合成生物学 ›› 2024, Vol. 5 ›› Issue (4): 700-718.DOI: 10.12211/2096-8280.2023-094
人胚胎早期发育与干细胞
艾宗勇1,2, 张成庭1,2, 牛宝华1,2, 尹宇1,2, 杨洁1, 李天晴1,2
- 1.昆明理工大学灵长类转化医学研究院,省部共建非人灵长类生物医学国家重点实验室,云南 昆明 650500
2.云南中科灵长类生物医学重点实验室,云南 昆明 650500
-
收稿日期:2023-11-30修回日期:2024-02-04出版日期:2024-08-31发布日期:2024-09-19 -
通讯作者:李天晴 -
作者简介:艾宗勇 (1984—),男,博士,副教授。研究方向为干细胞与胚胎发育。E-mail:aizy@lpbr.cn李天晴 (1975—),男,博士,教授,“国家高层次人才”特聘教授和科技部重点研发计划首席科学家。研究方向为干细胞与组织工程,从事灵长类干细胞和胚胎发育研究。E-mail:litq@lpbr.cn -
基金资助:国家自然科学基金(32360177)
Early human embryo development and stem cells
AI Zongyong1,2, ZHANG Chengting1,2, NIU Baohua1,2, YIN Yu1,2, YANG Jie1, LI Tianqing1,2
- 1.State Key Laboratory of Primate Biomedical Research,Institute of Primate Translational Medicine,Kunming University of Science and Technology,Kunming 650500,Yunnan,China
2.Yunnan Key Laboratory of Primate Biomedical Research,Kunming 650500,Yunnan,China
-
Received:2023-11-30Revised:2024-02-04Online:2024-08-31Published:2024-09-19 -
Contact:LI Tianqing
摘要:
人胚胎早期发育包括三个重要阶段:①从受精卵到晚期囊胚的着床前阶段;②从晚期囊胚到原肠运动前的围着床阶段;③从原肠运动到早期器官发生的原肠后阶段。后两个阶段统称为着床后早期发育阶段。妊娠过程中,不育(胚胎着床失败或流产)和胎儿出生缺陷,很大程度上是胚胎的着床后早期发育出现异常所致。人着床后早期胚胎,由于位于母体子宫,且尺寸较小,不易对其观察和研究,因此,这一阶段的胚胎发育过程长期处于“黑匣子”状态。近年来,随着单细胞组学技术和胚胎体外延长培养系统的建立,以及胚胎和胚外干细胞、类器官和类胚胎领域的快速发展,使得人胚胎着床后早期发育的神秘面纱被慢慢揭开。本文从人胚胎早期发育、胚胎和胚外干细胞、类胚胎和类器官研究的视角,结合细胞通信、谱系互作、信号梯度、黏附分子、生物力学和细胞外基质等因素对细胞分选、迁移重排和自我组织的影响,概述了人胚胎早期发育过程中的发育原理,当前胚胎和胚外干细胞的研究进展以及用其模拟人胚胎早期发育的研究现状、存在问题和发展方向,以期能够帮助理解人胚胎早期发育的奥秘。
中图分类号:
引用本文
艾宗勇, 张成庭, 牛宝华, 尹宇, 杨洁, 李天晴. 人胚胎早期发育与干细胞[J]. 合成生物学, 2024, 5(4): 700-718.
AI Zongyong, ZHANG Chengting, NIU Baohua, YIN Yu, YANG Jie, LI Tianqing. Early human embryo development and stem cells[J]. Synthetic Biology Journal, 2024, 5(4): 700-718.
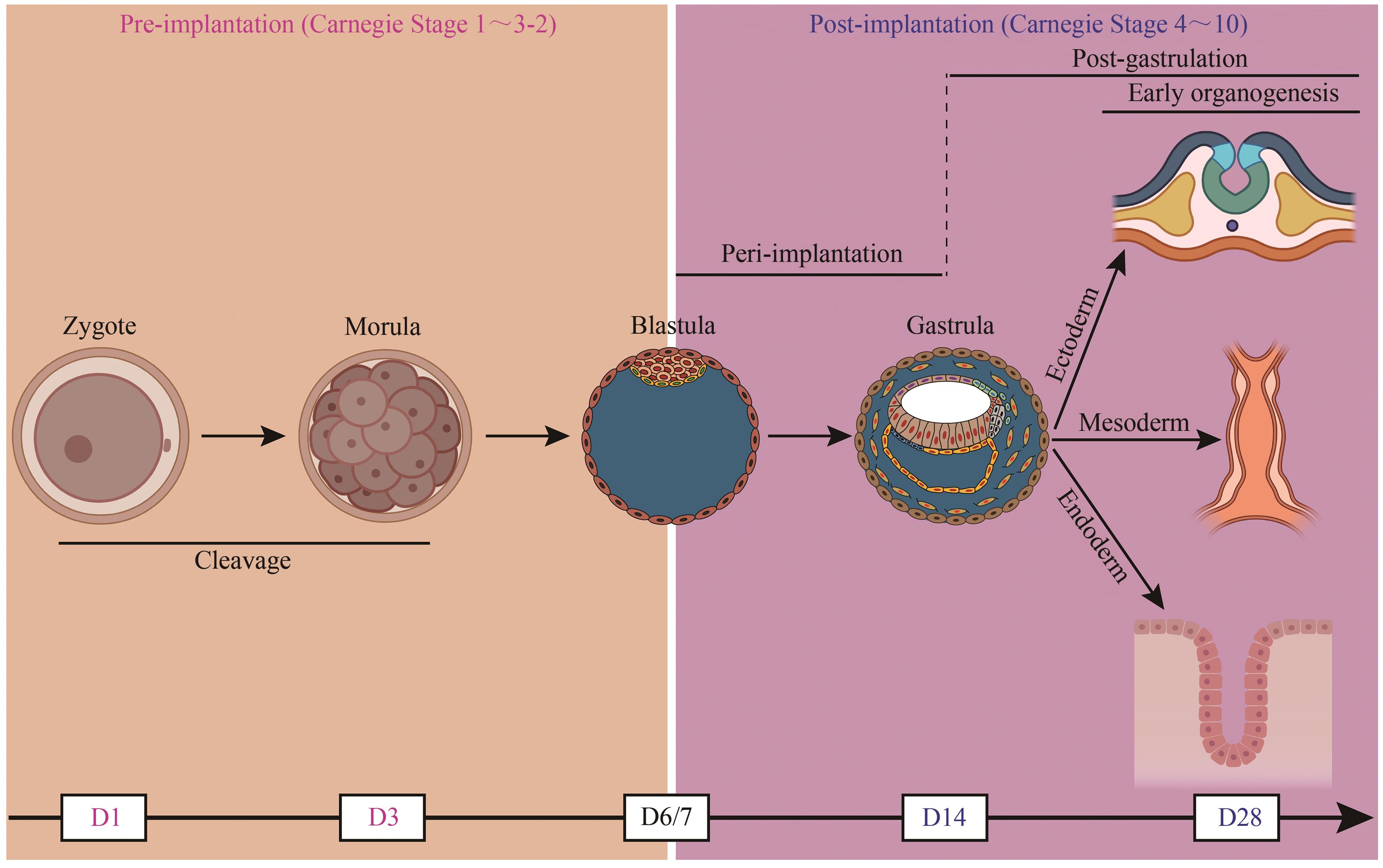
图1 人胚胎早期发育示意图(部分图形模块用BioRender制作。D—天)
Fig. 1 Schematic overview of early human embryogenesis(Some of the modules of the figure were created with BioRender. D—Day)
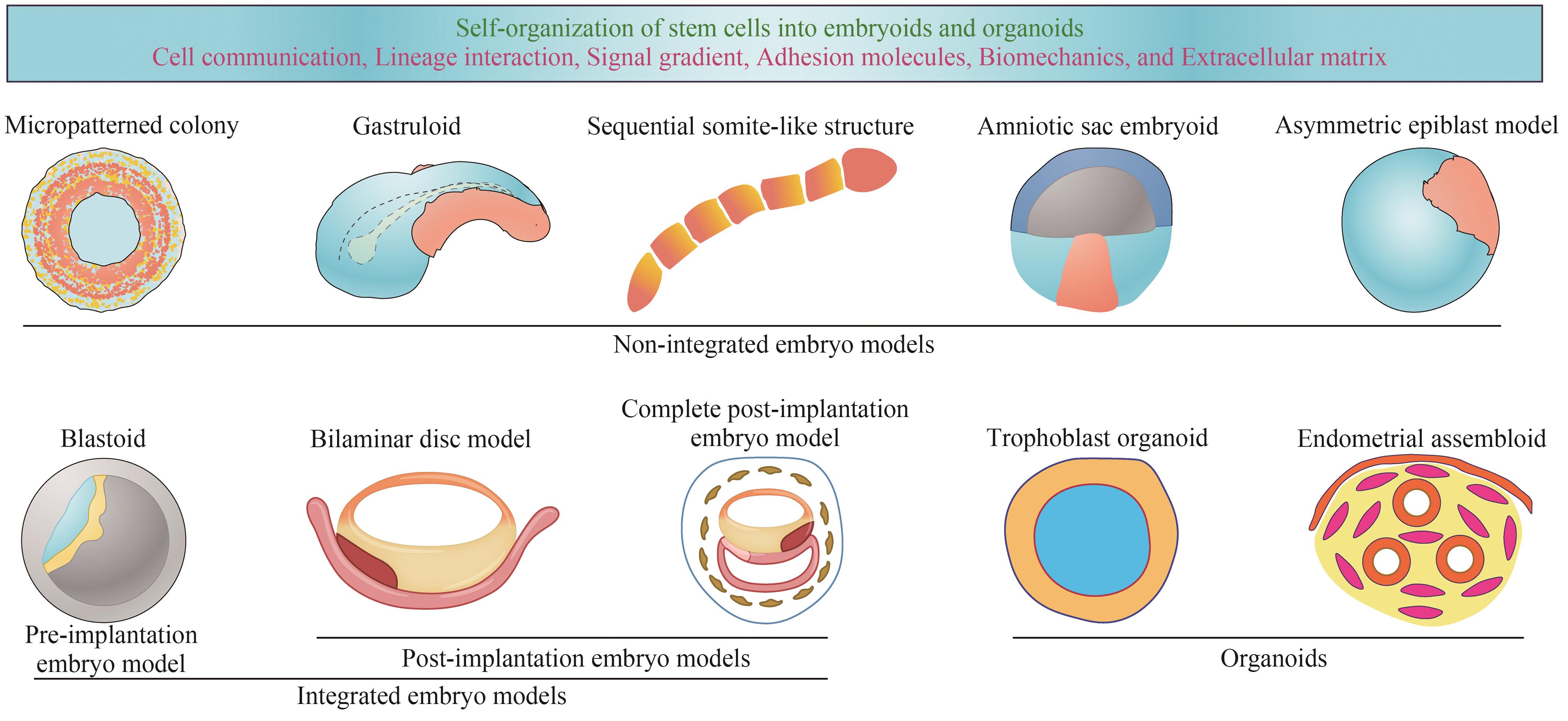
图3 人的类胚胎研究进展(在Fu等[115]和Zernicka-Goetz[116]的基础上进行改编)
Fig. 3 Research progress in human embryo models(Adapted from Fu et al.[115] and Zernicka-Goetz[116] with permission)
| 1 | MAÎTRE J L, TURLIER H, ILLUKKUMBURA R, et al. Asymmetric division of contractile domains couples cell positioning and fate specification[J]. Nature, 2016, 536(7616): 344-348. |
| 2 | JOHNSON M, ZIOMEK C A. The foundation of two distinct cell lineages within the mouse morula[J]. Cell, 1981, 24(1): 71-80. |
| 3 | JOHNSON M H, MCCONNELL J M L. Lineage allocation and cell polarity during mouse embryogenesis[J]. Seminars in Cell & Developmental Biology, 2004, 15(5): 583-597. |
| 4 | SASAKI H. Position- and polarity-dependent Hippo signaling regulates cell fates in preimplantation mouse embryos[J]. Seminars in Cell & Developmental Biology, 2015, 47-48: 80-87. |
| 5 | GERRI C, MCCARTHY A, ALANIS-LOBATO G, et al. Initiation of a conserved trophectoderm program in human, cow and mouse embryos[J]. Nature, 2020, 587(7834): 443-447. |
| 6 | ZENKER J, WHITE M D, GASNIER M, et al. Expanding actin rings zipper the mouse embryo for blastocyst formation[J]. Cell, 2018, 173(3): 776-791.e17. |
| 7 | CHAN C J, COSTANZO M, RUIZ-HERRERO T, et al. Hydraulic control of mammalian embryo size and cell fate[J]. Nature, 2019, 571(7763): 112-116. |
| 8 | ZHANG Y D, LI X, GAO S, et al. Genetic reporter for live tracing fluid flow forces during cell fate segregation in mouse blastocyst development[J]. Cell Stem Cell, 2023, 30(8): 1110-1123.e9. |
| 9 | BOROVIAK T, LOOS R, LOMBARD P, et al. Lineage-specific profiling delineates the emergence and progression of naive pluripotency in mammalian embryogenesis[J]. Developmental Cell, 2015, 35(3): 366-382. |
| 10 | LINNEBERG-AGERHOLM M, WONG Y F, ROMERO HERRERA J A, et al. Naïve human pluripotent stem cells respond to Wnt, Nodal and LIF signalling to produce expandable naïve extra-embryonic endoderm[J]. Development, 2019, 146(24): dev180620. |
| 11 | MEISTERMANN D, BRUNEAU A, LOUBERSAC S, et al. Integrated pseudotime analysis of human pre-implantation embryo single-cell transcriptomes reveals the dynamics of lineage specification[J]. Cell Stem Cell, 2021, 28(9): 1625-1640.e6. |
| 12 | YANAGIDA A, CORUJO-SIMON E, REVELL C K, et al. Cell surface fluctuations regulate early embryonic lineage sorting[J]. Cell, 2022, 185(5): 777-793.e20. |
| 13 | GELLERSEN B, BROSENS J J. Cyclic decidualization of the human endometrium in reproductive health and failure[J]. Endocrine Reviews, 2014, 35(6): 851-905. |
| 14 | RAWLINGS T M, MAKWANA K, TAYLOR D M, et al. Modelling the impact of decidual senescence on embryo implantation in human endometrial assembloids[J]. eLife, 2021, 10: e69603. |
| 15 | GARCIA-ALONSO L, HANDFIELD L F, ROBERTS K, et al. Mapping the temporal and spatial dynamics of the human endometrium in vivo and in vitro [J]. Nature Genetics, 2021, 53(12): 1698-1711. |
| 16 | WANG W X, VILELLA F, ALAMA P, et al. Single-cell transcriptomic atlas of the human endometrium during the menstrual cycle[J]. Nature Medicine, 2020, 26(10): 1644-1653. |
| 17 | RUANE P T, GARNER T, PARSONS L, et al. Trophectoderm differentiation to invasive syncytiotrophoblast is promoted by endometrial epithelial cells during human embryo implantation[J]. Human Reproduction, 2022, 37(4): 777-792. |
| 18 | HAIDER S, MEINHARDT G, SALEH L, et al. Notch1 controls development of the extravillous trophoblast lineage in the human placenta[J]. Proceedings of the National Academy of Sciences of the United States of America, 2016, 113(48): E7710-E7719. |
| 19 | KAGAWA H, JAVALI A, KHOEI H H, et al. Human blastoids model blastocyst development and implantation[J]. Nature, 2022, 601(7894): 600-605. |
| 20 | ENDERS A C, BLANKENSHIP T N, FAZLEABAS A T, et al. Structure of anchoring villi and the trophoblastic shell in the human, baboon and macaque placenta[J]. Placenta, 2001, 22(4): 284-303. |
| 21 | VENTO-TORMO R, EFREMOVA M, BOTTING R A, et al. Single-cell reconstruction of the early maternal-fetal interface in humans[J]. Nature, 2018, 563(7731): 347-353. |
| 22 | LIU Y W, FAN X Y, WANG R, et al. Single-cell RNA-seq reveals the diversity of trophoblast subtypes and patterns of differentiation in the human placenta[J]. Cell Research, 2018, 28(8): 819-832. |
| 23 | ARUTYUNYAN A, ROBERTS K, TROULÉ K, et al. Spatial multiomics map of trophoblast development in early pregnancy[J]. Nature, 2023, 616(7955): 143-151. |
| 24 | DEGLINCERTI A, CROFT G F, PIETILA L N, et al. Self-organization of the in vitro attached human embryo[J]. Nature, 2016, 533(7602): 251-254. |
| 25 | SHAHBAZI M N, JEDRUSIK A, VUORISTO S, et al. Self-organization of the human embryo in the absence of maternal tissues[J]. Nature Cell Biology, 2016, 18: 700-708. |
| 26 | XIANG L F, YIN Y, ZHENG Y, et al. A developmental landscape of 3D-cultured human pre-gastrulation embryos[J]. Nature, 2020, 577(7791): 537-542. |
| 27 | GASSER R F, CORK R J. The virtual human embryo[EB/OL].[2023-11-01]. . |
| 28 | ZHENG Y, XUE X F, SHAO Y, et al. Controlled modelling of human epiblast and amnion development using stem cells[J]. Nature, 2019, 573(7774): 421-425. |
| 29 | AI Z Y, NIU B H, YIN Y, et al. Dissecting peri-implantation development using cultured human embryos and embryo-like assembloids[J]. Cell Research, 2023, 33(9): 661-678. |
| 30 | BEDZHOV I, ZERNICKA-GOETZ M. Self-organizing properties of mouse pluripotent cells initiate morphogenesis upon implantation[J]. Cell, 2014, 156(5): 1032-1044. |
| 31 | COUCOUVANIS E, MARTIN G R. Signals for death and survival: a two-step mechanism for cavitation in the vertebrate embryo[J]. Cell, 1995, 83(2): 279-287. |
| 32 | TANIGUCHI K, SHAO Y, TOWNSHEND R F, et al. Lumen formation is an intrinsic property of isolated human pluripotent stem cells[J]. Stem Cell Reports, 2015, 5(6): 954-962. |
| 33 | SHAO Y, TANIGUCHI K, TOWNSHEND R F, et al. A pluripotent stem cell-based model for post-implantation human amniotic sac development[J]. Nature Communications, 2017, 8(1): 208. |
| 34 | SHAO Y, TANIGUCHI K, GURDZIEL K, et al. Self-organized amniogenesis by human pluripotent stem cells in a biomimetic implantation-like niche[J]. Nature Materials, 2017, 16(4): 419-425. |
| 35 | ENDERS A C, SCHLAFKE S, HENDRICKX A G. Differentiation of the embryonic disc, amnion, and yolk sac in the rhesus monkey[J]. American Journal of Anatomy, 1986, 177(2): 161-185. |
| 36 | ENDERS A C, LANTZ K C, SCHLAFKE S. Differentiation of the inner cell mass of the baboon blastocyst[J]. The Anatomical Record, 1990, 226(2): 237-248. |
| 37 | ROSS C, BOROVIAK T E. Origin and function of the yolk sac in primate embryogenesis[J]. Nature Communications, 2020, 11(1): 3760. |
| 38 | THOMAS P, BEDDINGTON R. Anterior primitive endoderm may be responsible for patterning the anterior neural plate in the mouse embryo[J]. Current Biology, 1996, 6(11): 1487-1496. |
| 39 | RIVERA-PÉREZ J A, MAGER J, MAGNUSON T. Dynamic morphogenetic events characterize the mouse visceral endoderm[J]. Developmental Biology, 2003, 261(2): 470-487. |
| 40 | MOLÈ M A, COORENS T H H, SHAHBAZI M N, et al. A single cell characterisation of human embryogenesis identifies pluripotency transitions and putative anterior hypoblast centre[J]. Nature Communications, 2021, 12(1): 3679. |
| 41 | SHAHBAZI M N, ZERNICKA-GOETZ M. Deconstructing and reconstructing the mouse and human early embryo[J]. Nature Cell Biology, 2018, 20(8): 878-887. |
| 42 | BOROVIAK T, NICHOLS J. Primate embryogenesis predicts the hallmarks of human naïve pluripotency[J]. Development, 2017, 144(2): 175-186. |
| 43 | JIANG X X, ZHAI J L, XIAO Z Y, et al. Identifying a dynamic transcriptomic landscape of the cynomolgus macaque placenta during pregnancy at single-cell resolution[J]. Developmental Cell, 2023, 58(9): 806-821.e7. |
| 44 | AI Z Y, YIN Y, NIU B H, et al. Deconstructing human peri-implantation embryogenesis based on embryos and embryoids[J]. Biology of Reproduction, 2022, 107(1): 212-225. |
| 45 | SAITOU M, BARTON S C, SURANI M A. A molecular programme for the specification of germ cell fate in mice[J]. Nature, 2002, 418(6895): 293-300. |
| 46 | OHINATA Y, PAYER B, O’CARROLL D, et al. Blimp1 is a critical determinant of the germ cell lineage in mice[J]. Nature, 2005, 436(7048): 207-213. |
| 47 | KOBAYASHI T, ZHANG H X, TANG W W C, et al. Principles of early human development and germ cell program from conserved model systems[J]. Nature, 2017, 546(7658): 416-420. |
| 48 | SASAKI K, NAKAMURA T, OKAMOTO I, et al. The germ cell fate of Cynomolgus monkeys is specified in the nascent amnion[J]. Developmental Cell, 2016, 39(2): 169-185. |
| 49 | TYSER R C V, MAHAMMADOV E, NAKANOH S, et al. Single-cell transcriptomic characterization of a gastrulating human embryo[J]. Nature, 2021, 600(7888): 285-289. |
| 50 | WITSCHI E. Migration of the germ cells of human embryos from the yolk sac to the primitive gonadal fold[J]. Contributions to Embryology, 1948, 32: 67-80. |
| 51 | CHEN D, SUN N, HOU L, et al. Human primordial germ cells are specified from lineage-primed progenitors[J]. Cell Reports, 2019, 29(13): 4568-4582.e5. |
| 52 | VICENTE C. An interview with Lewis Wolpert[J]. Development, 2015, 142(15): 2547-2548. |
| 53 | DALEY G Q, HYUN I, APPERLEY J F, et al. Setting global standards for stem cell research and clinical translation: the 2016 ISSCR guidelines[J]. Stem Cell Reports, 2016, 6(6): 787-797. |
| 54 | YANG R, GOEDEL A, KANG Y, et al. Amnion signals are essential for mesoderm formation in Primates[J]. Nature Communications, 2021, 12(1): 5126. |
| 55 | ARTZT K. Mammalian developmental genetics in the twentieth century[J]. Genetics, 2012, 192(4): 1151-1163. |
| 56 | HERTIG A T, ROCK J, ADAMS E C. A description of 34 human ova within the first 17 days of development[J]. American Journal of Anatomy, 1956, 98(3): 435-493. |
| 57 | HERTIG A T. On the development of the amnion and exoccelomic membrane in the previllous human ovum[J]. The Yale Journal of Biology and Medicine, 1945, 18: 107-115. |
| 58 | HERTIG A T. Angiogenesis in the early human chorion and in the primary placenta of the Macque monkey[J]. Contributions to Embryology, 1935, 25: 37-82. |
| 59 | BOYD J D, HAMILTON W J. The Human Placenta[M]. Combridge: Heffer, 1970, 217. |
| 60 | ZENG B, LIU Z Y, LU Y F, et al. The single-cell and spatial transcriptional landscape of human gastrulation and early brain development[J]. Cell Stem Cell, 2023, 30(6): 851-866.e7. |
| 61 | TAM P P L, LOEBEL D A F. Gene function in mouse embryogenesis: get set for gastrulation[J]. Nature Reviews Genetics, 2007, 8: 368-381. |
| 62 | YANG Y, LIU B, XU J, et al. Derivation of pluripotent stem cells with in vivo embryonic and extraembryonic potency[J]. Cell, 2017, 169(2): 243-257.e25. |
| 63 | GAO X F, NOWAK-IMIALEK M, CHEN X, et al. Establishment of porcine and human expanded potential stem cells[J]. Nature Cell Biology, 2019, 21(6): 687-699. |
| 64 | GUO G, STIRPARO G G, STRAWBRIDGE S E, et al. Human naive epiblast cells possess unrestricted lineage potential[J]. Cell Stem Cell, 2021, 28(6): 1040-1056.e6. |
| 65 | IO S, KABATA M, IEMURA Y, et al. Capturing human trophoblast development with naive pluripotent stem cells in vitro [J]. Cell Stem Cell, 2021, 28(6): 1023-1039.e13. |
| 66 | STIRPARO G G, BOROVIAK T, GUO G, et al. Integrated analysis of single-cell embryo data yields a unified transcriptome signature for the human pre-implantation epiblast[J]. Development, 2018, 145(3): dev158501. |
| 67 | GUO G, VON MEYENN F, ROSTOVSKAYA M, et al. Epigenetic resetting of human pluripotency[J]. Development, 2017, 144(15): 2748-2763. |
| 68 | MAZID M A, WARD C, LUO Z W, et al. Rolling back human pluripotent stem cells to an eight-cell embryo-like stage[J]. Nature, 2022, 605(7909): 315-324. |
| 69 | TAUBENSCHMID-STOWERS J, ROSTOVSKAYA M, SANTOS F, et al. 8C-like cells capture the human zygotic genome activation program in vitro [J]. Cell Stem Cell, 2022, 29(3): 449-459.e6. |
| 70 | MOYA-JÓDAR M, ULLATE-AGOTE A, BARLABÉ P, et al. Revealing cell populations catching the early stages of human embryo development in naive pluripotent stem cell cultures[J]. Stem Cell Reports, 2023, 18(1): 64-80. |
| 71 | YU X, LIANG S Q, CHEN M Q, et al. Recapitulating early human development with 8C-like cells[J]. Cell Reports, 2022, 39(12): 110994. |
| 72 | YOSHIHARA M, KIRJANOV I, NYKÄNEN S, et al. Transient DUX4 expression in human embryonic stem cells induces blastomere-like expression program that is marked by SLC34A2[J]. Stem Cell Reports, 2022, 17(7): 1743-1756. |
| 73 | YOSHIHARA M, KERE J. Transcriptomic differences between human 8-cell-like cells reprogrammed with different methods[J]. Stem Cell Reports, 2023, 18(8): 1621-1628. |
| 74 | THOMSON J A, ITSKOVITZ-ELDOR J, SHAPIRO S S, et al. Embryonic stem cell lines derived from human blastocysts[J]. Science, 1998, 282(5391): 1145-1147. |
| 75 | TAKAHASHI K, TANABE K, OHNUKI M, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors[J]. Cell, 2007, 131(5): 861-872. |
| 76 | YU J Y, VODYANIK M A, SMUGA-OTTO K, et al. Induced pluripotent stem cell lines derived from human somatic cells[J]. Science, 2007, 318(5858): 1917-1920. |
| 77 | GAFNI O, WEINBERGER L, MANSOUR A A, et al. Derivation of novel human ground state naive pluripotent stem cells[J]. Nature, 2013, 504: 282-286. |
| 78 | AI Z Y, NIU B H, DUAN K, et al. Modulation of Wnt and Activin/Nodal supports efficient derivation, cloning and suspension expansion of human pluripotent stem cells[J]. Biomaterials, 2020, 249: 120015. |
| 79 | CHAN Y S, GÖKE J, NG J H, et al. Induction of a human pluripotent state with distinct regulatory circuitry that resembles preimplantation epiblast[J]. Cell Stem Cell, 2013, 13(6): 663-675. |
| 80 | NICHOLS J, SMITH A. Naive and primed pluripotent states[J]. Cell Stem Cell, 2009, 4(6): 487-492. |
| 81 | SMITH A. Formative pluripotency: the executive phase in a developmental continuum[J]. Development, 2017, 144(3): 365-373. |
| 82 | TAKASHIMA Y, GUO G, LOOS R, et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human[J]. Cell, 2014, 158(6): 1254-1269. |
| 83 | THEUNISSEN T W, POWELL B E, WANG H Y, et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency[J]. Cell Stem Cell, 2014, 15(4): 471-487. |
| 84 | BREDENKAMP N, YANG J, CLARKE J, et al. Wnt inhibition facilitates RNA-mediated reprogramming of human somatic cells to naive pluripotency[J]. Stem Cell Reports, 2019, 13(6): 1083-1098. |
| 85 | THEUNISSEN T W, FRIEDLI M, HE Y P, et al. Molecular criteria for defining the naive human pluripotent state[J]. Cell Stem Cell, 2016, 19(4): 502-515. |
| 86 | BAYERL J, AYYASH M, SHANI T, et al. Principles of signaling pathway modulation for enhancing human naive pluripotency induction[J]. Cell Stem Cell, 2021, 28(9): 1549-1565.e12. |
| 87 | ROSTOVSKAYA M, STIRPARO G G, SMITH A. Capacitation of human naïve pluripotent stem cells for multi-lineage differentiation[J]. Development, 2019, 146(7): dev172916. |
| 88 | DI STEFANO B, UEDA M, SABRI S, et al. Reduced MEK inhibition preserves genomic stability in naive human embryonic stem cells[J]. Nature Methods, 2018, 15(9): 732-740. |
| 89 | KALKAN T, SMITH A. Mapping the route from naive pluripotency to lineage specification[J]. Philosophical Transactions of the Royal Society of London Series B, Biological Sciences, 2014, 369(1657): 20130540. |
| 90 | HAYASHI K, OHTA H, KURIMOTO K, et al. Reconstitution of the mouse germ cell specification pathway in culture by pluripotent stem cells[J]. Cell, 2011, 146(4): 519-532. |
| 91 | KINOSHITA M, BARBER M, MANSFIELD W, et al. Capture of mouse and human stem cells with features of formative pluripotency[J]. Cell Stem Cell, 2021, 28(3): 453-471.e8. |
| 92 | YU L Q, WEI Y L, SUN H X, et al. Derivation of intermediate pluripotent stem cells amenable to primordial germ cell specification[J]. Cell Stem Cell, 2021, 28(3): 550-567.e12. |
| 93 | KILENS S, MEISTERMANN D, MORENO D, et al. Parallel derivation of isogenic human primed and naive induced pluripotent stem cells[J]. Nature Communications, 2018, 9(1): 360. |
| 94 | AMITA M, ADACHI K, ALEXENKO A P, et al. Complete and unidirectional conversion of human embryonic stem cells to trophoblast by BMP4[J]. Proceedings of the National Academy of Sciences of the United States of America, 2013, 110(13): E1212-E1221. |
| 95 | XU R H, CHEN X, LI D S, et al. BMP4 initiates human embryonic stem cell differentiation to trophoblast[J]. Nature Biotechnology, 2002, 20(12): 1261-1264. |
| 96 | WU Z, ZHANG W, CHEN G B, et al. Combinatorial signals of activin/nodal and bone morphogenic protein regulate the early lineage segregation of human embryonic stem cells[J]. Journal of Biological Chemistry, 2008, 283(36): 24991-25002. |
| 97 | CASTEL G, MEISTERMANN D, BRETIN B, et al. Induction of human trophoblast stem cells from somatic cells and pluripotent stem cells[J]. Cell Reports, 2020, 33(8): 108419. |
| 98 | BERNARDO A S, FAIAL T, GARDNER L, et al. BRACHYURY and CDX2 mediate BMP-induced differentiation of human and mouse pluripotent stem cells into embryonic and extraembryonic lineages[J]. Cell Stem Cell, 2011, 9(2): 144-155. |
| 99 | CHHABRA S, WARMFLASH A. BMP-treated human embryonic stem cells transcriptionally resemble amnion cells in the monkey embryo[J]. Biology Open, 2021, 10(9): bio058617. |
| 100 | KUNATH T, YAMANAKA Y, DETMAR J, et al. Developmental differences in the expression of FGF receptors between human and mouse embryos[J]. Placenta, 2014, 35(12): 1079-1088. |
| 101 | SONCIN F, NATALE D, PARAST M M. Signaling pathways in mouse and human trophoblast differentiation: a comparative review[J]. Cellular and Molecular Life Sciences, 2015, 72(7): 1291-1302. |
| 102 | OKAE H, TOH H, SATO T, et al. Derivation of human trophoblast stem cells[J]. Cell Stem Cell, 2018, 22(1): 50-63.e6. |
| 103 | DONG C, BELTCHEVA M, GONTARZ P, et al. Derivation of trophoblast stem cells from naïve human pluripotent stem cells[J]. eLife, 2020, 9: e52504. |
| 104 | LIU X D, OUYANG J F, ROSSELLO F J, et al. Reprogramming roadmap reveals route to human induced trophoblast stem cells[J]. Nature, 2020, 586(7827): 101-107. |
| 105 | CINKORNPUMIN J K, KWON S Y, GUO Y X, et al. Naive human embryonic stem cells can give rise to cells with a trophoblast-like transcriptome and methylome[J]. Stem Cell Reports, 2020, 15(1): 198-213. |
| 106 | WEI Y X, WANG T Y, MA L S, et al. Efficient derivation of human trophoblast stem cells from primed pluripotent stem cells[J]. Science Advances, 2021, 7(33): eabf4416. |
| 107 | SONCIN F, MOREY R, BUI T, et al. Derivation of functional trophoblast stem cells from primed human pluripotent stem cells[J]. Stem Cell Reports, 2022, 17(6): 1303-1317. |
| 108 | VIUKOV S, SHANI T, BAYERL J, et al. Human primed and naïve PSCs are both able to differentiate into trophoblast stem cells[J]. Stem Cell Reports, 2022, 17(11): 2484-2500. |
| 109 | ZORZAN I, BETTO R M, ROSSIGNOLI G, et al. Chemical conversion of human conventional PSCs to TSCs following transient naive gene activation[J]. EMBO Reports, 2023, 24(4): e55235. |
| 110 | SEETHARAM A S, VU H T H, CHOI S, et al. The product of BMP-directed differentiation protocols for human primed pluripotent stem cells is placental trophoblast and not amnion[J]. Stem Cell Reports, 2022, 17(6): 1289-1302. |
| 111 | MACKINLAY K M, WEATHERBEE B A, SOUZA ROSA V, et al. An in vitro stem cell model of human epiblast and yolk sac interaction[J]. eLife, 2021, 10: e63930. |
| 112 | PHAM T X A, PANDA A, KAGAWA H, et al. Modeling human extraembryonic mesoderm cells using naive pluripotent stem cells[J]. Cell Stem Cell, 2022, 29(9): 1346-1365.e10. |
| 113 | WEI Y L, ZHANG E, YU L Q, et al. Dissecting embryonic and extraembryonic lineage crosstalk with stem cell co-culture[J]. Cell, 2023, 186(26): 5859-5875.e24. |
| 114 | OKUBO T, RIVRON N, KABATA M, et al. Hypoblast from human pluripotent stem cells regulates epiblast development[J]. Nature, 2024, 626: 357-366. |
| 115 | FU J P, WARMFLASH A, LUTOLF M P. Stem-cell-based embryo models for fundamental research and translation[J]. Nature Materials, 2021, 20(2): 132-144. |
| 116 | ZERNICKA-GOETZ M. The evolution of embryo models[J]. Nature Methods, 2023, 20(12): 1844-1848. |
| 117 | MARTYN I, KANNO T Y, RUZO A, et al. Self-organization of a human organizer by combined Wnt and Nodal signalling[J]. Nature, 2018, 558: 132-135. |
| 118 | SIMUNOVIC M, METZGER J J, ETOC F, et al. A 3D model of a human epiblast reveals BMP4-driven symmetry breaking[J]. Nature Cell Biology, 2019, 21(7): 900-910. |
| 119 | SIMUNOVIC M, SIGGIA E D, BRIVANLOU A H. In vitro attachment and symmetry breaking of a human embryo model assembled from primed embryonic stem cells[J]. Cell Stem Cell, 2022, 29(6): 962-972.e4. |
| 120 | YU L Q, LOGSDON D, PINZON-ARTEAGA C A, et al. Large-scale production of human blastoids amenable to modeling blastocyst development and maternal-fetal cross talk[J]. Cell Stem Cell, 2023, 30(9): 1246-1261.e9. |
| 121 | SPEMANN H, MANGOLD H. Induction of embryonic primordia by implantation of organizers from a different species. 1923[J]. The International Journal of Developmental Biology, 2001, 45(1): 13-38. |
| 122 | TURING A M. The chemical basis of morphogenesis. 1953[J]. Bulletin of Mathematical Biology, 1990, 52(1/2): 153-197. |
| 123 | WOLPERT L. Positional information and the spatial pattern of cellular differentiation[J]. Journal of Theoretical Biology, 1969, 25(1): 1-47. |
| 124 | YAMANAKA Y, HAMIDI S, YOSHIOKA-KOBAYASHI K, et al. Reconstituting human somitogenesis in vitro [J]. Nature, 2023, 614(7948): 509-520. |
| 125 | MIAO Y C, DJEFFAL Y, SIMONE A D, et al. Reconstruction and deconstruction of human somitogenesis in vitro [J]. Nature, 2023, 614(7948): 500-508. |
| 126 | OLDAK B, WILDSCHUTZ E, BONDARENKO V, et al. Complete human day 14 post-implantation embryo models from naive ES cells[J]. Nature, 2023, 622(7983): 562-573. |
| 127 | WEATHERBEE B A T, GANTNER C W, IWAMOTO-STOHL L K, et al. Pluripotent stem cell-derived model of the post-implantation human embryo[J]. Nature, 2023, 622(7983): 584-593. |
| 128 | TSAI T Y C, SIKORA M, XIA P, et al. An adhesion code ensures robust pattern formation during tissue morphogenesis[J]. Science, 2020, 370(6512): 113-116. |
| 129 | STEINBERG M S. Does differential adhesion govern self-assembly processes in histogenesis? Equilibrium configurations and the emergence of a hierarchy among populations of embryonic cells[J]. The Journal of Experimental Zoology, 1970, 173(4): 395-433. |
| 130 | BAO M, CORNWALL-SCOONES J, SANCHEZ-VASQUEZ E, et al. Stem cell-derived synthetic embryos self-assemble by exploiting cadherin codes and cortical tension[J]. Nature Cell Biology, 2022, 24(9): 1341-1349. |
| 131 | WEBERLING A, ZERNICKA-GOETZ M. Trophectoderm mechanics direct epiblast shape upon embryo implantation[J]. Cell Reports, 2021, 34(3): 108655. |
| 132 | LIU L Z, OURA S, MARKHAM Z, et al. Modeling post-implantation stages of human development into early organogenesis with stem-cell-derived peri-gastruloids[J]. Cell, 2023, 186(18): 3776-3792.e16. |
| 133 | TIAN J W, YANG J, CHEN T W, et al. Generation of human endometrial assembloids with a luminal epithelium using air-liquid interface culture methods[J]. Advanced Science, 2023, 10(30): e2301868. |
| 134 | TURCO M Y, GARDNER L, KAY R G, et al. Trophoblast organoids as a model for maternal-fetal interactions during human placentation[J]. Nature, 2018, 564(7735): 263-267. |
| 135 | HAIDER S, MEINHARDT G, SALEH L, et al. Self-renewing trophoblast organoids recapitulate the developmental program of the early human placenta[J]. Stem Cell Reports, 2018, 11(2): 537-551. |
| 136 | YANG L H, LIANG P F, YANG H H, et al. Trophoblast organoids with physiological polarity model placental structure and function[J]. Journal of Cell Science, 2024, 137(5): jcs261528. |
| 137 | ZHOU J, SHERIDAN M A, TIAN Y, et al. Development of properly-polarized trophoblast stem cell-derived organoids to model early human pregnancy[EB/OL]. BioRxiv, 2023: 2023.09.30.560327[2023-12-01]. . |
| 138 | ZHOU F, WANG R, YUAN P, et al. Reconstituting the transcriptome and DNA methylome landscapes of human implantation[J]. Nature, 2019, 572(7771): 660-664. |
| 139 | LV B, AN Q, ZENG Q, et al. Single-cell RNA sequencing reveals regulatory mechanism for trophoblast cell-fate divergence in human peri-implantation conceptuses[J]. PLoS Biology, 2019, 17(10): e3000187. |
| 140 | WEST R C, MING H, LOGSDON D M, et al. Dynamics of trophoblast differentiation in peri-implantation–stage human embryos[J]. Proceedings of the National Academy of Sciences of the United States of America, 2019, 116(45): 22635-22644. |
| 141 | SHAHBAZI M N, WANG T R, TAO X, et al. Developmental potential of aneuploid human embryos cultured beyond implantation[J]. Nature Communications, 2020, 11(1): 3987. |
| 142 | SHAHBAZI M N, SCIALDONE A, SKORUPSKA N, et al. Pluripotent state transitions coordinate morphogenesis in mouse and human embryos[J]. Nature, 2017, 552(7684): 239-243. |
| 143 | PEDROZA M, GASSALOGLU S I, DIAS N, et al. Self-patterning of human stem cells into post-implantation lineages[J]. Nature, 2023, 622(7983): 574-583. |
| 144 | KARVAS R M, ZEMKE J E, ALI S S, et al. 3D-cultured blastoids model human embryogenesis from pre-implantation to early gastrulation stages[J]. Cell Stem Cell, 2023, 30(9): 1148-1165.e7. |
| 145 | HISLOP J, SONG Q, KESHAVARZ F K, et al. Modelling post-implantation human development to yolk sac blood emergence[J]. Nature, 2024, 626: 367-376. |
| 146 | YUAN G G, WANG J C, LIU Z D, et al. Establishment of a novel non-integrated human pluripotent stem cell-based gastruloid model[EB/OL]. bioRxiv,2023: 2023. 06. 28.546720.[2023-12-01]. . |
| 147 | YANAGIDA A, SPINDLOW D, NICHOLS J, et al. Naive stem cell blastocyst model captures human embryo lineage segregation[J]. Cell Stem Cell, 2021, 28(6): 1016-1022.e4. |
| 148 | AMADEI G, HANDFORD C E, QIU C X, et al. Embryo model completes gastrulation to neurulation and organogenesis[J]. Nature, 2022, 610(7930): 143-153. |
| 149 | TARAZI S, AGUILERA-CASTREJON A, JOUBRAN C, et al. Post-gastrulation synthetic embryos generated ex utero from mouse naive ESCs[J]. Cell, 2022, 185(18): 3290-3306.e25. |
| 150 | LAU K Y C, RUBINSTEIN H, GANTNER C W, et al. Mouse embryo model derived exclusively from embryonic stem cells undergoes neurulation and heart development[J]. Cell Stem Cell, 2022, 29(10): 1445-1458.e8. |
| [1] | 陈汐玥, 王亚清, 包芳, 秦建华. 肝器官芯片在生物医学研究中的应用进展[J]. 合成生物学, 2024, 5(4): 813-830. |
| [2] | 曹荣凯, 秦建华, 王亚清. 胎盘芯片及其在生殖医学领域的研究进展[J]. 合成生物学, 2024, 5(4): 831-850. |
| [3] | 洪源, 刘妍. 脑类器官在再生医学中的研究进展[J]. 合成生物学, 2024, 5(4): 754-769. |
| [4] | 陈倩文, 赵思琪, 彭耀进. 类器官:技术创新与伦理争议[J]. 合成生物学, 2024, 5(4): 898-907. |
| [5] | 胡博文, 陈家斌, 刘晓东. 人类早期胚胎发育体外模型研究进展[J]. 合成生物学, 2024, 5(4): 719-733. |
| [6] | 蔡冰玉, 谭象天, 李伟. 合成生物学在干细胞工程化改造中的研究进展[J]. 合成生物学, 2024, 5(4): 782-794. |
| [7] | 张博航, 祁晓萱, 袁艳. 睾丸类器官在体外精子发生中的研究进展[J]. 合成生物学, 2024, 5(4): 770-781. |
| [8] | 韩宜钊, 郭佳, 邵玥. 干细胞模拟发育:细胞元件、胚胎模型与工程方法[J]. 合成生物学, 2024, 5(4): 734-753. |
| [9] | 李石开, 曾东鳌, 杜方舟, 张京钟, 余爽. 血管化类器官的构建方法及生物材料[J]. 合成生物学, 2024, 5(4): 851-866. |
| [10] | 胡可儿, 王汉奇, 黄儒麒, 张灿阳, 邢新会, 马少华. 整合设计策略下的工程化类器官与类器官芯片技术[J]. 合成生物学, 2024, 5(4): 883-897. |
| [11] | 陈子苓, 向阳飞. 类器官技术与合成生物学协同研究进展[J]. 合成生物学, 2024, 5(4): 795-812. |
| [12] | 孟倩, 尹聪, 黄卫人. 肿瘤类器官及其在合成生物学中的研究进展[J]. 合成生物学, 2024, 5(1): 191-201. |
| [13] | 朱骊宇, 赵玉龙, 李伟, 王立宾. 哺乳动物染色体工程研究进展[J]. 合成生物学, 2023, 4(2): 394-406. |
| [14] | 张璨, 施李杨, 戴建武. 细胞培养肉用生物材料的设计[J]. 合成生物学, 2022, 3(4): 676-689. |
| [15] | 宋成治, 孙阳, 曹毅. 力信号在干细胞命运决定过程中的影响[J]. 合成生物学, 2022, 3(4): 781-794. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||