合成生物学 ›› 2023, Vol. 4 ›› Issue (1): 5-29.DOI: 10.12211/2096-8280.2022-038
蛋白质稳定性计算设计与定向进化前沿工具
阮青云1, 黄莘1, 孟子钧1, 全舒1,2
- 1.华东理工大学生物工程学院,生物反应器工程国家重点实验室,上海生物制造技术协同创新中心,上海 200237
2.上海市细胞代谢光遗传学技术前沿科学研究基地,上海 200237
-
收稿日期:2022-07-02修回日期:2022-07-30出版日期:2023-02-28发布日期:2023-03-07 -
通讯作者:全舒 -
作者简介:阮青云(1996—),男,博士研究生。研究方向为蛋白质稳定性检测探针的开发与应用。阮青云 (1996—),男,博士研究生。研究方向为蛋白质稳定性检测探针的开发与应用。 E-mail: alessandroruan@mail.ecust.edu.cn全舒 (1982—),女,教授,博士生导师。课题组聚焦于蛋白质折叠领域的工具开发与机制解析,发展了一系列蛋白质体内稳定性检测探针,建立了以分子伴侣活力改造为基础的蛋白质折叠调控策略,为基础研究及蛋白质在生物催化、生物医药等领域的应用提供了支撑 E-mail: shuquan@ecust.edu.cn -
基金资助:国家自然科学基金面上项目(31870054)
Computational design and directed evolution strategies for optimizing protein stability
RUAN Qingyun1, HUANG Xin1, MENG Zijun1, QUAN Shu1,2
- 1.State Key Laboratory of Bioreactor Engineering,School of Biotechnology,East China University of Science and Technology,Shanghai Collaborative Innovation Center for Biomanufacturing(SCICB),Shanghai 200237,China
2.Shanghai Frontiers Science Center of Optogenetic Techniques for Cell Metabolism,East China University of Science and Technology,Shanghai 200237,China
-
Received:2022-07-02Revised:2022-07-30Online:2023-02-28Published:2023-03-07 -
Contact:QUAN Shu
摘要:
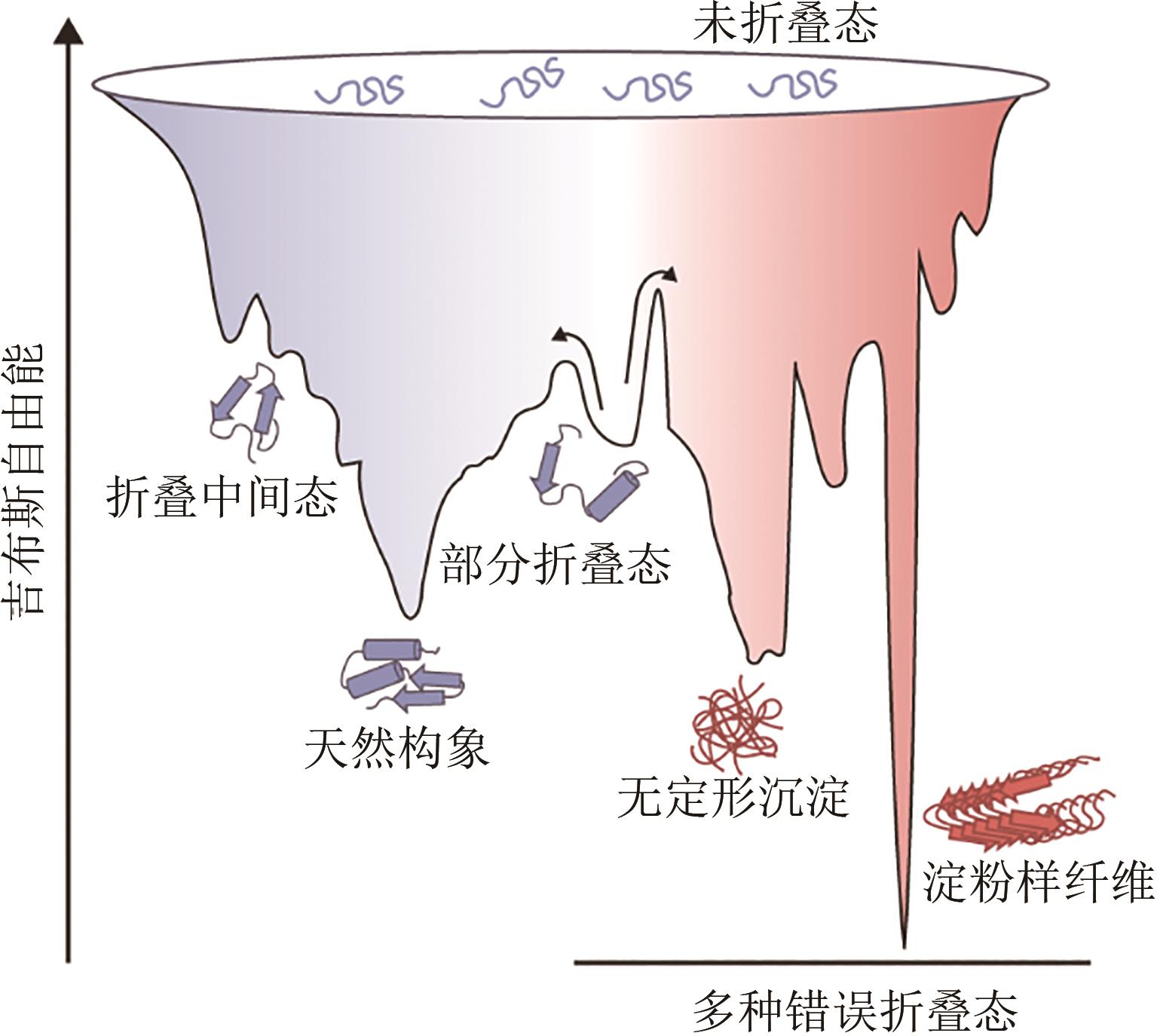
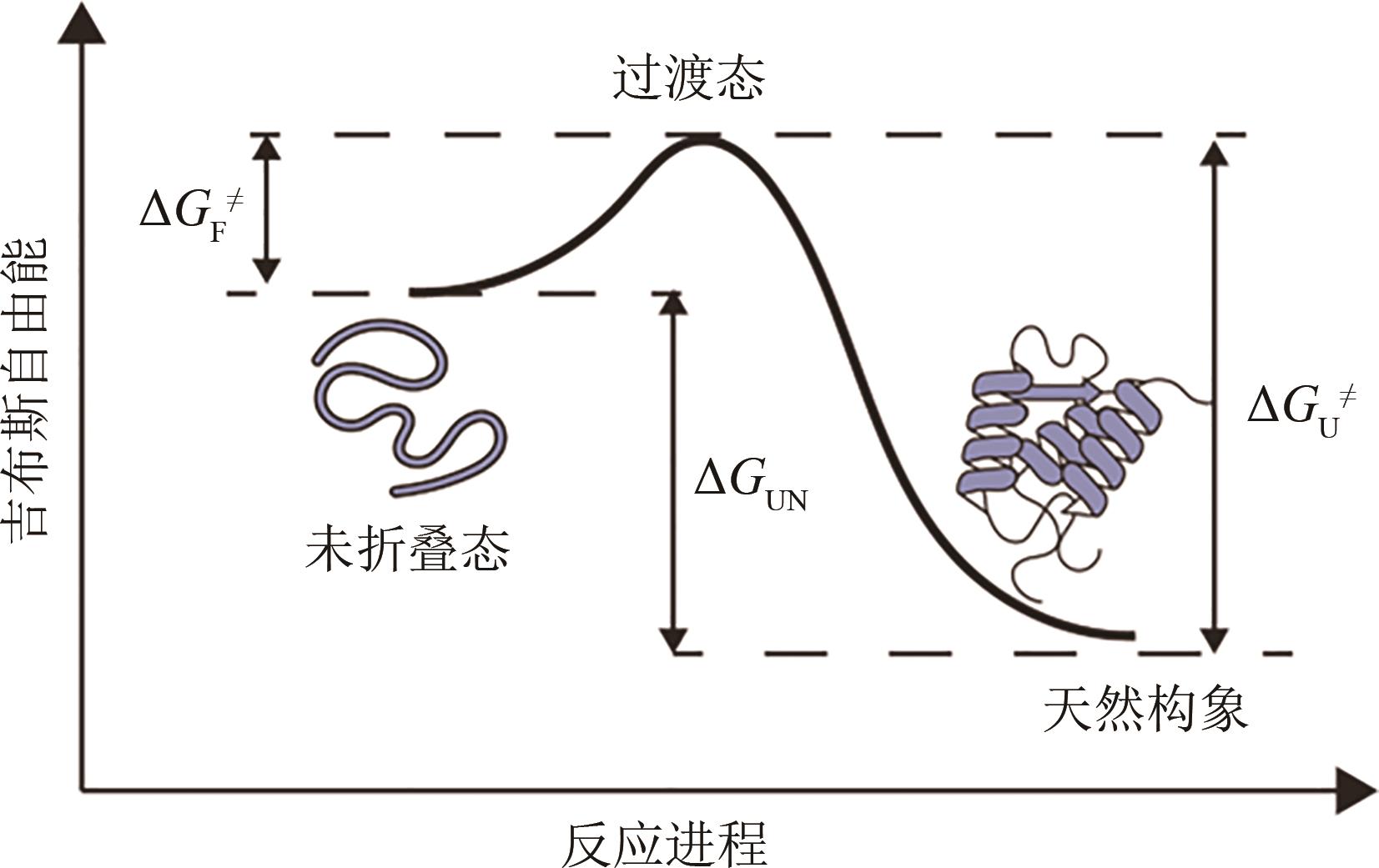
天然蛋白质具有临界稳定性的特征,这种较低的稳定性使蛋白质结构具有足够的灵活性,从而支持其发挥生物学功能。然而,临界稳定性使得蛋白质遭受胁迫压力后极易发生错误折叠并失去功能,导致天然蛋白质往往无法满足科学研究与工业应用的需求。此外,体内蛋白质在错误折叠后产生的聚集沉淀被认为是多种疾病发生发展的原因,包括阿尔兹海默病、帕金森综合征等。因此,优化蛋白质的稳定性是科学研究与工程应用领域亟待解决的关键问题。本文从蛋白质的折叠与稳定性机制出发,聚焦于序列优化与折叠环境优化两种改善蛋白质稳定性的手段,综述了基于理性设计、计算机辅助设计改善蛋白质稳定性的研究方法,介绍了用于高通量筛选蛋白质稳定化突变体或折叠相关因子的定向进化技术。通过多项蛋白质序列改良、折叠环境优化的案例介绍,展示了蛋白质稳定化技术在蛋白质工程与生物医药领域的广阔应用,包括酶的稳定化设计、疫苗蛋白质的构象控制、分子伴侣与蛋白质聚集抑制剂的筛选、蛋白质稳态药物的开发等。最后,展望了蛋白质稳定化技术未来的研究方向与前景,定制化的蛋白质稳定性检测技术将会迎来蓬勃发展。
中图分类号:
引用本文
阮青云, 黄莘, 孟子钧, 全舒. 蛋白质稳定性计算设计与定向进化前沿工具[J]. 合成生物学, 2023, 4(1): 5-29.
RUAN Qingyun, HUANG Xin, MENG Zijun, QUAN Shu. Computational design and directed evolution strategies for optimizing protein stability[J]. Synthetic Biology Journal, 2023, 4(1): 5-29.
| 符号 | 稳定性类型 | 度量 | 定义 |
|---|---|---|---|
| ΔGUN | 热力学 | 折叠自由能 | 蛋白质未折叠状态到天然构象的吉布斯自由能变化 |
| ΔGNU | 热力学 | 去折叠自由能 | 蛋白质天然构象到未折叠状态的吉布斯自由能变化 |
| ΔΔG | 热力学 | 折叠自由能变化 | 蛋白质突变前后折叠自由能的变化 |
| Tm | 热力学 | 熔解温度 | 使一半的蛋白质解折叠时的温度 |
| C1/2 | 热力学 | 半变性浓度 | 使一半的蛋白质解折叠时的变性剂浓度 |
| KU | 热力学 | 解折叠平衡常数 | 未折叠状态与天然状态的浓度比值 |
| kf | 动力学 | 折叠速率常数 | 蛋白质折叠过程的速率常数 |
| ku | 动力学 | 去折叠速率常数 | 蛋白质去折叠过程的速率常数 |
| kd, obs | 动力学 | 表观失活速率常数 | 从天然状态到完全失活(deactivation)的表观速率常数 |
| T50 | ― | 半数失活温度 | 在一定时间内酶活降至一半时的温度 |
| t1/2 | 动力学 | 半衰期 | 酶活降至初始的一半时所需的时间 |
表1 不同表征蛋白质稳定性参数的定义
Table 1 Definitions of stability parameters
| 符号 | 稳定性类型 | 度量 | 定义 |
|---|---|---|---|
| ΔGUN | 热力学 | 折叠自由能 | 蛋白质未折叠状态到天然构象的吉布斯自由能变化 |
| ΔGNU | 热力学 | 去折叠自由能 | 蛋白质天然构象到未折叠状态的吉布斯自由能变化 |
| ΔΔG | 热力学 | 折叠自由能变化 | 蛋白质突变前后折叠自由能的变化 |
| Tm | 热力学 | 熔解温度 | 使一半的蛋白质解折叠时的温度 |
| C1/2 | 热力学 | 半变性浓度 | 使一半的蛋白质解折叠时的变性剂浓度 |
| KU | 热力学 | 解折叠平衡常数 | 未折叠状态与天然状态的浓度比值 |
| kf | 动力学 | 折叠速率常数 | 蛋白质折叠过程的速率常数 |
| ku | 动力学 | 去折叠速率常数 | 蛋白质去折叠过程的速率常数 |
| kd, obs | 动力学 | 表观失活速率常数 | 从天然状态到完全失活(deactivation)的表观速率常数 |
| T50 | ― | 半数失活温度 | 在一定时间内酶活降至一半时的温度 |
| t1/2 | 动力学 | 半衰期 | 酶活降至初始的一半时所需的时间 |
| 类别 | 软件名称 | 输入 | 网站/本地安装 | 参考文献 |
|---|---|---|---|---|
| 基于结构分析 | ||||
| 优化蛋白质表面电荷 | TKSA-MC | 结构 | http://tksamc.df.ibilce.unesp.br | [ |
| PHEPS | 结构 | http://pheps.orgchm.bas.bg/home.html | [ | |
| 基于温度因子 | B-FITTER | 结构 | Windows系统本地安装 | [ |
| 设计二硫键 | Disulfide By Design | 结构 | http://cptweb.cpt.wayne.edu/DbD2 | [ |
| DISULFIDE | 结构 | http://disulfind.disi.unitn.it | [ | |
| 破坏表面大面积疏水区域 | AGGRESCAN3D | 结构/序列 | http://biocomp.chem.uw.edu.pl/A3D2 | [ |
| 基于进化分析 | ||||
| 同源序列比对 | 3DM | 序列 | https://3dm.bio-prodict.com | [ |
| Consensus Finder | 序列 | http://kazlab.umn.edu | [ | |
| 祖先酶重构 | Ancestors 1.0 | 序列 | http://ancestors.bioinfo.uqam.ca/ancestorWeb | [ |
| FireProt ASR | 序列 | https://loschmidt.chemi.muni.cz/fireprotasr | [ | |
| 基于折叠自由能计算 | ||||
| 单纯ΔΔG计算 | FoldX | 结构 | 全平台本地安装 | [ |
| Rosetta | 结构 | Linux系统本地安装 | [ | |
| PoPMuSiC | 结构 | http://babylone.ulb.ac.be/popmusic | [ | |
| 组合其他策略的ΔΔG计算 | PROSS | 结构 | http://pross.weizmann.ac.il/step/pross-terms | [ |
| FireProt | 结构/序列 | https://loschmidt.chemi.muni.cz/fireprot | [ | |
| 基于机器学习 | ||||
| 支持向量机 | I-mutant | 结构/序列 | http://gpcr.biocomp.unibo.it/cgi/predictors/I-Mutant2.0/I-Mutant2.0.cgi | [ |
| SRP神经网络 | DeepDDG | 结构 | http://protein.org.cn/ddg.html | [ |
| 卷积神经网络 | MutCompute | 结构 | https://mutcompute.com | [ |
表2 理性设计方法的代表性软件[31-33, 38-40, 43, 45, 47-56]
Table 2 Summary of software for protein stability rational design
| 类别 | 软件名称 | 输入 | 网站/本地安装 | 参考文献 |
|---|---|---|---|---|
| 基于结构分析 | ||||
| 优化蛋白质表面电荷 | TKSA-MC | 结构 | http://tksamc.df.ibilce.unesp.br | [ |
| PHEPS | 结构 | http://pheps.orgchm.bas.bg/home.html | [ | |
| 基于温度因子 | B-FITTER | 结构 | Windows系统本地安装 | [ |
| 设计二硫键 | Disulfide By Design | 结构 | http://cptweb.cpt.wayne.edu/DbD2 | [ |
| DISULFIDE | 结构 | http://disulfind.disi.unitn.it | [ | |
| 破坏表面大面积疏水区域 | AGGRESCAN3D | 结构/序列 | http://biocomp.chem.uw.edu.pl/A3D2 | [ |
| 基于进化分析 | ||||
| 同源序列比对 | 3DM | 序列 | https://3dm.bio-prodict.com | [ |
| Consensus Finder | 序列 | http://kazlab.umn.edu | [ | |
| 祖先酶重构 | Ancestors 1.0 | 序列 | http://ancestors.bioinfo.uqam.ca/ancestorWeb | [ |
| FireProt ASR | 序列 | https://loschmidt.chemi.muni.cz/fireprotasr | [ | |
| 基于折叠自由能计算 | ||||
| 单纯ΔΔG计算 | FoldX | 结构 | 全平台本地安装 | [ |
| Rosetta | 结构 | Linux系统本地安装 | [ | |
| PoPMuSiC | 结构 | http://babylone.ulb.ac.be/popmusic | [ | |
| 组合其他策略的ΔΔG计算 | PROSS | 结构 | http://pross.weizmann.ac.il/step/pross-terms | [ |
| FireProt | 结构/序列 | https://loschmidt.chemi.muni.cz/fireprot | [ | |
| 基于机器学习 | ||||
| 支持向量机 | I-mutant | 结构/序列 | http://gpcr.biocomp.unibo.it/cgi/predictors/I-Mutant2.0/I-Mutant2.0.cgi | [ |
| SRP神经网络 | DeepDDG | 结构 | http://protein.org.cn/ddg.html | [ |
| 卷积神经网络 | MutCompute | 结构 | https://mutcompute.com | [ |
| 1 | DILL K A, MACCALLUM J L. The protein-folding problem, 50 years on[J]. Science, 2012, 338(6110): 1042-1046. |
| 2 | KIM Y E, HIPP M S, BRACHER A, et al. Molecular chaperone functions in protein folding and proteostasis[J]. Annual Review of Biochemistry, 2013, 82: 323-355. |
| 3 | BALCHIN D, HAYER-HARTL M, HARTL F U. In vivo aspects of protein folding and quality control[J]. Science, 2016, 353(6294): aac4354. |
| 4 | HARTL F U, BRACHER A, HAYER-HARTL M. Molecular chaperones in protein folding and proteostasis[J]. Nature, 2011, 475(7356): 324-332. |
| 5 | HARTL F U, HAYER-HARTL M. Converging concepts of protein folding in vitro and in vivo [J]. Nature Structural & Molecular Biology, 2009, 16(6): 574-581. |
| 6 | TRAXLMAYR M W, OBINGER C. Directed evolution of proteins for increased stability and expression using yeast display[J]. Archives of Biochemistry and Biophysics, 2012, 526(2): 174-180. |
| 7 | SANCHEZ-RUIZ J M. Protein kinetic stability[J]. Biophysical Chemistry, 2010, 148(1/2/3): 1-15. |
| 8 | POLIZZI K M, BOMMARIUS A S, BROERING J M, et al. Stability of biocatalysts[J]. Current Opinion in Chemical Biology, 2007, 11(2): 220-225. |
| 9 | TAVERNA D M, GOLDSTEIN R A. Why are proteins marginally stable? [J]. Proteins: Structure, Function, and Genetics, 2002, 46(1): 105-109. |
| 10 | 唐宇琦, 叶松涛, 刘嘉, 等. 分子伴侣作用下的蛋白质稳定与进化[J]. 合成生物学, 2022, 3(3): 445-464. |
| TANG Y Q, YE S T, LIU J, et al. Molecular chaperones promote protein stability and evolution[J]. Synthetic Biology Journal, 2022, 3(3): 445-464. | |
| 11 | BENILOVA I, KARRAN E, DE STROOPER B. The toxic Aβ oligomer and Alzheimer's disease: an emperor in need of clothes[J]. Nature Neuroscience, 2012, 15(3): 349-357. |
| 12 | SPILLANTINI M G, GOEDERT M. Tau pathology and neurodegeneration[J]. The Lancet Neurology, 2013, 12(6): 609-622. |
| 13 | SINGLETON A B, FARRER M, JOHNSON J, et al. α-Synuclein locus triplication causes Parkinson's disease[J]. Science, 2003, 302(5646): 841. |
| 14 | ZRAIKA S, HULL R L, VERCHERE C B, et al. Toxic oligomers and islet beta cell death: guilty by association or convicted by circumstantial evidence? [J]. Diabetologia, 2010, 53(6): 1046-1056. |
| 15 | COELHO T, MERLINI G, BULAWA C E, et al. Mechanism of action and clinical application of tafamidis in hereditary transthyretin amyloidosis[J]. Neurology and Therapy, 2016, 5(1): 1-25. |
| 16 | BULAWA C E, CONNELLY S, DEVIT M, et al. Tafamidis, a potent and selective transthyretin kinetic stabilizer that inhibits the amyloid cascade[J]. Proceedings of the National Academy of Sciences of the United States of America, 2012, 109(24): 9629-9634. |
| 17 | HIROSE M, KURODA Y, MURATA E. NGF/TrkA signaling as a therapeutic target for pain[J]. Pain Practice, 2016, 16(2): 175-182. |
| 18 | EBO J S, SAUNDERS J C, DEVINE P W A, et al. An in vivo platform to select and evolve aggregation-resistant proteins[J]. Nature Communications, 2020, 11: 1816. |
| 19 | SETIAWAN D, BRENDER J, ZHANG Y. Recent advances in automated protein design and its future challenges[J]. Expert Opinion on Drug Discovery, 2018, 13(7): 587-604. |
| 20 | GOIHBERG E, DYM O, TEL-OR S, et al. A single proline substitution is critical for the thermostabilization of Clostridium beijerinckii alcohol dehydrogenase[J]. Proteins: Structure, Function, and Bioinformatics, 2007, 66(1): 196-204. |
| 21 | STERNKE M, TRIPP K W, BARRICK D. Consensus sequence design as a general strategy to create hyperstable, biologically active proteins[J]. Proceedings of the National Academy of Sciences of the United States of America, 2019, 116(23): 11275-11284. |
| 22 | GUMULYA Y, BAEK J M, WUN S J, et al. Engineering highly functional thermostable proteins using ancestral sequence reconstruction[J]. Nature Catalysis, 2018, 1(11): 878-888. |
| 23 | WANG C H, HUANG R B, HE B F, et al. Improving the thermostability of alpha-amylase by combinatorial coevolving-site saturation mutagenesis[J]. BMC Bioinformatics, 2012, 13: 263. |
| 24 | STRICKLER S S, GRIBENKO A V, GRIBENKO A V, et al. Protein stability and surface electrostatics: a charged relationship[J]. Biochemistry, 2006, 45(9): 2761-2766. |
| 25 | CHAN C H, WILBANKS C C, MAKHATADZE G I, et al. Electrostatic contribution of surface charge residues to the stability of a thermophilic protein: benchmarking experimental and predicted pKa values[J]. PLoS One, 2012, 7(1): e30296. |
| 26 | LAWRENCE M S, PHILLIPS K J, LIU D R. Supercharging proteins can impart unusual resilience[J]. Journal of the American Chemical Society, 2007, 129(33): 10110-10112. |
| 27 | REETZ M T, CARBALLEIRA J D, VOGEL A. Iterative saturation mutagenesis on the basis of B factors as a strategy for increasing protein thermostability[J]. Angewandte Chemie International Edition, 2006, 45(46): 7745-7751. |
| 28 | YU H R, HUANG H. Engineering proteins for thermostability through rigidifying flexible sites[J]. Biotechnology Advances, 2014, 32(2): 308-315. |
| 29 | ZHANG C, SAMAD M, YU H R, et al. Computational design to reduce conformational flexibility and aggregation rates of an antibody fab fragment[J]. Molecular Pharmaceutics, 2018, 15(8): 3079-3092. |
| 30 | FANG L, CHOW K M, HOU S R, et al. Rational design, preparation, and characterization of a therapeutic enzyme mutant with improved stability and function for cocaine detoxification[J]. ACS Chemical Biology, 2014, 9(8): 1764-1772. |
| 31 | GIL-GARCIA M, BAÑÓ-POLO M, VAREJÃO N, et al. Combining structural aggregation propensity and stability predictions to redesign protein solubility[J]. Molecular Pharmaceutics, 2018, 15(9): 3846-3859. |
| 32 | DELGADO J, RADUSKY L G, CIANFERONI D, et al. FoldX 5.0: working with RNA, small molecules and a new graphical interface[J]. Bioinformatics, 2019, 35(20): 4168-4169. |
| 33 | SIMONS K T, BONNEAU R, RUCZINSKI I, et al. Ab initio protein structure prediction of CASP III targets using ROSETTA[J]. Proteins: Structure, Function, and Genetics, 1999(S3): 171-176. |
| 34 | GUEROIS R, NIELSEN J E, SERRANO L. Predicting changes in the stability of proteins and protein complexes: a study of more than 1000 mutations[J]. Journal of Molecular Biology, 2002, 320(2): 369-387. |
| 35 | BUß O, MULLER D, JAGER S, et al. Improvement in the thermostability of a β-amino acid converting ω-transaminase by using FoldX[J]. ChemBioChem, 2018, 19(4): 379-387. |
| 36 | YU H R, HERNÁNDEZ LÓPEZ R I, STEADMAN D, et al. Engineering transketolase to accept both unnatural donor and acceptor substrates and produce α-hydroxyketones[J]. The FEBS Journal, 2020, 287(9): 1758-1776. |
| 37 | WIJMA H J, FLOOR R J, JEKEL P A, et al. Computationally designed libraries for rapid enzyme stabilization[J]. Protein Engineering, Design and Selection, 2014, 27(2): 49-58. |
| 38 | GOLDENZWEIG A, GOLDSMITH M, HILL S E, et al. Automated structure- and sequence-based design of proteins for high bacterial expression and stability[J]. Molecular Cell, 2016, 63(2): 337-346. |
| 39 | MUSIL M, STOURAC J, BENDL J, et al. FireProt: web server for automated design of thermostable proteins[J]. Nucleic Acids Research, 2017, 45(W1): W393-W399. |
| 40 | SHROFF R, COLE A W, DIAZ D J, et al. Discovery of novel gain-of-function mutations guided by structure-based deep learning[J]. ACS Synthetic Biology, 2020, 9(11): 2927-2935. |
| 41 | LU H Y, DIAZ D J, CZARNECKI N J, et al. Machine learning-aided engineering of hydrolases for PET depolymerization[J]. Nature, 2022, 604(7907): 662-667. |
| 42 | BAVA K A, GROMIHA M M, UEDAIRA H, et al. ProTherm, version 4.0: thermodynamic database for proteins and mutants[J]. Nucleic Acids Research, 2004, 32(): D120-D121. |
| 43 | CAPRIOTTI E, FARISELLI P, CASADIO R. I-Mutant2.0: predicting stability changes upon mutation from the protein sequence or structure[J]. Nucleic Acids Research, 2005, 33(): W306-W310. |
| 44 | RASHID P M A, SALIH G F. Molecular and computational analysis of spike protein of newly emerged omicron variant in comparison to the delta variant of SARS-CoV-2 in Iraq[J]. Molecular Biology Reports, 2022, 49(8): 7437-7445. |
| 45 | CAO H L, WANG J X, HE L P, et al. DeepDDG: predicting the stability change of protein point mutations using neural networks[J]. Journal of Chemical Information and Modeling, 2019, 59(4): 1508-1514. |
| 46 | ROMERO P A, KRAUSE A, ARNOLD F H. Navigating the protein fitness landscape with Gaussian processes[J]. Proceedings of the National Academy of Sciences of the United States of America, 2013, 110(3): E193-E201. |
| 47 | CONTESSOTO V G, DE OLIVEIRA V M, FERNANDES B R, et al. TKSA-MC: A web server for rational mutation through the optimization of protein charge interactions[J]. Proteins: Structure, Function, and Bioinformatics, 2018, 86(11): 1184-1188. |
| 48 | KANTARDJIEV A A, ATANASOV B P. PHEPS: web-based pH-dependent protein electrostatics server[J]. Nucleic Acids Research, 2006, 34(): W43-W47. |
| 49 | REETZ M T, SONI P, FERNÁNDEZ L, et al. Increasing the stability of an enzyme toward hostile organic solvents by directed evolution based on iterative saturation mutagenesis using the B-FIT method[J]. Chemical Communications, 2010, 46(45): 8657-8658. |
| 50 | CRAIG D B, DOMBKOWSKI A A. Disulfide by Design 2.0: a web-based tool for disulfide engineering in proteins[J]. BMC Bioinformatics, 2013, 14: 346. |
| 51 | CERONI A, PASSERINI A, VULLO A, et al. DISULFIND: a disulfide bonding state and cysteine connectivity prediction server[J]. Nucleic Acids Research, 2006, 34(): W177-W181. |
| 52 | KUIPERS R K, JOOSTEN H J, VAN BERKEL W J H, et al. 3DM: systematic analysis of heterogeneous superfamily data to discover protein functionalities[J]. Proteins: Structure, Function, and Bioinformatics, 2010, 78(9): 2101-2113. |
| 53 | JONES B J, KAN C N E, LUO C, et al. Consensus Finder web tool to predict stabilizing substitutions in proteins[J]. Methods in Enzymology, 2020, 643: 129-148. |
| 54 | DIALLO A B, MAKARENKOV V, BLANCHETTE M. Ancestors 1.0: a web server for ancestral sequence reconstruction[J]. Bioinformatics, 2010, 26(1): 130-131. |
| 55 | MUSIL M, KHAN R T, BEIER A, et al. FireProtASR: a web server for fully automated ancestral sequence reconstruction[J]. Briefings in Bioinformatics, 2021, 22(4): bbaa337. |
| 56 | DEHOUCK Y, KWASIGROCH J M, GILIS D, et al. PoPMuSiC 2.1: a web server for the estimation of protein stability changes upon mutation and sequence optimality[J]. BMC Bioinformatics, 2011, 12: 151. |
| 57 | KOWALSKI J M, PAREKH R N, MAO J, et al. Protein folding stability can determine the efficiency of escape from endoplasmic reticulum quality control[J]. Journal of Biological Chemistry, 1998, 273(31): 19453-19458. |
| 58 | COUGHLAN C M, WALKER J L, COCHRAN J C, et al. Degradation of mutated bovine pancreatic trypsin inhibitor in the yeast vacuole suggests post-endoplasmic reticulum protein quality control[J]. Journal of Biological Chemistry, 2004, 279(15): 15289-15297. |
| 59 | SHUSTA E V, KIEKE M C, PARKE E, et al. Yeast polypeptide fusion surface display levels predict thermal stability and soluble secretion efficiency[J]. Journal of Molecular Biology, 1999, 292(5): 949-956. |
| 60 | KIEKE M C, SHUSTA E V, BODER E T, et al. Selection of functional T cell receptor mutants from a yeast surface-display library[J]. Proceedings of the National Academy of Sciences of the United States of America, 1999, 96(10): 5651-5656. |
| 61 | TRAXLMAYR M W, FAISSNER M, STADLMAYR G, et al. Directed evolution of stabilized IgG1-Fc scaffolds by application of strong heat shock to libraries displayed on yeast[J]. Biochimica et Biophysica Acta (BBA)-Proteins and Proteomics, 2012, 1824(4): 542-549. |
| 62 | ORR B A, CARR L M, WITTRUP K D, et al. Rapid method for measuring ScFv thermal stability by yeast surface display[J]. Biotechnology Progress, 2003, 19(2): 631-638. |
| 63 | HAGIHARA Y, KIM P S. Toward development of a screen to identify randomly encoded, foldable sequences[J]. Proceedings of the National Academy of Sciences of the United States of America, 2002, 99(10): 6619-6624. |
| 64 | SMITH G P. Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface[J]. Science, 1985, 228(4705): 1315-1317. |
| 65 | PACKER M S, LIU D R. Methods for the directed evolution of proteins[J]. Nature Reviews Genetics, 2015, 16(7): 379-394. |
| 66 | KRISTENSEN P, WINTER G. Proteolytic selection for protein folding using filamentous bacteriophages[J]. Folding and Design, 1998, 3(5): 321-328. |
| 67 | SIEBER V, PLÜCKTHUN A, SCHMID F X. Selecting proteins with improved stability by a phage-based method[J]. Nature Biotechnology, 1998, 16(10): 955-960. |
| 68 | MARTIN A, SCHMID F X, SIEBER V. Proside: A phage-based method for selecting thermostable proteins[M]//Directed enzyme evolution. New Jersey: Humana Press, 2003: 57-70. |
| 69 | KATHER I, JAKOB R P, DOBBEK H, et al. Increased folding stability of TEM-1 beta-lactamase by in vitro selection[J]. Journal of Molecular Biology, 2008, 383(1): 238-251. |
| 70 | PERSHAD K, KAY B K. Generating thermal stable variants of protein domains through phage display[J]. Methods, 2013, 60(1): 38-45. |
| 71 | ESVELT K M, CARLSON J C, LIU D R. A system for the continuous directed evolution of biomolecules[J]. Nature, 2011, 472(7344): 499-503. |
| 72 | WANG T N, BADRAN A H, HUANG T P, et al. Continuous directed evolution of proteins with improved soluble expression[J]. Nature Chemical Biology, 2018, 14(10): 972-980. |
| 73 | 刘陈立, 赖旺生, 陈茜, 等. 一种可视的连续空间定向进化方法: CN109971693A[P]. 2019-07-05. |
| LIU C L, LAI W S, CHEN Q, et al. Visual continuous space directional evolution method: CN109971693A[P]. 2019-07-05. | |
| 74 | PAN J G, KIM E J, YUN C H. Bacillus spore display[J]. Trends in Biotechnology, 2012, 30(12): 610-612. |
| 75 | ZHANG G Y, AN Y F, ZABED H, et al. Bacillus subtilis spore surface display technology: a review of its development and applications[J]. Journal of Microbiology and Biotechnology, 2019, 29(2): 179-190. |
| 76 | SHENG S L, JIA H, TOPIOL S, et al. Engineering CotA laccase for acidic pH stability using Bacillus subtilis spore display[J]. Journal of Microbiology and Biotechnology, 2017, 27(3): 507-513. |
| 77 | REN C, WEN X, MENCIUS J, et al. Selection and screening strategies in directed evolution to improve protein stability[J]. Bioresources and Bioprocessing, 2019, 6: 53. |
| 78 | JERMUTUS L, HONEGGER A, SCHWESINGER F, et al. Tailoring in vitro evolution for protein affinity or stability[J]. Proceedings of the National Academy of Sciences of the United States of America, 2001, 98(1): 75-80. |
| 79 | NEMOTO N, MIYAMOTO-SATO E, HUSIMI Y, et al. In vitro virus bonding of mRNA bearing puromycin at the 3 terminal [J]. Federation of European Biochemical Societies, 1997, 414(2): 405-408. |
| 80 | UENO S, NEMOTO N. cDNA display: rapid stabilization of mRNA display[J]. Methods in Molecular Biology, 2012, 805: 113-135. |
| 81 | KURZ M, GU K, AL-GAWARI A, et al. cDNA-protein fusions: covalent protein-gene conjugates for the in vitro selection of peptides and proteins[J]. Chembiochem: a European Journal of Chemical Biology, 2001, 2(9): 666-672. |
| 82 | WALDO G S, STANDISH B M, BERENDZEN J, et al. Rapid protein-folding assay using green fluorescent protein[J]. Nature Biotechnology, 1999, 17(7): 691-695. |
| 83 | KAWASAKI M, INAGAKI F. Random PCR-based screening for soluble domains using green fluorescent protein[J]. Biochemical and Biophysical Research Communications, 2001, 280(3): 842-844. |
| 84 | VAN DEN BERG S, LOFDAHL P A, HARD T, et al. Improved solubility of TEV protease by directed evolution[J]. Journal of Biotechnology, 2006, 121(3): 291-298. |
| 85 | BÉHAR G, SOLÉ V, DEFONTAINE A, et al. Evolution of interleukin-15 for higher E. coli expression and solubility[J]. Protein Engineering, Design and Selection, 2011, 24(3): 283-290. |
| 86 | CHAUTARD H, BLAS-GALINDO E, MENGUY T, et al. An activity-independent selection system of thermostable protein variants[J]. Nature Methods, 2007, 4(11): 919-921. |
| 87 | LIU J W, BOUCHER Y, STOKES H W, et al. Improving protein solubility: the use of the Escherichia coli dihydrofolate reductase gene as a fusion reporter[J]. Protein Expression and Purification, 2006, 47(1): 258-263. |
| 88 | CABANTOUS S, TERWILLIGER T C, WALDO G S. Protein tagging and detection with engineered self-assembling fragments of green fluorescent protein[J]. Nature Biotechnology, 2005, 23(1): 102-107. |
| 89 | RODRÍGUEZ-BANQUERI A, ERRASTI-MURUGARREN E, BARTOCCIONI P, et al. Stabilization of a prokaryotic LAT transporter by random mutagenesis[J]. Journal of General Physiology, 2016, 147(4): 353-368. |
| 90 | WIGLEY W C, STIDHAM R D, SMITH N M, et al. Protein solubility and folding monitored in vivo by structural complementation of a genetic marker protein[J]. Nature Biotechnology, 2001, 19(2): 131-136. |
| 91 | CABANTOUS S, PÉDELACQ J D, MARK B L, et al. Recent advances in GFP folding reporter and split-GFP solubility reporter technologies. application to improving the folding and solubility of recalcitrant proteins from Mycobacterium tuberculosis [J]. Journal of Structural and Functional Genomics, 2005, 6(2/3): 113-119. |
| 92 | NAGUMO Y, FUJIWARA K, HORISAWA K, et al. PURE mRNA display for in vitro selection of single-chain antibodies[J]. The Journal of Biochemistry, 2016, 159(5): 519-526. |
| 93 | FOIT L, MORGAN G J, KERN M J, et al. Optimizing protein stability in vivo [J]. Molecular Cell, 2009, 36(5): 861-871. |
| 94 | QUAN S, KOLDEWEY P, TAPLEY T, et al. Genetic selection designed to stabilize proteins uncovers a chaperone called Spy[J]. Nature Structural & Molecular Biology, 2011, 18(3): 262-269. |
| 95 | SACHSENHAUSER V, DENG X X, KIM H H, et al. Yeast tripartite biosensors sensitive to protein stability and aggregation propensity[J]. ACS Chemical Biology, 2020, 15(4): 1078-1088. |
| 96 | MALIK A, MUELLER-SCHICKERT A, BARDWELL J C A. Cytosolic selection systems to study protein stability[J]. Journal of Bacteriology, 2014, 196(24): 4333-4343. |
| 97 | PITTMAN A M C, LAGE M D, POLTORATSKY V, et al. Rapid profiling of disease alleles using a tunable reporter of protein misfolding[J]. Genetics, 2012, 192(3): 831-842. |
| 98 | REN C, WEN X, MENCIUS J, et al. An enzyme-based biosensor for monitoring and engineering protein stability in vivo [J]. Proceedings of the National Academy of Sciences of the United States of America, 2021, 118(13): e2101618118. |
| 99 | FISHER A C, KIM W, DELISA M P. Genetic selection for protein solubility enabled by the folding quality control feature of the twin-arginine translocation pathway[J]. Protein Science, 2006, 15(3): 449-458. |
| 100 | BOOCK J T, KING B C, TAW M N, et al. Repurposing a bacterial quality control mechanism to enhance enzyme production in living cells[J]. Journal of Molecular Biology, 2015, 427(6): 1451-1463. |
| 101 | KIM D S, SONG H N, NAM H J, et al. Directed evolution of human heavy chain variable domain (VH) using in vivo protein fitness filter[J]. PLoS One, 2014, 9(6): e98178. |
| 102 | WARAHO D, DELISA M P. Versatile selection technology for intracellular protein-protein interactions mediated by a unique bacterial hitchhiker transport mechanism[J]. Proceedings of the National Academy of Sciences of the United States of America, 2009, 106(10): 3692-3697. |
| 103 | TAW M N, LI M J, KIM D, et al. Engineering a supersecreting strain of Escherichia coli by directed coevolution of the multiprotein Tat translocation machinery[J]. ACS Synthetic Biology, 2021, 10(11): 2947-2958. |
| 104 | RICHTER K, HASLBECK M, BUCHNER J, et al. The heat shock response: life on the verge of death[J]. Molecular Cell, 2010, 40(2): 253-266. |
| 105 | LIM B, MIYAZAKI R, NEHER S, et al. Heat shock transcription factor σ32 co-opts the signal recognition particle to regulate protein homeostasis in E. coli [J]. PLoS Biology, 2013, 11(12): e1001735. |
| 106 | ZUTZ A, HAMBORG L, PEDERSEN L E, et al. A dual-reporter system for investigating and optimizing protein translation and folding in E. coli [J]. Nature Communications, 2021, 12: 6093. |
| 107 | ROMERO-SUAREZ D, WULFF T, RONG Y X, et al. A reporter system for cytosolic protein aggregates in yeast[J]. ACS Synthetic Biology, 2021, 10(3): 466-477. |
| 108 | CORNVIK T, DAHLROTH S L, MAGNUSDOTTIR A, et al. Colony filtration blot: a new screening method for soluble protein expression in Escherichia coli [J]. Nature Methods, 2005, 2(7): 507-509. |
| 109 | ASIAL I, CHENG Y X, ENGMAN H, et al. Engineering protein thermostability using a generic activity-independent biophysical screen inside the cell[J]. Nature Communications, 2013, 4: 2901. |
| 110 | RILEY T P, CHOU H T, HU R Z, et al. Enhancing the prefusion conformational stability of SARS-CoV-2 spike protein through structure-guided design[J]. Frontiers in Immunology, 2021, 12: 660198. |
| 111 | SANDERS R W, VESANEN M, SCHUELKE N, et al. Stabilization of the soluble, cleaved, trimeric form of the envelope glycoprotein complex of human immunodeficiency virus type 1[J]. Journal of Virology, 2002, 76(17): 8875-8889. |
| 112 | KIRCHDOERFER R N, WANG N S, PALLESEN J, et al. Stabilized coronavirus spikes are resistant to conformational changes induced by receptor recognition or proteolysis[J]. Scientific Reports, 2018, 8: 15701. |
| 113 | GRAHAM B S, GILMAN M S A, MCLELLAN J S. Structure-based vaccine antigen design[J]. Annual Review of Medicine, 2019, 70: 91-104. |
| 114 | PALLESEN J, WANG N, CORBETT K S, et al. Immunogenicity and structures of a rationally designed prefusion MERS-CoV spike antigen[J]. Proceedings of the National Academy of Sciences of the United States of America, 2017, 114(35): E7348-E7357. |
| 115 | CORBETT K S, EDWARDS D K, LEIST S R, et al. SARS-CoV-2 mRNA vaccine design enabled by prototype pathogen preparedness[J]. Nature, 2020, 586(7830): 567-571. |
| 116 | KHOURY D S, CROMER D, REYNALDI A, et al. Neutralizing antibody levels are highly predictive of immune protection from symptomatic SARS-CoV-2 infection[J]. Nature Medicine, 2021, 27(7): 1205-1211. |
| 117 | XIONG X L, QU K, CIAZYNSKA K A, et al. A thermostable, closed SARS-CoV-2 spike protein trimer[J]. Nature Structural & Molecular Biology, 2020, 27(10): 934-941. |
| 118 | HENDERSON R, EDWARDS R J, MANSOURI K, et al. Controlling the SARS-CoV-2 spike glycoprotein conformation[J]. Nature Structural & Molecular Biology, 2020, 27(10): 925-933. |
| 119 | QUAN S, BARDWELL J C A. Chaperone discovery[J]. BioEssays, 2012, 34(11): 973-981. |
| 120 | HE W, ZHANG J Y, SACHSENHAUSER V, et al. Increased surface charge in the protein chaperone Spy enhances its anti-aggregation activity[J]. Journal of Biological Chemistry, 2020, 295(42): 14488-14500. |
| 121 | QUAN S, WANG L L, PETROTCHENKO E V, et al. Super Spy variants implicate flexibility in chaperone action[J]. eLife, 2014, 3: e01584. |
| 122 | CORTEZ L, SIM V. The therapeutic potential of chemical chaperones in protein folding diseases[J]. Prion, 2014, 8(2): 197-202. |
| 123 | HAILU T T, FOIT L, BARDWELL J C A, et al. In vivo detection and quantification of chemicals that enhance protein stability[J]. Analytical Biochemistry, 2013, 434(1): 181-186. |
| 124 | WANG Z, SNPs MOULT J., structure protein, and disease [J]. Human Mutation, 2001, 17(4): 263-270. |
| 125 | SAUNDERS J C, YOUNG L M, MAHOOD R A, et al. An in vivo platform for identifying inhibitors of protein aggregation[J]. Nature Chemical Biology, 2016, 12(2): 94-101. |
| 126 | CHEN S, WU J L, LIANG Y, et al. Arsenic trioxide rescues structural p53 mutations through a cryptic allosteric site[J]. Cancer Cell, 2021, 39(2): 225-239.e8. |
| 127 | QI S M, DONG J Y, XU Z Y, et al. PROTAC: an effective targeted protein degradation strategy for cancer therapy[J]. Frontiers in Pharmacology, 2021, 12: 692574. |
| 128 | RAMADAS B, KUMAR PAIN P, MANNA D. LYTACs: an emerging tool for the degradation of non-cytosolic proteins[J]. ChemMedChem, 2021, 16(19): 2951-2953. |
| 129 | LU M C, LIU T, JIAO Q, et al. Discovery of a Keap1-dependent peptide PROTAC to knockdown Tau by ubiquitination-proteasome degradation pathway[J]. European Journal of Medicinal Chemistry, 2018, 146: 251-259. |
| 130 | HENNING N J, BOIKE L, SPRADLIN J N, et al. Deubiquitinase-targeting chimeras for targeted protein stabilization[J]. Nature Chemical Biology, 2022, 18(4): 412-421. |
| 131 | TEH S Y, LIN R, HUNG L H, et al. Droplet microfluidics [J]. Lab on a Chip, 2008, 8(2): 198-220. |
| [1] | 付雨, 钟芳锐. 化学原理驱动的光生物不对称催化研究进展[J]. 合成生物学, 2024, 5(5): 1021-1049. |
| [2] | 汤志军, 胡友财, 刘文. 酶促4+2和2+2环加成反应:区域与立体选择性的理解与应用[J]. 合成生物学, 2024, 5(3): 401-407. |
| [3] | 孙梦楚, 陆亮宇, 申晓林, 孙新晓, 王佳, 袁其朋. 基于荧光检测的高通量筛选技术和装备助力细胞工厂构建[J]. 合成生物学, 2023, 4(5): 947-965. |
| [4] | 明阳, 陈彬, 黄小强. 光酶催化合成进展[J]. 合成生物学, 2023, 4(4): 651-675. |
| [5] | 康里奇, 谈攀, 洪亮. 人工智能时代下的酶工程[J]. 合成生物学, 2023, 4(3): 524-534. |
| [6] | 陈青黎, 童贻刚. 工程噬菌体的合成生物学“智造”[J]. 合成生物学, 2023, 4(2): 283-300. |
| [7] | 祁延萍, 朱晋, 张凯, 刘彤, 王雅婕. 定向进化在蛋白质工程中的应用研究进展[J]. 合成生物学, 2022, 3(6): 1081-1108. |
| [8] | 唐宇琦, 叶松涛, 刘嘉, 张鑫. 分子伴侣作用下的蛋白质稳定与进化[J]. 合成生物学, 2022, 3(3): 445-464. |
| [9] | 张建志, 付立豪, 唐婷, 张嵩亚, 朱静, 李拓, 王子宁, 司同. 基于合成生物学策略的酶蛋白元件规模化挖掘[J]. 合成生物学, 2020, 1(3): 319-336. |
| [10] | 陈欣懋, 欧阳颀. 生物逆向工程设计在合成生物学中的应用[J]. 合成生物学, 2020, 1(1): 29-43. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||