK M J, et al. An integrated biotechnology platform for developing sustainable chemical processes[J]. Journal of Industrial Microbiology & Biotechnology, 2015, 42(3): 349-360. |
| 61 | LIU H W, LU T. Autonomous production of 1, 4-butanediol via a de novo biosynthesis pathway in engineered Escherichia coli [J]. Metabolic Engineering, 2015, 29: 135-141. |
| 62 | WANG J, JAIN R, SHEN X L, et al. Rational engineering of diol dehydratase enables 1,4-butanediol biosynthesis from xylose[J]. Metabolic Engineering, 2017, 40: 148-156. |
| 63 | LIU X, FABOS V, TAYLOR S, et al. One-step production of 1,3-butadiene from 2,3-butanediol dehydration[J]. Chemistry, 2016, 22(35): 12290-12294. |
| 64 | BAEK H S, WOO B Y, YOO S J, et al. Composition containing meso-2,3-butanediol: US 10525017[P/OL]. 2020-01-07[2024-01-01]. . |
| 65 | OTAGIRI M, UI S, TAKUSAGAWA Y, et al. Structural basis for chiral substrate recognition by two 2,3-butanediol dehydrogenases[J]. FEBS Letters, 2010, 584(1): 219-223. |
| 66 | YU B, SUN J B, BOMMAREDDY R R, et al. Novel (2R,3R)-2,3-butanediol dehydrogenase from potential industrial strain Paenibacillus polymyxa ATCC 12321[J]. Applied and Environmental Microbiology, 2011, 77(12): 4230-4233. |
| 67 | LEE J W, LEE Y G, JIN Y S, et al. Metabolic engineering of non-pathogenic microorganisms for 2,3-butanediol production[J]. Applied Microbiology and Biotechnology, 2021, 105(14/15): 5751-5767. |
| 68 | MIZOBATA A, MITSUI R, YAMADA R, et al. Improvement of 2,3-butanediol tolerance in Saccharomyces cerevisiae by using a novel mutagenesis strategy[J]. Journal of Bioscience and Bioengineering, 2021, 131(3): 283-289. |
| 69 | NOVAK K, KUTSCHA R, PFLÜGL S. Microbial upgrading of acetate into 2,3-butanediol and acetoin by E. coli W[J]. Biotechnology for Biofuels, 2020, 13: 177. |
| 70 | REHMAN S, LENG L, ZHUANG H C, et al. Genomic insights to facilitate the construction of a high-xylose-utilization Enterococcus faecalis OPS2 for 2,3-BDO production[J]. Chemical Engineering Journal, 2022, 448: 137617. |
| 71 | LIN H, XU J Y, SUN W L, et al. Efficient 1-hydroxy-2-butanone production from 1,2-butanediol by whole cells of engineered E. coli [J]. Catalysts, 2021, 11(10): 1184. |
| 72 | LU H Y, DIAZ D J, CZARNECKI N J, et al. Machine learning-aided engineering of hydrolases for PET depolymerization[J]. Nature, 2022, 604(7907): 662-667. |
| 73 | JIANG S, WANG R R, WANG D H, et al. Metabolic reprogramming and biosensor-assisted mutagenesis screening for high-level production of L-arginine in Escherichia coli [J]. Metabolic Engineering, 2023, 76: 146-157. |
| 74 | LIN J L, WAGNER J M, ALPER H S. Enabling tools for high-throughput detection of metabolites: metabolic engineering and directed evolution applications[J]. Biotechnology Advances, 2017, 35(8): 950-970. |
| 75 | CHAE T U, CHOI S Y, KIM J W, et al. Recent advances in systems metabolic engineering tools and strategies[J]. Current Opinion in Biotechnology, 2017, 47: 67-82. |
| 76 | ZHANG Y W, YANG J W, YANG S Q, et al. Programming cells by multicopy chromosomal integration using CRISPR-associated transposases[J]. The CRISPR Journal, 2021, 4(3): 350-359. |
| 77 | DONG Y F, ZHANG Y, LIU D H, et al. Strain and process engineering toward continuous industrial fermentation[J]. Frontiers of Chemical Science and Engineering, 2023, 17(10): 1336-1353. |
| 78 | JIANG L L, LIU H F, MU Y, et al. High tolerance to glycerol and high production of 1,3-propanediol in batch fermentations by microbial consortium from marine sludge[J]. Engineering in Life Sciences, 2017, 17(6): 635-644. |
| 79 | LEE S Y, KIM H U, CHAE T U, et al. A comprehensive metabolic map for production of bio-based chemicals[J]. Nature Catalysis, 2019, 2: 18-33. |
| 80 | UTESCH T, SABRA W, PRESCHER C, et al. Enhanced electron transfer of different mediators for strictly opposite shifting of metabolism in Clostridium pasteurianum grown on glycerol in a new electrochemical bioreactor[J]. Biotechnology and Bioengineering, 2019, 116(7): 1627-1643. |
| [1] | 郭姝媛, 张倩楠, 姑丽克孜·买买提热夏提, 杨一群, 于涛. 液体生物燃料合成与炼制的研究进展[J]. 合成生物学, 2025, 6(1): 18-44. |
| [2] | 仲泉周, 单依怡, 裴清云, 金艳芸, 王艺涵, 孟璐远, 王歆韵, 张雨鑫, 刘坤媛, 王慧中, 冯尚国. 生物合成法生产α-熊果苷的研究进展[J]. 合成生物学, 2025, 6(1): 118-135. |
| [3] | 邵明威, 孙思勉, 杨时茂, 陈国强. 基于极端微生物的生物制造[J]. 合成生物学, 2024, 5(6): 1419-1436. |
| [4] | 赵亮, 李振帅, 付丽平, 吕明, 王士安, 张全, 刘立成, 李福利, 刘自勇. 生物转化一碳化合物原料产油脂与单细胞蛋白研究进展[J]. 合成生物学, 2024, 5(6): 1300-1318. |
| [5] | 刘益宁, 蒲伟, 杨金星, 王钰. ω-氨基酸与内酰胺的生物合成研究进展[J]. 合成生物学, 2024, 5(6): 1350-1366. |
| [6] | 李庚, 申晓林, 孙新晓, 王佳, 袁其朋. 过氧化物酶的重组表达和应用研究进展[J]. 合成生物学, 2024, 5(6): 1498-1517. |
| [7] | 郑皓天, 李朝风, 刘良叙, 王嘉伟, 李恒润, 倪俊. 负碳人工光合群落的设计、优化与应用[J]. 合成生物学, 2024, 5(5): 1189-1210. |
| [8] | 程晓雷, 刘天罡, 陶慧. 萜类化合物的非常规生物合成研究进展[J]. 合成生物学, 2024, 5(5): 1050-1071. |
| [9] | 刘子健, 穆柏杨, 段志强, 王璇, 陆晓杰. 与核酸兼容的化学反应开发进展[J]. 合成生物学, 2024, 5(5): 1102-1124. |
| [10] | 禹伟, 高教琪, 周雍进. 一碳生物转化合成有机酸的研究进展[J]. 合成生物学, 2024, 5(5): 1169-1188. |
| [11] | 张守祺, 王涛, 孔尧, 邹家胜, 刘元宁, 徐正仁. 天然产物的化学-酶法合成:方法与策略的演进[J]. 合成生物学, 2024, 5(5): 913-940. |
| [12] | 谢向前, 郭雯, 王欢, 李进. 含氨基乙烯半胱氨酸核糖体肽的生物合成与化学合成[J]. 合成生物学, 2024, 5(5): 981-996. |
| [13] | 汤志军, 胡友财, 刘文. 酶促4+2和2+2环加成反应:区域与立体选择性的理解与应用[J]. 合成生物学, 2024, 5(3): 401-407. |
| [14] | 张俊, 金诗雪, 云倩, 瞿旭东. 聚酮化合物非天然延伸单元的生物合成与结构改造应用[J]. 合成生物学, 2024, 5(3): 561-570. |
| [15] | 陈锡玮, 张华然, 邹懿. 真菌源非核糖体肽类药物生物合成及代谢工程[J]. 合成生物学, 2024, 5(3): 571-592. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
