Synthetic Biology Journal ›› 2022, Vol. 3 ›› Issue (6): 1277-1291.DOI: 10.12211/2096-8280.2022-004
• Invited Review • Previous Articles
Efficient synthesis of gentamicin and its related products in industrial chassis cells
WU Liangliang1, CHANG Yingying1, DENG Zixin1,2, LIU Tiangang1,2
- 1.Key Laboratory of Combinatorial Biosynthesis and Drug Discovery,Ministry of Education,Wuhan University School of Pharmaceutical Sciences,Wuhan 430071,Hubei,China
2.Hubei Engineering Laboratory for Synthetic Microbiology,Wuhan Institute of Biotechnology,Wuhan 430075,Hubei,China
-
Received:2022-01-05Revised:2022-02-23Online:2023-01-17Published:2022-12-31 -
Contact:LIU Tiangang
庆大霉素及其相关产物在工业底盘细胞中的高效合成
吴亮亮1, 常莹莹1, 邓子新1,2, 刘天罡1,2
- 1.武汉大学药学院,组合生物合成与新药发现教育部重点实验室,湖北 武汉 430071
2.武汉生物技术研究院,合成微生物技术湖北省工程实验室,湖北 武汉 430075
-
通讯作者:刘天罡 -
作者简介:吴亮亮 (1996—),男,硕士研究生。研究方向为定向合成代谢指导工业菌株中庆大霉素的产量提升。E-mail:wuliangliang@whu.edu.cn常莹莹 (1994—),女,工程师。研究方向为定向合成代谢指导工业菌株中庆大霉素的产量提升。E-mail:yingyingchang@whu.edu.cn刘天罡 (1979—),男,教授,博士生导师。研究方向为天然产物高效合成与创新发现。E-mail:liutg@whu.edu.cn -
基金资助:国家重点研发计划“合成生物学”重点专项(2018YFA0900400)
Cite this article
WU Liangliang, CHANG Yingying, DENG Zixin, LIU Tiangang. Efficient synthesis of gentamicin and its related products in industrial chassis cells[J]. Synthetic Biology Journal, 2022, 3(6): 1277-1291.
吴亮亮, 常莹莹, 邓子新, 刘天罡. 庆大霉素及其相关产物在工业底盘细胞中的高效合成[J]. 合成生物学, 2022, 3(6): 1277-1291.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2022-004
| Strain | Description | Reference |
|---|---|---|
| M. echinospora J1-020 | Wild-type | This study |
| DH10B | F–mcrA ∆(mrr-hsdRMS-mcrBC) | Gibco-BRL |
| ET12567(pUZ8002) | damdcmhsdS/pUZ8002, for intergeneric conjugation | [ |
| ET12567(pUB307) | dam dcm hsdS/pUB307, for intergeneric conjugation | [ |
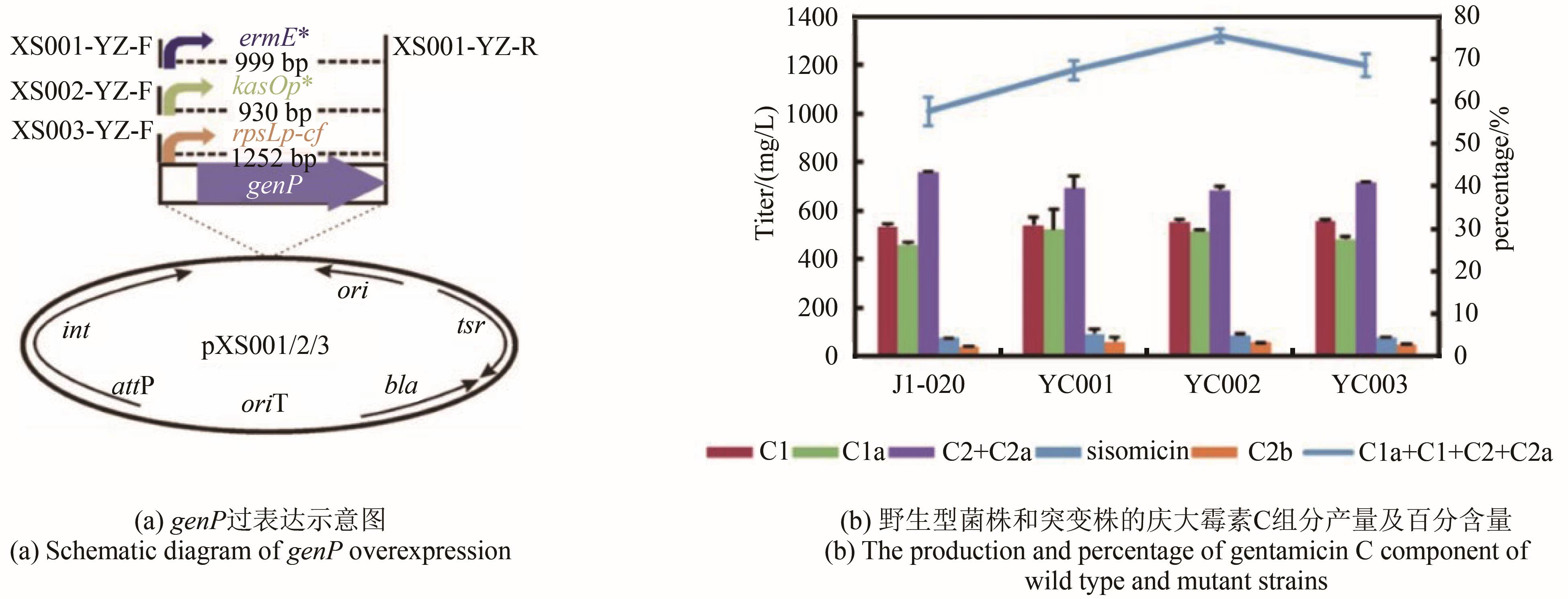
| YC001 | genP gene overexpression strain | This study |
| YC002 | genP gene overexpression strain | This study |
| YC003 | genP gene overexpression strain | This study |
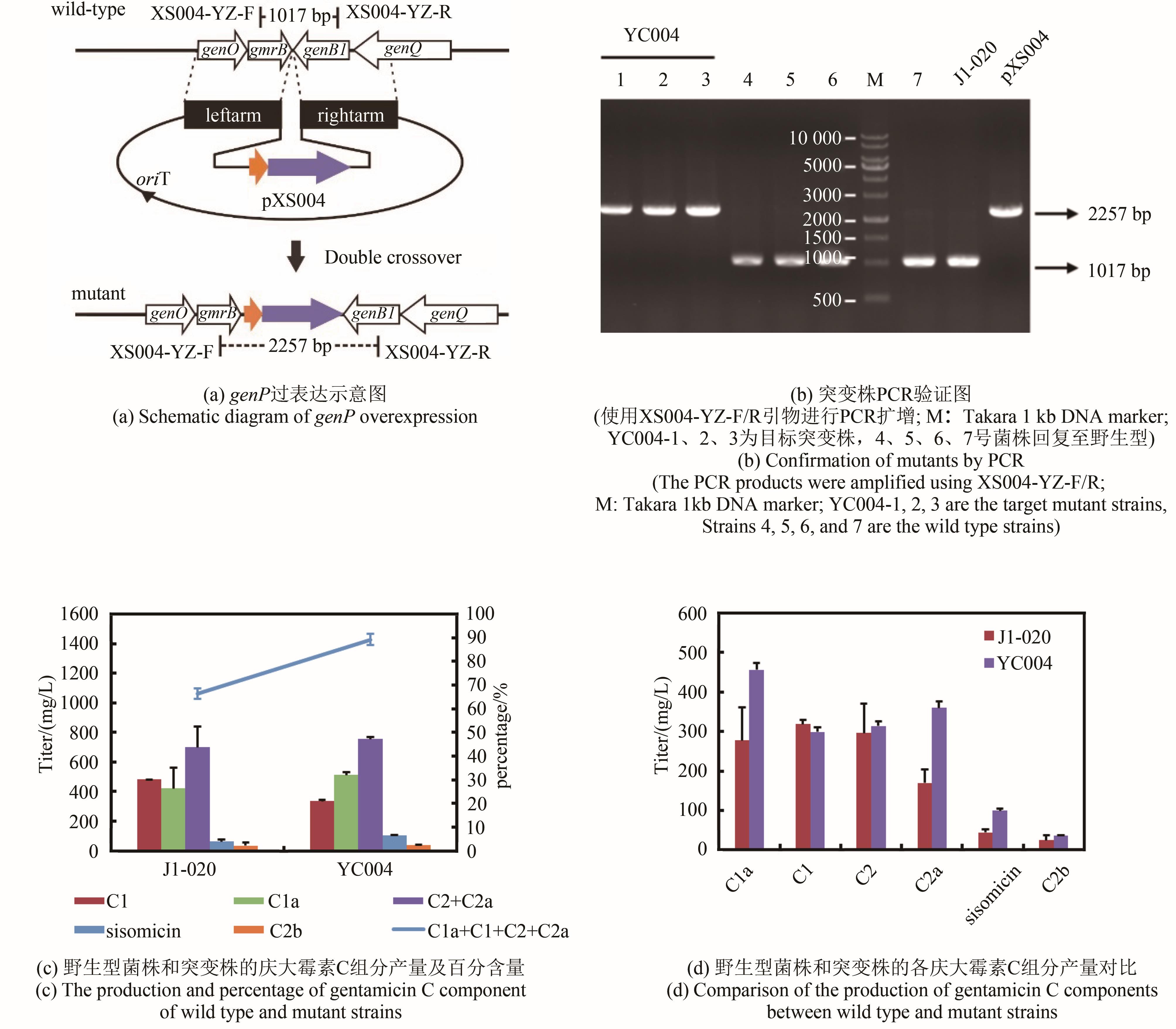
| YC004 | genP gene overexpression strain | This study |
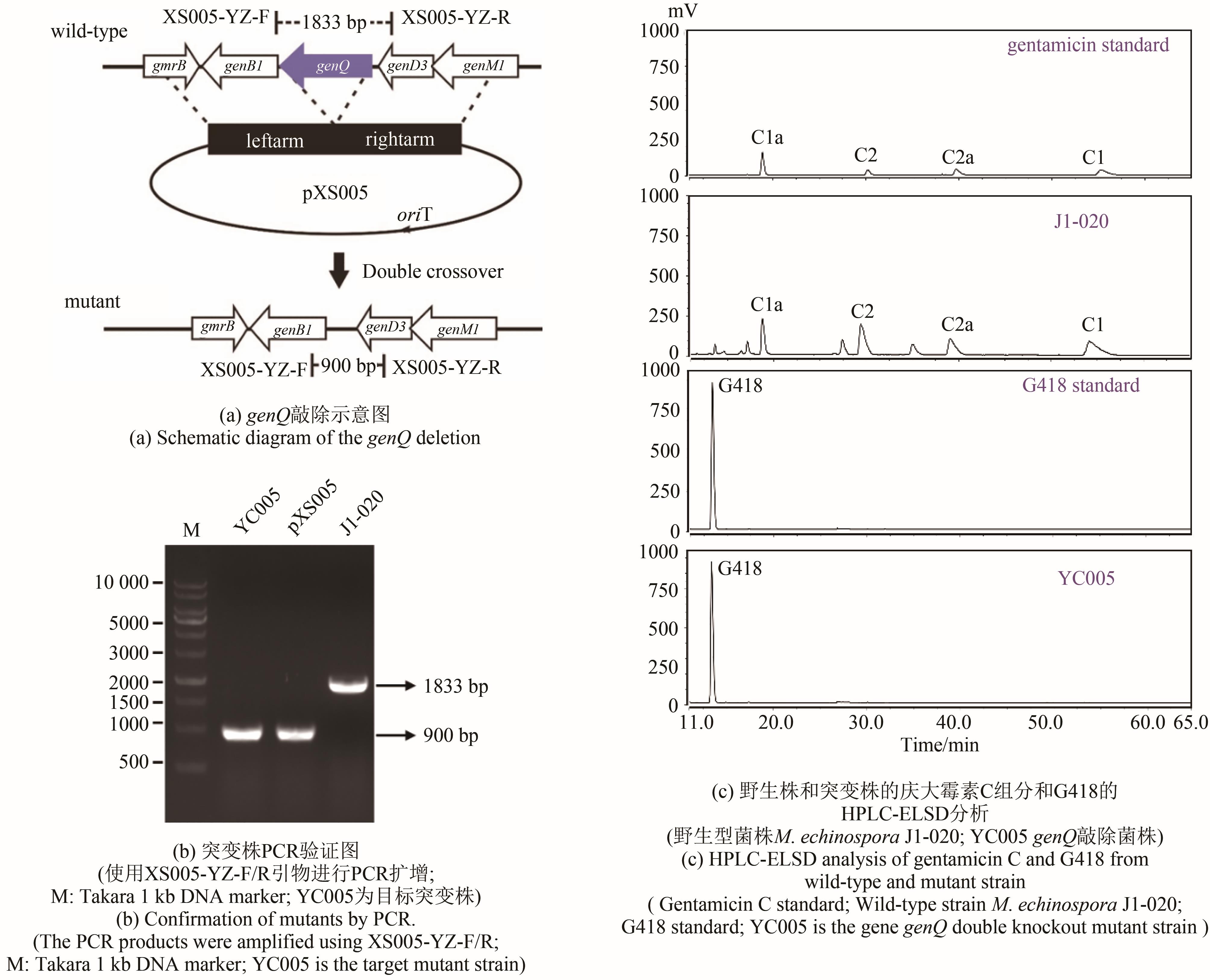
| YC005 | genQ gene knockout strain | This study |
| YC006 | genB4 gene knockout strain | This study |
| YC007 | genB4 and genK gene knockout strain | This study |
| Plasmid | ||
| pWHU77 | int, att, tsr, ermE* | [ |
| pYH7 | sti-, rep*, orf85-, R1-, tsr, aac(3)IV | [ |
| pXS001 | E.coli-actinomycete integrated shuttle plasmid, pWHU77 carries genP under the control of the ermE* promoter | This study |
| pXS002 | E.coli-actinomycete integrated shuttle plasmid, pWHU77 carries genP under the control of the kasOp* promoter | This study |
| pXS003 | E.coli-actinomycete integrated shuttle plasmid, pWHU77 carries genP under the control of the rpsLp-cf promoter | This study |
| pXS004 | E.coli-actinomycete replicating shuttle plasmid, pYH7 carries genP under the control of the rpsLp-cf promoter | This study |
| pXS005 | E.coli-actinomycete replicating shuttle plasmid, genQ gene knockout plasmid | This study |
| pXS006 | E.coli-actinomycete replicating shuttle plasmid, genB4 gene knockout plasmid | This study |
| pXS007 | E.coli-actinomycete replicating shuttle plasmid, genK gene knockout plasmid | This study |
Tab. 1 Strains and plasmids used in this study
| Strain | Description | Reference |
|---|---|---|
| M. echinospora J1-020 | Wild-type | This study |
| DH10B | F–mcrA ∆(mrr-hsdRMS-mcrBC) | Gibco-BRL |
| ET12567(pUZ8002) | damdcmhsdS/pUZ8002, for intergeneric conjugation | [ |
| ET12567(pUB307) | dam dcm hsdS/pUB307, for intergeneric conjugation | [ |
| YC001 | genP gene overexpression strain | This study |
| YC002 | genP gene overexpression strain | This study |
| YC003 | genP gene overexpression strain | This study |
| YC004 | genP gene overexpression strain | This study |
| YC005 | genQ gene knockout strain | This study |
| YC006 | genB4 gene knockout strain | This study |
| YC007 | genB4 and genK gene knockout strain | This study |
| Plasmid | ||
| pWHU77 | int, att, tsr, ermE* | [ |
| pYH7 | sti-, rep*, orf85-, R1-, tsr, aac(3)IV | [ |
| pXS001 | E.coli-actinomycete integrated shuttle plasmid, pWHU77 carries genP under the control of the ermE* promoter | This study |
| pXS002 | E.coli-actinomycete integrated shuttle plasmid, pWHU77 carries genP under the control of the kasOp* promoter | This study |
| pXS003 | E.coli-actinomycete integrated shuttle plasmid, pWHU77 carries genP under the control of the rpsLp-cf promoter | This study |
| pXS004 | E.coli-actinomycete replicating shuttle plasmid, pYH7 carries genP under the control of the rpsLp-cf promoter | This study |
| pXS005 | E.coli-actinomycete replicating shuttle plasmid, genQ gene knockout plasmid | This study |
| pXS006 | E.coli-actinomycete replicating shuttle plasmid, genB4 gene knockout plasmid | This study |
| pXS007 | E.coli-actinomycete replicating shuttle plasmid, genK gene knockout plasmid | This study |
| Primers | Sequence(5′ to 3′) | Restriction site |
|---|---|---|
| XS001-genP-F | CCA | NdeⅠ |
| XS001-genP-R | TAC | EcoRⅠ |
| XS001-YZ-F | GCGAGTGTCCGTTCGAGTGG | |
| XS001-YZ-R | TCAGAGAAATTCGTCCAGCAG | |
| XS002-kasOp*-F | GCAGGTCGACTCTAGTATGCAT | XbaⅠ |
| XS002- kasOp*-R | GTGCTGCAACCATCTTCATATGGCGTATCCCCTTTCAGATACC | |
| XS002-genP-F | TCTGAAAGGGGATACGCCATATGAAGATGGTTGCAGCACC | |
| XS002-genP-R | GGAAACAGCTATGACATGATTAC | EcoRⅠ |
| XS002-YZ-F | GGAACGATCGTTGGCTGTGTTC | |
| XS003-rpsLp-cf-F | AGGTCGACTCTAGTATGCAT | XbaⅠ |
| XS003-rpsLp-cf-R | GGTGCTGCAACCATCTTCATATGGCGTATCCCCTTTCAGATAC | |
| XS003-genP-F | TCTGAAAGGGGATACGCCATATGAAGATGGTTGCAGCACC | |
| XS003-genP-R | TTTCACACAGGAAACAGCTATGACATGATTAC | EcoRⅠ |
| XS003-YZ-F | GGAACGATCGTTGGCTGCCCGCCGCGGGCGCTG | |
| XS004-leftarm-F | ACCTGCAGGTCGACTCTAGACACGTCTGAA | NheⅠ |
| XS004-leftarm-R | CCTCCAGCGCCCGCGGCGGGCAGCCAACGATCGTTCCTCACGCCTTGTGGATCGCCACC | |
| XS004-rpsLp-cf-F | CCTGGTGGCGATCCACAAGGCGTGAGGAACGATCGTTGGCTGCCCGCCGCGGGCGCTGG | |
| XS004-genP-R | AGACCACCGCGATCGTCGAGCGCCTCTGGGAGGACTGATCAGAGAAATTCGTCCAGCAG | |
| XS004-rightarm-F | GCTGACCTACATCCAACTGCTGGACGAATTTCTCTGATCAGTCCTCCCAGAGGCGCTCG | |
| XS004-rightarm-R | TAGGCGTATCACGAGGCCCTTTCGTCTTCAA | EcoRⅠ |
| XS004-YZ-F | CCGTTCACCGTGCCCTGGCTGCGCGAGGTG | |
| XS004-YZ-R | CTCGACCCGGCCGTCTGGATCGTGGCGAAG | |
| XS005-leftarm-F | CGGCCATCGTGCCTCCCCACTCCTGC | HindⅢ |
| XS005-leftarm-R | GTGCGGCCTTCCGCGAATTCCGGGACCGGGCCCGACTGGAG | |
| XS005-rightarm-F | CGGGCCCGGTCCCGGAATTCGCGGAAGGCCGCACCGCCGAAG | |
| XS005-rightarm-R | AGCGGAAAAGATCCGTCGACCTGCAGGCATGC | BglⅡ |
| XS005-YZ-F | TCCGCTCGATTCGTTCCGTTCCGAC | |
| XS005-YZ-R | GCACGGGACCACCGGGCAGGTGCTC | |
| XS006-leftarm-F | CTGCAGGTCGACTCTAGACACGTCTGAA | NheⅠ |
| XS006-leftarm-R | AAGCTGACCTACATCCAACTGCTGGACGAATTTCTCTGATCAGCGCTGGTAGGTGCTCG | |
| XS006-rightarm-F | AGCGCTGGCGGCTGCGCGCACCTACCAGCGCTGCTAAGAGAAATTCGTCCAGCAGTTGG | |
| XS006-rightarm-R | GGCGTATCACGAGGCCCTTTCGTCTTCAA | EcoRⅠ |
| XS006-YZ-F | GGCAGCCGGACTGGGCGACCATCCGGATCG | |
| XS006-YZ-R | CCGAGGACGATCTCGTCGTCTGCCACGGTG | |
| XS007-leftarm-F | CTGCAGGTCGACTCTAGACACGTCTGAA | NheⅠ |
| XS007-leftarm-R | CCCCCTACTACGAGAGCGCCTACGAGCTGGCCCGGATGATGGCGCAGCTCGACCCGGAG | |
| XS007-rightarm-F | GTTCCTCCGGGCTCTCCGGGTCGAGCtGCGCCATCATCCGGGCCAGCTCGAGGCGCTC | |
| XS007-rightarm-R | ATAGGCGTATCACGAGGCCCTTTCGTCTTCAA | EcoRⅠ |
| XS007-YZ-F | CGTCACGCGATGGGAGCAGGGCGGAGTATC | |
| XS007-YZ-R | CCGAGCATCACGCTGCCGAAGGAGTTGGAG |
Tab. 2 Oligonucleotide primers used in this study
| Primers | Sequence(5′ to 3′) | Restriction site |
|---|---|---|
| XS001-genP-F | CCA | NdeⅠ |
| XS001-genP-R | TAC | EcoRⅠ |
| XS001-YZ-F | GCGAGTGTCCGTTCGAGTGG | |
| XS001-YZ-R | TCAGAGAAATTCGTCCAGCAG | |
| XS002-kasOp*-F | GCAGGTCGACTCTAGTATGCAT | XbaⅠ |
| XS002- kasOp*-R | GTGCTGCAACCATCTTCATATGGCGTATCCCCTTTCAGATACC | |
| XS002-genP-F | TCTGAAAGGGGATACGCCATATGAAGATGGTTGCAGCACC | |
| XS002-genP-R | GGAAACAGCTATGACATGATTAC | EcoRⅠ |
| XS002-YZ-F | GGAACGATCGTTGGCTGTGTTC | |
| XS003-rpsLp-cf-F | AGGTCGACTCTAGTATGCAT | XbaⅠ |
| XS003-rpsLp-cf-R | GGTGCTGCAACCATCTTCATATGGCGTATCCCCTTTCAGATAC | |
| XS003-genP-F | TCTGAAAGGGGATACGCCATATGAAGATGGTTGCAGCACC | |
| XS003-genP-R | TTTCACACAGGAAACAGCTATGACATGATTAC | EcoRⅠ |
| XS003-YZ-F | GGAACGATCGTTGGCTGCCCGCCGCGGGCGCTG | |
| XS004-leftarm-F | ACCTGCAGGTCGACTCTAGACACGTCTGAA | NheⅠ |
| XS004-leftarm-R | CCTCCAGCGCCCGCGGCGGGCAGCCAACGATCGTTCCTCACGCCTTGTGGATCGCCACC | |
| XS004-rpsLp-cf-F | CCTGGTGGCGATCCACAAGGCGTGAGGAACGATCGTTGGCTGCCCGCCGCGGGCGCTGG | |
| XS004-genP-R | AGACCACCGCGATCGTCGAGCGCCTCTGGGAGGACTGATCAGAGAAATTCGTCCAGCAG | |
| XS004-rightarm-F | GCTGACCTACATCCAACTGCTGGACGAATTTCTCTGATCAGTCCTCCCAGAGGCGCTCG | |
| XS004-rightarm-R | TAGGCGTATCACGAGGCCCTTTCGTCTTCAA | EcoRⅠ |
| XS004-YZ-F | CCGTTCACCGTGCCCTGGCTGCGCGAGGTG | |
| XS004-YZ-R | CTCGACCCGGCCGTCTGGATCGTGGCGAAG | |
| XS005-leftarm-F | CGGCCATCGTGCCTCCCCACTCCTGC | HindⅢ |
| XS005-leftarm-R | GTGCGGCCTTCCGCGAATTCCGGGACCGGGCCCGACTGGAG | |
| XS005-rightarm-F | CGGGCCCGGTCCCGGAATTCGCGGAAGGCCGCACCGCCGAAG | |
| XS005-rightarm-R | AGCGGAAAAGATCCGTCGACCTGCAGGCATGC | BglⅡ |
| XS005-YZ-F | TCCGCTCGATTCGTTCCGTTCCGAC | |
| XS005-YZ-R | GCACGGGACCACCGGGCAGGTGCTC | |
| XS006-leftarm-F | CTGCAGGTCGACTCTAGACACGTCTGAA | NheⅠ |
| XS006-leftarm-R | AAGCTGACCTACATCCAACTGCTGGACGAATTTCTCTGATCAGCGCTGGTAGGTGCTCG | |
| XS006-rightarm-F | AGCGCTGGCGGCTGCGCGCACCTACCAGCGCTGCTAAGAGAAATTCGTCCAGCAGTTGG | |
| XS006-rightarm-R | GGCGTATCACGAGGCCCTTTCGTCTTCAA | EcoRⅠ |
| XS006-YZ-F | GGCAGCCGGACTGGGCGACCATCCGGATCG | |
| XS006-YZ-R | CCGAGGACGATCTCGTCGTCTGCCACGGTG | |
| XS007-leftarm-F | CTGCAGGTCGACTCTAGACACGTCTGAA | NheⅠ |
| XS007-leftarm-R | CCCCCTACTACGAGAGCGCCTACGAGCTGGCCCGGATGATGGCGCAGCTCGACCCGGAG | |
| XS007-rightarm-F | GTTCCTCCGGGCTCTCCGGGTCGAGCtGCGCCATCATCCGGGCCAGCTCGAGGCGCTC | |
| XS007-rightarm-R | ATAGGCGTATCACGAGGCCCTTTCGTCTTCAA | EcoRⅠ |
| XS007-YZ-F | CGTCACGCGATGGGAGCAGGGCGGAGTATC | |
| XS007-YZ-R | CCGAGCATCACGCTGCCGAAGGAGTTGGAG |
| Type | Morphology of actinomycetes | Plasmid | The mixing ratio (Donor: Receptor) | Incubation time/h | Number of conjugants (Cultivate to 9d) |
|---|---|---|---|---|---|
| Diparental conjugation | mycelia | pWHU77 | 10∶1、4∶1、1∶1 | 14、16 | 100~200 |
| mycelia | pYH7 | 20∶1、10∶1 | 1 | ||
| Triparental conjugation | mycelia | pWHU77 | 4∶4∶1、1∶1∶1 | 100~200 | |
| mycelia | pYH7 | 4∶4∶1、1∶1∶1 | 50~100 | ||
| spore | pYH7 | 10∶10∶1、1∶1∶1 | 13 | >200 |
Tab. 3 Exploration of the conjugation conditions of M. echinospora J1-020
| Type | Morphology of actinomycetes | Plasmid | The mixing ratio (Donor: Receptor) | Incubation time/h | Number of conjugants (Cultivate to 9d) |
|---|---|---|---|---|---|
| Diparental conjugation | mycelia | pWHU77 | 10∶1、4∶1、1∶1 | 14、16 | 100~200 |
| mycelia | pYH7 | 20∶1、10∶1 | 1 | ||
| Triparental conjugation | mycelia | pWHU77 | 4∶4∶1、1∶1∶1 | 100~200 | |
| mycelia | pYH7 | 4∶4∶1、1∶1∶1 | 50~100 | ||
| spore | pYH7 | 10∶10∶1、1∶1∶1 | 13 | >200 |
| 1 | LLEWELLYN N M, SPENCER J B. Biosynthesis of 2-deoxystreptamine-containing aminoglycoside antibiotics[J]. Natural Product Reports, 2006, 23(6): 864-874. |
| 2 | Waksman SA. Antibiotics and chemotherapy[J]. California medicine, 1953,78(5):417-423. |
| 3 | SWART E A, HUTCHISON D, WAKSMAN S A. Neomycin, recovery and purification[J]. Archives of Biochemistry, 1949, 24(1): 92-103. |
| 4 | UMEZAWA H, UEDA M, MAEDA K, et al. Production and isolation of a new antibiotic: kanamycin[J]. The Journal of Antibiotics, 1957, 10(5): 181-188. |
| 5 | WEINSTEIN M J, LUEDEMANN G M, ODEN E M, et al. Gentamicin, a new antibiotic complex from micromonospora[J]. Journal of Medicinal Chemistry, 1963, 6: 463-464. |
| 6 | FOURMY D, RECHT M I, BLANCHARD S C, et al. Structure of the A site of Escherichia coli 16S ribosomal RNA complexed with an aminoglycoside antibiotic[J]. Science, 1996, 274(5291): 1367-1371. |
| 7 | TESTA R T, TILLEY B C. Biotransformation, a new approach to aminoglycoside biosynthesis: II. Gentamicin[J]. The Journal of Antibiotics, 1976, 29(2): 140-146. |
| 8 | KHAREL M K, BASNET D B, LEE H C, et al. Molecular cloning and characterization of a 2-deoxystreptamine biosynthetic gene cluster in gentamicin-producing Micromonospora echinospora ATCC 15835[J]. Molecules and Cells, 2004, 18(1): 71-78. |
| 9 | PARK J W, HONG J S J, PARAJULI N, et al. Genetic dissection of the biosynthetic route to gentamicin A2 by heterologous expression of its minimal gene set[J]. Proceedings of the National Academy of Sciences of the United States of America, 2008, 105(24): 8399-8404. |
| 10 | KIM J Y, SUH J W, KANG S H, et al. Gene inactivation study of gntE reveals its role in the first step of pseudotrisaccharide modifications in gentamicin biosynthesis[J]. Biochemical and Biophysical Research Communications, 2008, 372(4): 730-734. |
| 11 | HONG W R, YAN L B. Identification of gntK, a gene required for the methylation of purpurosamine C-6' in gentamicin biosynthesis[J]. The Journal of General and Applied Microbiology, 2012, 58(5): 349-356. |
| 12 | KIM H J, MCCARTY R M, OGASAWARA Y, et al. GenK-catalyzed C-6′ methylation in the biosynthesis of gentamicin: isolation and characterization of a cobalamin-dependent radical SAM enzyme[J]. Journal of the American Chemical Society, 2013, 135(22): 8093-8096. |
| 13 | HUANG C, HUANG F L, MOISON E, et al. Delineating the biosynthesis of gentamicin X2, the common precursor of the gentamicin C antibiotic complex[J]. Chemistry & Biology, 2015, 22(2): 251-261. |
| 14 | GUO J H, HUANG F L, HUANG C, et al. Specificity and promiscuity at the branch point in gentamicin biosynthesis[J]. Chemistry & Biology, 2014, 21(5): 608-618. |
| 15 | SHAO L, CHEN J S, WANG C X, et al. Characterization of a key aminoglycoside phosphotransferase in gentamicin biosynthesis[J]. Bioorganic & Medicinal Chemistry Letters, 2013, 23(5): 1438-1441. |
| 16 | ZHOU S T, CHEN X T, NI X P, et al. Pyridoxal-5′-phosphate-dependent enzyme GenB3 catalyzes C-3′, 4′-dideoxygenation in gentamicin biosynthesis[J]. Microbial Cell Factories, 2021, 20(1): 65. |
| 17 | CHEN X T, ZHANG H, ZHOU S T, et al. The bifunctional enzyme, GenB4, catalyzes the last step of gentamicin 3′, 4′-di-deoxygenation via reduction and transamination activities[J]. Microbial Cell Factories, 2020, 19(1): 62. |
| 18 | LI S, GUO J, REVA A, et al. Methyltransferases of gentamicin biosynthesis[J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(6): 1340-1345. |
| 19 | BAN Y H, SONG M C, HWANG J Y, et al. Complete reconstitution of the diverse pathways of gentamicin B biosynthesis[J]. Nature Chemical Biology, 2019, 15(3): 295-303. |
| 20 | CHANG Y Y, CHAI B Z, DING Y K, et al. Overproduction of gentamicin B in industrial strain Micromonospora echinospora CCTCC M 2018898 by cloning of the missing genes genR and genS [J]. Metabolic Engineering Communications, 2019, 9: e00096. |
| 21 | MACNEIL D J, OCCI J L, GEWAIN K M, et al. Complex organization of the Streptomyces avermitilis genes encoding the avermectin polyketide synthase[J]. Gene, 1992, 115(1/2): 119-125. |
| 22 | FLETT F, MERSINIAS V, SMITH C P. High efficiency intergeneric conjugal transfer of plasmid DNA from Escherichia coli to methyl DNA-restricting streptomycetes[J]. FEMS Microbiology Letters, 1997, 155(2): 223-229. |
| 23 | SUN Y H, HONG H, SAMBORSKYY M, et al. Organization of the biosynthetic gene cluster in Streptomyces sp. DSM 4137 for the novel neuroprotectant polyketide meridamycin[J]. Microbiology, 2006, 152(Pt 12): 3507-3515. |
| 24 | WEBER T, BLIN K, DUDDELA S, et al. antiSMASH 3.0—a comprehensive resource for the genome mining of biosynthetic gene clusters[J]. Nucleic Acids Research, 2015, 43(W1): W237-W243. |
| 25 | BILYK O, LUZHETSKYY A. Metabolic engineering of natural product biosynthesis in actinobacteria[J]. Current Opinion in Biotechnology, 2016, 42: 98-107. |
| 26 | SASAKI K, MIZUSAWA H, ISHIDATE M, et al. Regulation of G418 selection efficiency by cell-cell interaction in transfection[J]. Somatic Cell and Molecular Genetics, 1992, 18(6): 517-527. |
| 27 | BAIAZITOV R Y, FRIESEN W, JOHNSON B, et al. Chemical modifications of G418 (geneticin): synthesis of novel readthrough aminoglycosides results in an improved in vitro safety window but no improvements in vivo [J]. Carbohydrate Research, 2020, 495: 108058. |
| 28 | NI X P, SUN Z P, ZHANG H Y, et al. Genetic engineering combined with random mutagenesis to enhance G418 production in Micromonospora echinospora [J]. Journal of Industrial Microbiology & Biotechnology, 2014, 41(9): 1383-1390. |
| 29 | SHAEER K M, ZMARLICKA M T, CHAHINE E B, et al. Plazomicin: a next-generation aminoglycoside[J]. Pharmacotherapy, 2019, 39(1): 77-93. |
| 30 | NI X P, SUN Z P, GU Y W, et al. Assembly of a novel biosynthetic pathway for gentamicin B production in Micromonospora echinospora [J]. Microbial Cell Factories, 2016, 15: 1. |
| 31 | WU Z, GAO W L, ZHOU S T, et al. Improving gentamicin B and gentamicin C1a production by engineering the glycosyltransferases that transfer primary metabolites into secondary metabolites biosynthesis[J]. Microbiological Research, 2017, 203: 40-46. |
| 32 | LI D, LI H, NI X P, et al. Construction of a gentamicin C1a-overproducing strain of Micromonospora purpurea by inactivation of the gacD gene[J]. Microbiological research, 2013, 168(5): 263-267. |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||