合成生物学 ›› 2025, Vol. 6 ›› Issue (4): 829-845.DOI: 10.12211/2096-8280.2025-030
基于转录因子生物传感器的构建与应用进展
王宏1, 陆孔泳2, 郑洋洋1, 陈涛1, 王智文1,2
- 1.天津大学化工学院,天津 300350
2.宁夏大学生命科学学院,宁夏 银川 750021
-
收稿日期:2025-03-28修回日期:2025-05-29出版日期:2025-08-31发布日期:2025-09-03 -
通讯作者:王智文 -
作者简介:王宏 (1999—),男,硕士研究生。研究方向为代谢工程与合成生物学。 E-mail:wh17826529977@163.com陆孔泳 (1989—),女,博士,副教授,硕士生导师。研究方向为合成生物学、生物发酵与代谢调控、微藻生物技术、工业微生物技术等。E-mail:lky@nxu.edu.cn王智文 (1981—),男,博士,教授,博士生导师。研究方向为基因组编辑与合成生物学元件开发、基因组尺度网络模型构建与途径模拟设计、合成高附加值生物医药与生物基化学品人工细胞工厂构建、微生物资源挖掘与利用等。E-mail:zww@tju.edu.cn
第一联系人:共同第一作者 -
基金资助:宁夏重点研发基金资助项目(2024BEE02005)
Construction and advances in the applications of transcription factor-based biosensors
WANG Hong1, LU Kongyong2, ZHENG Yangyang1, CHEN Tao1, WANG Zhiwen1,2
- 1.School of Chemical Engineering and Technology,Tianjin University,Tianjin 300350,China
2.College of Life Science,Ningxia University,Yinchuan 750021,Ningxia,China
-
Received:2025-03-28Revised:2025-05-29Online:2025-08-31Published:2025-09-03 -
Contact:WANG Zhiwen
摘要:
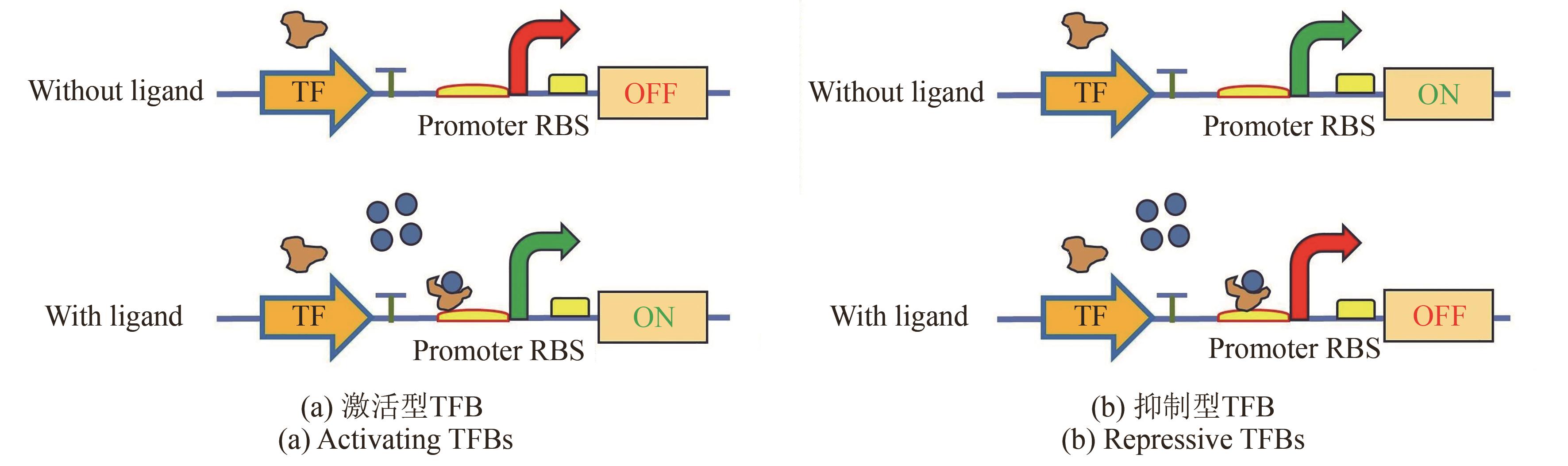
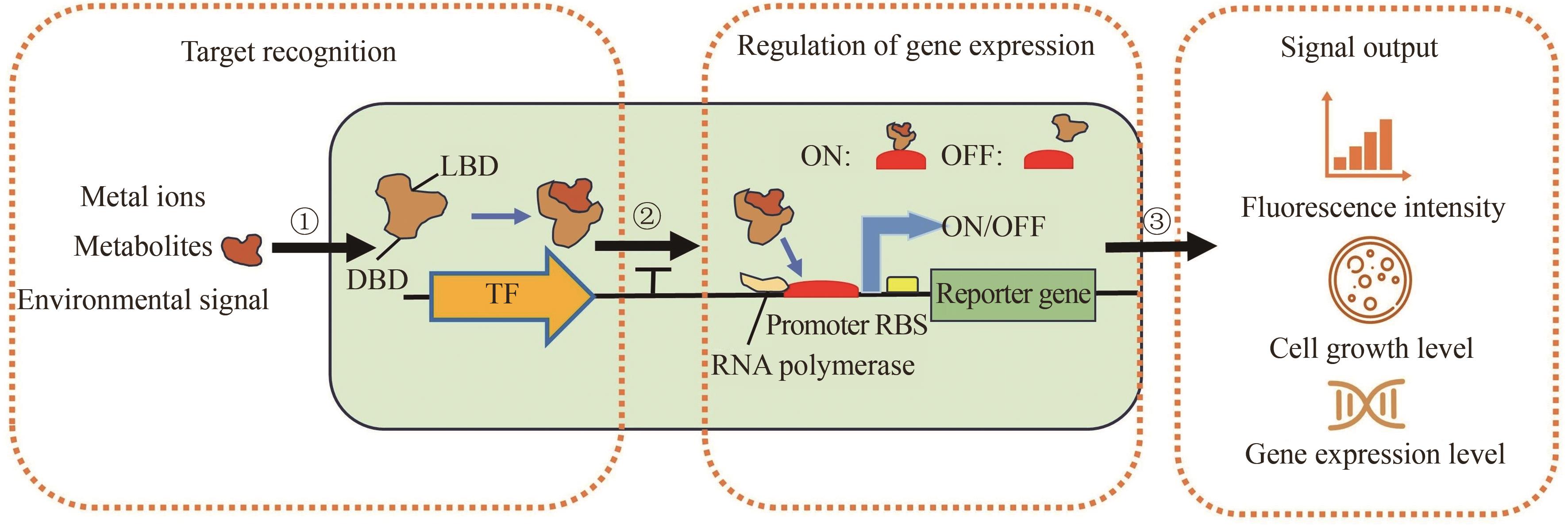
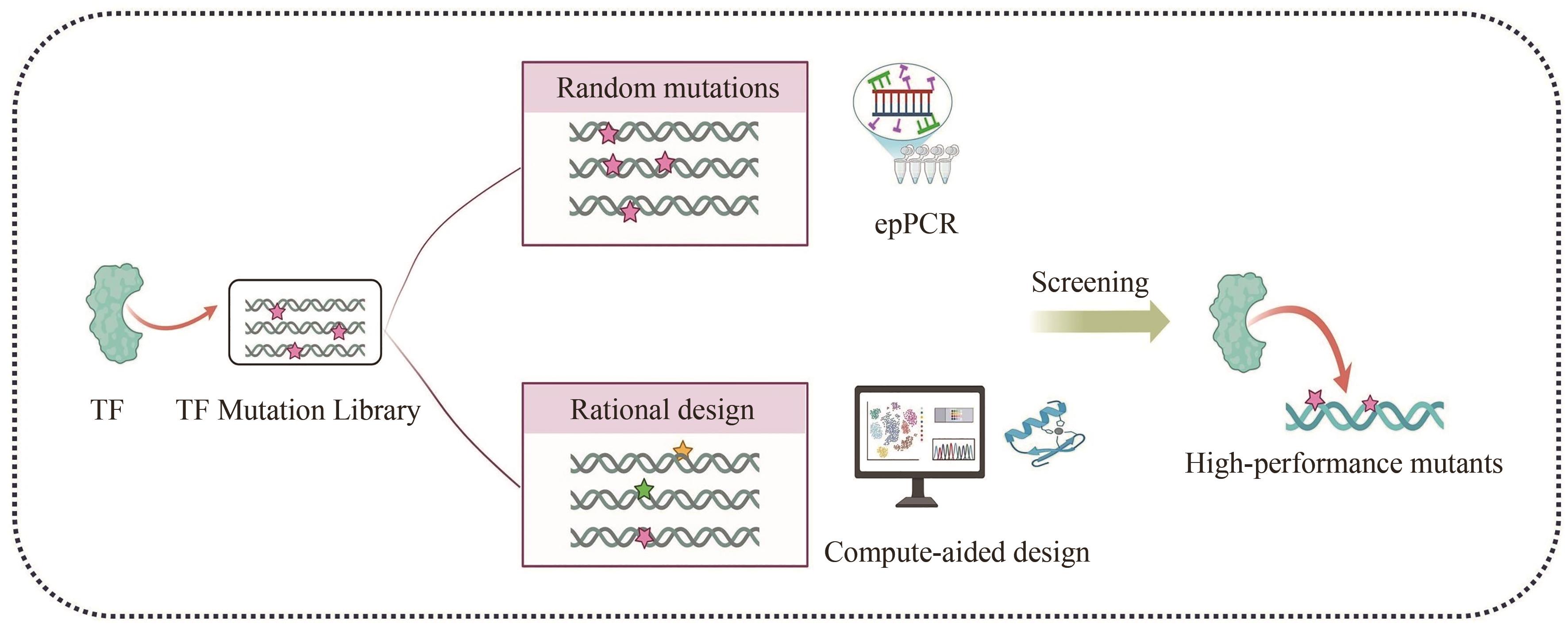
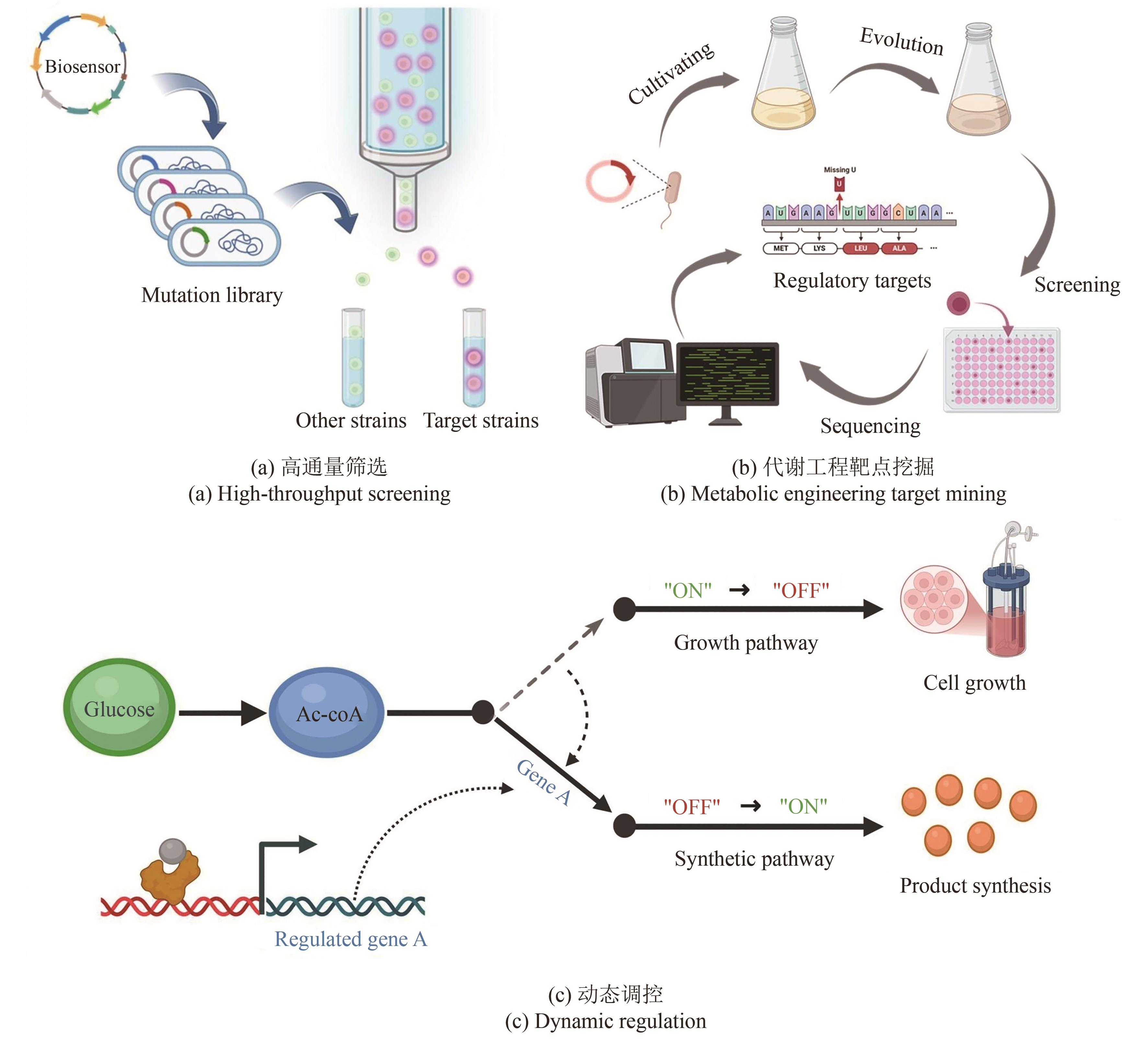
微生物细胞工厂作为绿色生物制造的重要实现形式,广泛应用于食品、化工、医药和能源等领域。然而,利用传统代谢工程策略改造微生物细胞工厂生产目标产品时,仍面临静态代谢调控的局限性与代谢通量实时监测的滞后性等问题,制约着生物基产品的高效生物合成。基于转录因子生物传感器通过实时感知代谢物浓度信号或环境信号,自动调控目的基因表达,为微生物细胞工厂的高效构建与智能化调控提供了创新性解决方案。本文介绍了基于转录因子生物传感器的组成、分类及作用机制,围绕传感器配体识别模块的设计和信号输出模块的元件重构,总结了基于转录因子生物传感器的构建策略,对基于转录因子生物传感器在微生物细胞工厂中的应用进展进行了综述,包括高通量筛选、代谢工程靶点挖掘以及动态调控。聚焦目前基于转录因子生物传感器面临的代谢物响应元件匮乏、检测范围受限、配体识别特异性不足、转录依赖的耗时性和传感器元件鲁棒性缺陷等挑战,对未来的研究方向进行展望,为未来基于转录因子生物传感器的构建与应用提供借鉴。
中图分类号:
引用本文
王宏, 陆孔泳, 郑洋洋, 陈涛, 王智文. 基于转录因子生物传感器的构建与应用进展[J]. 合成生物学, 2025, 6(4): 829-845.
WANG Hong, LU Kongyong, ZHENG Yangyang, CHEN Tao, WANG Zhiwen. Construction and advances in the applications of transcription factor-based biosensors[J]. Synthetic Biology Journal, 2025, 6(4): 829-845.
| 转录因子 | 宿主 | 响应物质 | 主要构建策略 | 传感器性能 | 参考文献 |
|---|---|---|---|---|---|
| EryD | 解脂耶氏酵母 | 赤藓糖醇 | 启动子筛选替换 | 检测范围扩展至0~400 mmol/L,500 mmol/L时信号饱和 | [ |
| CouR | 酿酒酵母 | 对香豆酰辅酶 A | 启动子元件重构 | 动态范围高达21.4 倍,具有高度底物特异性 | [ |
| TrpR | 大肠杆菌 | 色氨酸 | TrpR及启动子的定向进化 | 检测范围从125 mg/L扩展到500 mg/L | [ |
| BsNadR | 大肠杆菌 | 烟酸 | BsNadR的定向进化 | 检测范围扩展至0~50 mmol/L | [ |
| RamR | 大肠杆菌 | 苄基异喹啉生物碱 | RamR的定向进化 | 对5种BIA具有高特异性和灵敏度 | [ |
| AlkS | 大肠杆菌 | 短链氯代脂肪烃 | AlkS的定向进化 | 荧光输出增加150倍,检测限降低至0.03 mg/kg | [ |
| BenM | 大肠杆菌 | 己二酸 | BenM的理性设计 | 配体特异性改变,灵敏度提高约3倍 | [ |
| MarR | 大肠杆菌 | 阿司匹林 | MarR的理性设计 | 配体特异性改变,检测限低至0.01 mmol/L | [ |
| FeaR | 大肠杆菌 | 芳香胺 | FeaR的理性设计 | 动态范围高达580倍,对苯乙胺特异性响应 | [ |
| MyrR | 大肠杆菌 | β-蒎烯 | 启动子元件重构 | 动态范围提高54倍,检测范围扩展至0~160 mg/L | [ |
| TtgV | 大肠杆菌 | 3-甲基吲哚 | 启动子及质粒拷贝数优化 | 检测限低至10 μmol/L,检测范围扩展至10~1750 μmol/L | [ |
| LldR | 大肠杆菌 | 乳酸 | 启动子元件重构 | 检测限低至2.34 mmol/L,动态范围提高14倍 | [ |
| BreR | 大肠杆菌 | 胆汁酸 | 启动子元件重构 | 动态范围提高至470倍,检测限低至0.61 μmol/L | [ |
| MphR | 大肠杆菌 | 红霉素 | RBS替换优化MphR表达 | 获得了灵敏度差异超过10倍的传感器变体 | [ |
| CdaR | 大肠杆菌 | 葡萄糖酸 | 交叉RBS组合文库筛选 | 动态范围从最初的9倍提升到最高247倍 | [ |
| FapR | 大肠杆菌 | 丙二酰辅酶A | 启动子与RBS替换 | 低浓度mCoA下表现出更高的荧光输出 | [ |
| CamR | 恶臭假单胞菌 | 丁醇类 | CamR的定向进化 | 特异性响应正丁醇,展现出显著底物区分能力 | [ |
| LysG | 谷氨酸棒状杆菌 | γ-氨基丁酸 | LysG的定向进化 | 检测限降低至0.2 μmol/L,动态范围扩展至350倍 | [ |
| PdhR | 枯草芽孢杆菌 | 丙酮酸 | 启动子元件重构 | 动态范围从0.6倍提升至30.7倍 | [ |
表1 基于转录因子构建的生物传感器
Table 1 Biosensors based on transcription factors
| 转录因子 | 宿主 | 响应物质 | 主要构建策略 | 传感器性能 | 参考文献 |
|---|---|---|---|---|---|
| EryD | 解脂耶氏酵母 | 赤藓糖醇 | 启动子筛选替换 | 检测范围扩展至0~400 mmol/L,500 mmol/L时信号饱和 | [ |
| CouR | 酿酒酵母 | 对香豆酰辅酶 A | 启动子元件重构 | 动态范围高达21.4 倍,具有高度底物特异性 | [ |
| TrpR | 大肠杆菌 | 色氨酸 | TrpR及启动子的定向进化 | 检测范围从125 mg/L扩展到500 mg/L | [ |
| BsNadR | 大肠杆菌 | 烟酸 | BsNadR的定向进化 | 检测范围扩展至0~50 mmol/L | [ |
| RamR | 大肠杆菌 | 苄基异喹啉生物碱 | RamR的定向进化 | 对5种BIA具有高特异性和灵敏度 | [ |
| AlkS | 大肠杆菌 | 短链氯代脂肪烃 | AlkS的定向进化 | 荧光输出增加150倍,检测限降低至0.03 mg/kg | [ |
| BenM | 大肠杆菌 | 己二酸 | BenM的理性设计 | 配体特异性改变,灵敏度提高约3倍 | [ |
| MarR | 大肠杆菌 | 阿司匹林 | MarR的理性设计 | 配体特异性改变,检测限低至0.01 mmol/L | [ |
| FeaR | 大肠杆菌 | 芳香胺 | FeaR的理性设计 | 动态范围高达580倍,对苯乙胺特异性响应 | [ |
| MyrR | 大肠杆菌 | β-蒎烯 | 启动子元件重构 | 动态范围提高54倍,检测范围扩展至0~160 mg/L | [ |
| TtgV | 大肠杆菌 | 3-甲基吲哚 | 启动子及质粒拷贝数优化 | 检测限低至10 μmol/L,检测范围扩展至10~1750 μmol/L | [ |
| LldR | 大肠杆菌 | 乳酸 | 启动子元件重构 | 检测限低至2.34 mmol/L,动态范围提高14倍 | [ |
| BreR | 大肠杆菌 | 胆汁酸 | 启动子元件重构 | 动态范围提高至470倍,检测限低至0.61 μmol/L | [ |
| MphR | 大肠杆菌 | 红霉素 | RBS替换优化MphR表达 | 获得了灵敏度差异超过10倍的传感器变体 | [ |
| CdaR | 大肠杆菌 | 葡萄糖酸 | 交叉RBS组合文库筛选 | 动态范围从最初的9倍提升到最高247倍 | [ |
| FapR | 大肠杆菌 | 丙二酰辅酶A | 启动子与RBS替换 | 低浓度mCoA下表现出更高的荧光输出 | [ |
| CamR | 恶臭假单胞菌 | 丁醇类 | CamR的定向进化 | 特异性响应正丁醇,展现出显著底物区分能力 | [ |
| LysG | 谷氨酸棒状杆菌 | γ-氨基丁酸 | LysG的定向进化 | 检测限降低至0.2 μmol/L,动态范围扩展至350倍 | [ |
| PdhR | 枯草芽孢杆菌 | 丙酮酸 | 启动子元件重构 | 动态范围从0.6倍提升至30.7倍 | [ |
| 转录因子 | 宿主 | 响应物质 | 应用领域 | 应用结果 | 参考文献 |
|---|---|---|---|---|---|
| TtgR | 大肠杆菌 | 2S-柚皮素 | 高通量筛选 | 筛选出催化活性提高2.34倍的查尔酮合酶突变体 | [ |
| BmoR | 大肠杆菌 | 乙二醇 | 高通量筛选 | 筛选获得的SMM3F突变体催化活性提高1.52倍 | [ |
| YqhC | 大肠杆菌 | 香兰素 | 高通量筛选 | 筛选获得的Mu176突变体催化活性提高7倍 | [ |
| XylS | 大肠杆菌 | 3-甲基水杨酸 | 高通量筛选 | 筛选出催化效率比野生型提高15倍的突变酶 | [ |
| AlkS | 大肠杆菌 | 异戊醇 | 高通量筛选 | 筛选出的异戊醇产量提高45倍的突变菌株 | [ |
| LysG | 谷氨酸棒状杆菌 | L-组氨酸 | 高通量筛选 | 筛选出100个独立的L-组氨酸高产菌株 | [ |
| Leu3p | 酿酒酵母 | α-异丙基苹果酸 | 高通量筛选 | 筛选出异丁醇产量高达725 mg/L的突变菌株 | [ |
| EryD | 解脂耶氏酵母 | 赤藓糖醇 | 高通量筛选 | 筛选出赤藓糖醇产量较原始菌株提升4.4倍的突变菌株 | [ |
| PadR | 大肠杆菌 | 对香豆酸 | 代谢工程靶点挖掘 | 挖掘到与对香豆酸生产相关的靶点pfkA和ptsI | [ |
| LldR | 运动发酵单胞菌 | D-乳酸 | 代谢工程靶点挖掘 | 挖掘到与D-乳酸生产相关的靶点ZMO1323和ZMO1530 | [ |
| Lrp | 谷氨酸棒状杆菌 | 支链氨基酸 | 代谢工程靶点挖掘 | 挖掘到与支链氨基酸合成相关的靶点AHAS | [ |
| CouR | 酿酒酵母 | 对香豆酰辅酶A | 动态调控 | 柚皮素产量达47.3 mg/L,与未调控相比提高15倍 | [ |
| Mlc | 大肠杆菌 | 葡萄糖 | 动态调控 | 动态调控大肠杆菌葡萄糖摄取速率 | [ |
| GlcC | 大肠杆菌 | 乙醇酸 | 动态调控 | 动态调控gltA、ycdW和aceA的表达水平,乙醇酸产量达到52.2 g/L | [ |
| ivbL、BmoR | 大肠杆菌 | 氨基酸、高级醇 | 动态调控 | 动态平衡氨基酸向高级醇转化,异丁醇产量达40.4 g/L | [ |
| PadR | 大肠杆菌 | 对香豆酸 | 动态调控 | 动态调控丙二酰辅酶A合成,覆盆子酮产量提高32.4倍 | [ |
| LacI | 枯草芽孢杆菌 | 乳糖 | 动态调控 | 动态调控glcK表达,2'-岩藻糖基乳糖产量达到30.1 g/L | [ |
| Rex | 希瓦氏菌 | NADH/NAD⁺ | 动态调控 | 动态调控异丁醇合成途径,异丁醇产量提高10.8倍 | [ |
| ChnR | 谷氨酸棒状杆菌 | 戊内酰胺 | 动态调控 | 动态上调Act的表达水平,戊内酰胺产量提高10倍以上 | [ |
表2 基于转录因子生物传感器在微生物细胞工厂中的应用
Table 2 Applications of transcription factor-based biosensors in microbial cell factory
| 转录因子 | 宿主 | 响应物质 | 应用领域 | 应用结果 | 参考文献 |
|---|---|---|---|---|---|
| TtgR | 大肠杆菌 | 2S-柚皮素 | 高通量筛选 | 筛选出催化活性提高2.34倍的查尔酮合酶突变体 | [ |
| BmoR | 大肠杆菌 | 乙二醇 | 高通量筛选 | 筛选获得的SMM3F突变体催化活性提高1.52倍 | [ |
| YqhC | 大肠杆菌 | 香兰素 | 高通量筛选 | 筛选获得的Mu176突变体催化活性提高7倍 | [ |
| XylS | 大肠杆菌 | 3-甲基水杨酸 | 高通量筛选 | 筛选出催化效率比野生型提高15倍的突变酶 | [ |
| AlkS | 大肠杆菌 | 异戊醇 | 高通量筛选 | 筛选出的异戊醇产量提高45倍的突变菌株 | [ |
| LysG | 谷氨酸棒状杆菌 | L-组氨酸 | 高通量筛选 | 筛选出100个独立的L-组氨酸高产菌株 | [ |
| Leu3p | 酿酒酵母 | α-异丙基苹果酸 | 高通量筛选 | 筛选出异丁醇产量高达725 mg/L的突变菌株 | [ |
| EryD | 解脂耶氏酵母 | 赤藓糖醇 | 高通量筛选 | 筛选出赤藓糖醇产量较原始菌株提升4.4倍的突变菌株 | [ |
| PadR | 大肠杆菌 | 对香豆酸 | 代谢工程靶点挖掘 | 挖掘到与对香豆酸生产相关的靶点pfkA和ptsI | [ |
| LldR | 运动发酵单胞菌 | D-乳酸 | 代谢工程靶点挖掘 | 挖掘到与D-乳酸生产相关的靶点ZMO1323和ZMO1530 | [ |
| Lrp | 谷氨酸棒状杆菌 | 支链氨基酸 | 代谢工程靶点挖掘 | 挖掘到与支链氨基酸合成相关的靶点AHAS | [ |
| CouR | 酿酒酵母 | 对香豆酰辅酶A | 动态调控 | 柚皮素产量达47.3 mg/L,与未调控相比提高15倍 | [ |
| Mlc | 大肠杆菌 | 葡萄糖 | 动态调控 | 动态调控大肠杆菌葡萄糖摄取速率 | [ |
| GlcC | 大肠杆菌 | 乙醇酸 | 动态调控 | 动态调控gltA、ycdW和aceA的表达水平,乙醇酸产量达到52.2 g/L | [ |
| ivbL、BmoR | 大肠杆菌 | 氨基酸、高级醇 | 动态调控 | 动态平衡氨基酸向高级醇转化,异丁醇产量达40.4 g/L | [ |
| PadR | 大肠杆菌 | 对香豆酸 | 动态调控 | 动态调控丙二酰辅酶A合成,覆盆子酮产量提高32.4倍 | [ |
| LacI | 枯草芽孢杆菌 | 乳糖 | 动态调控 | 动态调控glcK表达,2'-岩藻糖基乳糖产量达到30.1 g/L | [ |
| Rex | 希瓦氏菌 | NADH/NAD⁺ | 动态调控 | 动态调控异丁醇合成途径,异丁醇产量提高10.8倍 | [ |
| ChnR | 谷氨酸棒状杆菌 | 戊内酰胺 | 动态调控 | 动态上调Act的表达水平,戊内酰胺产量提高10倍以上 | [ |
| [1] | SHI A Q, ZHU X N, LU J, et al. Activating transhydrogenase and NAD kinase in combination for improving isobutanol production[J]. Metabolic Engineering, 2013, 16: 1-10. |
| [2] | LUCKIE B A, KASHYAP M, PEARSON A N, et al. Development of Corynebacterium glutamicum as a monoterpene production platform[J]. Metabolic Engineering, 2024, 81: 110-122. |
| [3] | PADDON C J, WESTFALL P J, PITERA D J, et al. High-level semi-synthetic production of the potent antimalarial artemisinin[J]. Nature, 2013, 496(7446): 528-532. |
| [4] | 于政, 申晓林, 孙新晓, 等. 动态调控策略在代谢工程中的应用研究进展[J]. 合成生物学, 2020, 1(4): 440-453. |
| YU Z, SHEN X L, SUN X X, et al. Application of dynamic regulation strategies in metabolic engineering[J]. Synthetic Biology Journal, 2020, 1(4): 440-453. | |
| [5] | MACHADO D, COSTA R S, FERREIRA E C, et al. Exploring the gap between dynamic and constraint-based models of metabolism[J]. Metabolic Engineering, 2012, 14(2): 112-119. |
| [6] | 程术, 邓子新, 卞光凯, 等. 萜类高效合成平台的搭建与萜类产物批量挖掘[J]. 生命科学, 2019, 31(5): 449-457. |
| CHENG S, DENG Z X, BIAN G K, et al. Construction of high-efficient terpenoid platform and the application in terpenoid discovery[J]. Chinese Bulletin of Life Sciences, 2019, 31(5): 449-457. | |
| [7] | XU X H, LV X Q, BI X Y, et al. Genetic circuits for metabolic flux optimization[J]. Trends in Microbiology, 2024, 32(8): 791-806. |
| [8] | 洪霞, 田开仁, 乔建军, 等. 基因编码型生物传感器在微生物细胞工厂中的应用进展[J]. 中国生物工程杂志, 2023, 43(9): 62-76. |
| HONG X, TIAN K R, QIAO J J, et al. Application progress of genetically encoded biosensors in microbial cell factory[J]. China Biotechnology, 2023, 43(9): 62-76. | |
| [9] | YU W W, XU X H, JIN K, et al. Genetically encoded biosensors for microbial synthetic biology: from conceptual frameworks to practical applications[J]. Biotechnology Advances, 2023, 62: 108077. |
| [10] | VERMA A K, NOUMANI A, YADAV A K, et al. FRET based biosensor: principle applications recent advances and challenges[J]. Diagnostics, 2023, 13(8): 1375. |
| [11] | FERREIRA S S, ANTUNES M S. Re-engineering plant phenylpropanoid metabolism with the aid of synthetic biosensors[J]. Frontiers in Plant Science, 2021, 12: 701385. |
| [12] | CARPENTER A C, PAULSEN I T, WILLIAMS T C. Blueprints for biosensors: design, limitations, and applications[J]. Genes, 2018, 9(8): 375. |
| [13] | LI C F, WANG C, ZHU J, et al. Advances and prospects of transcription-factor-based biosensors in high-throughput screening for cell factories construction[J]. Food Bioengineering, 2022, 1(2): 135-147. |
| [14] | TU R, ZHANG Y, HUA E B, et al. Droplet-based microfluidic platform for high-throughput screening of Streptomyces [J]. Communications Biology, 2021, 4: 647. |
| [15] | YI D, BAYER T, BADENHORST C P S, et al. Recent trends in biocatalysis[J]. Chemical Society Reviews, 2021, 50(14): 8003-8049. |
| [16] | KIM H, JU J, LEE H N, et al. Genetically encoded biosensors based on fluorescent proteins[J]. Sensors, 2021, 21(3): 795. |
| [17] | 赵静宇, 张健, 祁庆生, 等. 基于细菌双组分系统的生物传感器的研究进展[J]. 合成生物学, 2024, 5(1): 38-52. |
| ZHAO J Y, ZHANG J, QI Q S, et al. Research progress in biosensors based on bacterial two-component systems[J]. Synthetic Biology Journal, 2024, 5(1): 38-52. | |
| [18] | CHEN L, ZHANG Z H, LI Z H, et al. Learning protein fitness landscapes with deep mutational scanning data from multiple sources[J]. Cell Systems, 2023, 14(8): 706-721.e5. |
| [19] | LI J W, QIN Z Q, ZHANG B H, et al. Development of transcriptional factor-based whole-cell biosensors to monitor and degrade antibiotics using mutant cells obtained via adaptive laboratory evolution[J]. Journal of Hazardous Materials, 2024, 473: 134536. |
| [20] | KANG Z Q, ZHANG M M, GAO K Y, et al. An l-2-hydroxyglutarate biosensor based on specific transcriptional regulator LhgR[J]. Nature Communications, 2021, 12: 3619. |
| [21] | 周子莹, 宋晓东, 刘洋儿, 等. 变构转录因子生物传感器构建策略及在食品安全中的应用进展[J]. 生物技术通报, 2024, 40(12): 20-33. |
| ZHOU Z Y, SONG X D, LIU Y E, et al. Construction strategies of allosteric transcription factor biosensors and their application advances in food safety[J]. Biotechnology Bulletin, 2024, 40(12): 20-33. | |
| [22] | LI M, CHEN Z Y, HUO Y X. Application evaluation and performance-directed improvement of the native and engineered biosensors[J]. ACS Sensors, 2024, 9(10): 5002-5024. |
| [23] | XIAO C F, PAN Y Y, HUANG M T. Advances in the dynamic control of metabolic pathways in Saccharomyces cerevisiae [J]. Engineering Microbiology, 2023, 3(4): 100103. |
| [24] | MITCHLER M M, GARCIA J M, MONTERO N E, et al. Transcription factor-based biosensors: a molecular-guided approach for natural product engineering[J]. Current Opinion in Biotechnology, 2021, 69: 172-181. |
| [25] | LIU Y E, ZHOU Z Y, WU Y F, et al. Engineered transcription factor-binding diversed functional nucleic acid-based synthetic biosensor[J]. Biotechnology Advances, 2024, 77: 108463. |
| [26] | DE PAEPE B, DE MEY M. Biological switches: past and future milestones of transcription factor-based biosensors[J]. ACS Synthetic Biology, 2025, 14(1): 72-86. |
| [27] | CHAISUPA P, WRIGHT R C. State-of-the-art in engineering small molecule biosensors and their applications in metabolic engineering[J]. SLAS Technology, 2024, 29(2): 100113. |
| [28] | HUTTANUS H M, TRIOLA E H, VELASQUEZ-GUZMAN J C, et al. Targeted mutagenesis and high-throughput screening of diversified gene and promoter libraries for isolating gain-of-function mutations[J]. Frontiers in Bioengineering and Biotechnology, 2023, 11: 1202388. |
| [29] | ZHANG Y F, CORTEZ J D, HAMMER S K, et al. Biosensor for branched-chain amino acid metabolism in yeast and applications in isobutanol and isopentanol production[J]. Nature Communications, 2022, 13: 270. |
| [30] | LI H M, ZHANG W, HAN Y Y, et al. Programming a bacterial biosensor for directed evolution of tryptophan hydroxylase via high-throughput droplet sorting[J]. Biosensors & Bioelectronics, 2025, 271: 117072. |
| [31] | 刘静, 李龙, 王云霞, 等. 细菌DeoR家族转录调控因子的研究进展[J]. 微生物学报, 2022, 62(3): 906-917. |
| LIU J, LI L, WANG Y X, et al. Progress on the DeoR family transcriptional regulators in bacteria[J]. Acta Microbiologica Sinica, 2022, 62(3): 906-917. | |
| [32] | YEOM S J, KIM M, KWON K K, et al. A synthetic microbial biosensor for high-throughput screening of lactam biocatalysts[J]. Nature Communications, 2018, 9: 5053. |
| [33] | PHAM C, STOGIOS P J, SAVCHENKO A, et al. Advances in engineering and optimization of transcription factor-based biosensors for plug-and-play small molecule detection[J]. Current Opinion in Biotechnology, 2022, 76: 102753. |
| [34] | SHEN Y P, PAN Y Y, NIU F X, et al. Biosensor-assisted evolution for high-level production of 4-hydroxyphenylacetic acid in Escherichia coli [J]. Metabolic Engineering, 2022, 70: 1-11. |
| [35] | SU H F, CHEN S J, CHEN X L, et al. Utilizing a high-throughput visualization screening technology to develop a genetically encoded biosensor for monitoring 5-aminolevulinic acid production in engineered Escherichia coli [J]. Biosensors and Bioelectronics, 2025, 267: 116806. |
| [36] | CHEN D D, XU S M, LI S L, et al. Directly evolved AlkS-based biosensor platform for monitoring and high-throughput screening of alkane production[J]. ACS Synthetic Biology, 2023, 12(3): 832-841. |
| [37] | PU W, CHEN J Z, LIU P, et al. Directed evolution of linker helix as an efficient strategy for engineering LysR-type transcriptional regulators as whole-cell biosensors[J]. Biosensors and Bioelectronics, 2023, 222: 115004. |
| [38] | TENG Y X, GONG X Y, ZHANG J L, et al. Investigating and engineering an 1, 2-propanediol-responsive transcription factor-based biosensor[J]. ACS Synthetic Biology, 2024, 13(7): 2177-2187. |
| [39] | ROTTINGHAUS A G, XI C G, AMROFELL M B, et al. Engineering ligand-specific biosensors for aromatic amino acids and neurochemicals[J]. Cell Systems, 2022, 13(3): 204-214.e4. |
| [40] | COULSON T J D, PATTEN C L. The TyrR transcription factor regulates the divergent akr-ipdC operons of Enterobacter cloacae UW5[J]. PLoS One, 2015, 10(3): e0121241. |
| [41] | D’OELSNITZ S, NGUYEN V, ALPER H S, et al. Evolving a generalist biosensor for bicyclic monoterpenes[J]. ACS Synthetic Biology, 2022, 11(1): 265-272. |
| [42] | DABIRIAN Y, LI X W, CHEN Y, et al. Expanding the dynamic range of a transcription factor-based biosensor in Saccharomyces cerevisiae [J]. ACS Synthetic Biology, 2019, 8(9): 1968-1975. |
| [43] | CHEN C, LIU J J, YAO G, et al. A novel, genetically encoded whole-cell biosensor for directed evolution of myrcene synthase in Escherichia coli [J]. Biosensors and Bioelectronics, 2023, 228: 115176. |
| [44] | LEBOVICH M, ANDREWS L B. Surveying the genetic design space for transcription factor-based metabolite biosensors: synthetic gamma-aminobutyric acid and propionate biosensors in E. coli Nissle 1917[J]. Frontiers in Bioengineering and Biotechnology, 2022, 10: 938056. |
| [45] | WEI W P, SHANG Y Z, ZHANG P, et al. Engineering prokaryotic transcriptional activator XylR as a xylose-inducible biosensor for transcription activation in yeast[J]. ACS Synthetic Biology, 2020, 9(5): 1022-1029. |
| [46] | BEABOUT K, EHRENWORTH BREEDON A M, BLUM S M, et al. Detection of bile acids in complex matrices using a transcription factor-based biosensor[J]. ACS Biomaterials Science & Engineering, 2023, 9(9): 5151-5162. |
| [47] | DING N N, YUAN Z Q, ZHANG X J, et al. Programmable cross-ribosome-binding sites to fine-tune the dynamic range of transcription factor-based biosensor[J]. Nucleic Acids Research, 2020, 48(18): 10602-10613. |
| [48] | LI C Y, ZHOU Y Y, ZOU Y S, et al. Identifying, characterizing, and engineering a phenolic acid-responsive transcriptional factor from Bacillus amyloliquefaciens [J]. ACS Synthetic Biology, 2023, 12(8): 2382-2392. |
| [49] | ZHAO J Y, SUN H H, WANG G G, et al. Engineering chimeric chemoreceptors and two-component systems for orthogonal and leakless biosensing of extracellular γ-aminobutyric acid[J]. Journal of Agricultural and Food Chemistry, 2024, 72(25): 14216-14228. |
| [50] | LI L, ZHANG Q Q, SHI R R, et al. Multidimensional combinatorial screening for high-level production of erythritol in Yarrowia lipolytica [J]. Bioresource Technology, 2024, 406: 131035. |
| [51] | LIU D, SICA M S, MAO J W, et al. A p-coumaroyl-CoA biosensor for dynamic regulation of naringenin biosynthesis in Saccharomyces cerevisiae [J]. ACS Synthetic Biology, 2022, 11(10): 3228-3238. |
| [52] | GONG X Y, ZHANG R H, WANG J, et al. Engineering of a TrpR-based biosensor for altered dynamic range and ligand preference[J]. ACS Synthetic Biology, 2022, 11(6): 2175-2183. |
| [53] | HAN L C, LIU X Y, CHENG Z Y, et al. Construction and application of a high-throughput in vivo screening platform for the evolution of nitrile metabolism-related enzymes based on a desensitized repressive biosensor[J]. ACS Synthetic Biology, 2022, 11(4): 1577-1587. |
| [54] | D’OELSNITZ S, KIM W, BURKHOLDER N T, et al. Using fungible biosensors to evolve improved alkaloid biosyntheses[J]. Nature Chemical Biology, 2022, 18(9): 981-989. |
| [55] | CHEN D D, ZHAO J D, XU S M, et al. Detection of short-chain chlorinated aliphatic hydrocarbons through an engineered biosensor with tailored ligand specificity[J]. Analytical Chemistry, 2024, 96(39): 15614-15623. |
| [56] | PHAM C, STOGIOS P J, SAVCHENKO A, et al. Computation-guided transcription factor biosensor specificity engineering for adipic acid detection[J]. Computational and Structural Biotechnology Journal, 2024, 23: 2211-2219. |
| [57] | KIM Y, JEON Y, SONG K, et al. Development of an Escherichia coli cell-based biosensor for aspirin monitoring by genetic engineering of MarR[J]. Biosensors, 2024, 14(11): 547. |
| [58] | PHAM C, STOGIOS P J, SAVCHENKO A, et al. Design and characterization of a generalist biosensor for indole derivatives[J]. ACS Synthetic Biology, 2024, 13(7): 2246-2252. |
| [59] | XIAO D, HU C X, XU X Z, et al. A D, L-lactate biosensor based on allosteric transcription factor LldR and amplified luminescent proximity homogeneous assay[J]. Biosensors and Bioelectronics, 2022, 211: 114378. |
| [60] | WANG Y, LI S X, XUE N, et al. Modulating sensitivity of an erythromycin biosensor for precise high-throughput screening of strains with different characteristics[J]. ACS Synthetic Biology, 2023, 12(6): 1761-1771. |
| [61] | KALKREUTER E, KEELER A M, MALICO A A, et al. Development of a genetically encoded biosensor for detection of polyketide synthase extender units in Escherichia coli [J]. ACS Synthetic Biology, 2019, 8(6): 1391-1400. |
| [62] | XU X H, LI X L, LIU Y F, et al. Pyruvate-responsive genetic circuits for dynamic control of central metabolism[J]. Nature Chemical Biology, 2020, 16(11): 1261-1268. |
| [63] | 杨璐, 吴楠, 白茸茸, 等. 基因回路型全细胞微生物传感器的设计、优化与应用[J]. 合成生物学, 2022, 3(6): 1061-1080. |
| YANG L, WU N, BAI R R, et al. Design, optimization and application of whole-cell microbial biosensors with engineered genetic circuits[J]. Synthetic Biology Journal, 2022, 3(6): 1061-1080. | |
| [64] | DING N N, YUAN Z N, MA Z, et al. AI-assisted rational design and activity prediction of biological elements for optimizing transcription-factor-based biosensors[J]. Molecules, 2024, 29(15): 3512. |
| [65] | 赵梅, 罗佳璐, 王震, 等. 基于转录因子的生物传感器研究进展[J]. 食品与发酵工业, 2024, 50(12): 362-369. |
| ZHAO M, LUO J L, WANG Z, et al. Research progress on transcription-factor-based biosensors[J]. Food and Fermentation Industries, 2024, 50(12): 362-369. | |
| [66] | ZHEN Z, XIANG L, LI S Z, et al. Designing a whole-cell biosensor applicable for S-adenosyl-L-methionine-dependent methyltransferases[J]. Biosensors and Bioelectronics, 2025, 268: 116904. |
| [67] | LI C, GAO X, QI H B, et al. Substantial improvement of an epimerase for the synthesis of D-allulose by biosensor-based high-throughput microdroplet screening[J]. Angewandte Chemie International Edition, 2023, 62(10): e202216721. |
| [68] | QIU X L, XU P, ZHAO X R, et al. Combining genetically-encoded biosensors with high throughput strain screening to maximize erythritol production in Yarrowia lipolytica [J]. Metabolic Engineering, 2020, 60: 66-76. |
| [69] | GAO J S, DU M H, ZHAO J H, et al. Design of a genetically encoded biosensor to establish a high-throughput screening platform for L-cysteine overproduction[J]. Metabolic Engineering, 2022, 73: 144-157. |
| [70] | TRIVEDI V D, MOHAN K, CHAPPELL T C, et al. Cheating the cheater: suppressing false-positive enrichment during biosensor-guided biocatalyst engineering[J]. ACS Synthetic Biology, 2022, 11(1): 420-429. |
| [71] | LI S S, LI Z L, TAN G Y, et al. In vitro allosteric transcription factor-based biosensing[J]. Trends in Biotechnology, 2023, 41(8): 1080-1095. |
| [72] | MORASKIE M, ROSHID M H O, O’CONNOR G, et al. Microbial whole-cell biosensors: current applications, challenges, and future perspectives[J]. Biosensors and Bioelectronics, 2021, 191: 113359. |
| [73] | 贾男, 臧国伟, 李春, 等. 辅因子在微生物细胞工厂中的代谢调控与应用[J]. 中国生物工程杂志, 2022, 42(7): 79-89. |
| JIA N, ZANG G W, LI C, et al. Metabolic regulations and applications of cofactors in microbial cell factories[J]. China Biotechnology, 2022, 42(7): 79-89. | |
| [74] | DONG C, SCHULTZ J C, LIU W, et al. Identification of novel metabolic engineering targets for S-adenosyl-L-methionine production in Saccharomyces cerevisiae via genome-scale engineering[J]. Metabolic Engineering, 2021, 66: 319-327. |
| [75] | LI X Y, ZHOU M H, ZENG D W, et al. Membrane transport engineering for efficient yeast biomanufacturing[J]. Bioresource Technology, 2025, 418: 131890. |
| [76] | WANG J, LI C Y, JIANG T, et al. Biosensor-assisted titratable CRISPRi high-throughput (BATCH) screening for over-production phenotypes[J]. Metabolic Engineering, 2023, 75: 58-67. |
| [77] | PENG Q Q, BAO W W, GENG B N, et al. Biosensor-assisted CRISPRi high-throughput screening to identify genetic targets in Zymomonas mobilis for high d-lactate production[J]. Synthetic and Systems Biotechnology, 2024, 9(2): 242-249. |
| [78] | BAUMANN L, BRUDER S, KABISCH J, et al. High-throughput screening of an octanoic acid producer strain library enables detection of new targets for increasing titers in Saccharomyces cerevisiae [J]. ACS Synthetic Biology, 2021, 10(5): 1077-1086. |
| [79] | STELLA R G, GERTZEN C G W, SMITS S H J, et al. Biosensor-based growth-coupling and spatial separation as an evolution strategy to improve small molecule production of Corynebacterium glutamicum [J]. Metabolic Engineering, 2021, 68: 162-173. |
| [80] | KRÜGER A, GÖDDECKE J, OSTHEGE M, et al. Biosensor-based growth-coupling as an evolutionary strategy to improve heme export in Corynebacterium glutamicum [J]. Microbial Cell Factories, 2024, 23(1): 276. |
| [81] | GEORGE K W, THOMPSON M G, KIM J, et al. Integrated analysis of isopentenyl pyrophosphate (IPP) toxicity in isoprenoid-producing Escherichia coli [J]. Metabolic Engineering, 2018, 47: 60-72. |
| [82] | LU L Y, WANG X L, WANG T, et al. A bacterial platform for producing aromatic esters from glycerol[J]. Nature Chemical Engineering, 2024, 1(12): 751-764. |
| [83] | ZHANG Q W, XU X H, ZHANG W, et al. De novo 2'-fucosyllactose biosynthesis using glucose as the sole carbon source by multiple engineered Bacillus subtilis [J]. Metabolic Engineering, 2025, 88: 85-93. |
| [84] | YU F, LI C Y, ZHANG T, et al. Developing a novel heme biosensor to produce high-active hemoproteins in Pichia pastoris through comparative transcriptomics[J]. Metabolic Engineering, 2024, 84: 59-68. |
| [85] | DING D Q, ZHU Y R, BAI D Y, et al. Monitoring and dynamically controlling glucose uptake rate and central metabolism[J]. Nature Chemical Engineering, 2025, 2(1): 50-62. |
| [86] | TONG Y J, LI N, ZHOU S H, et al. Improvement of Chalcone synthase activity and high-efficiency fermentative production of (2S)-naringenin via in vivo biosensor-guided directed evolution[J]. ACS Synthetic Biology, 2024, 13(5): 1454-1466. |
| [87] | LI M, CHEN Z Y, ZHANG W Y, et al. Customization of ethylene glycol (EG)-induced BmoR-based biosensor for the directed evolution of PET degrading enzymes[J]. Advanced Science, 2025, 12(13): e2413205. |
| [88] | DONG P Y, FAN Y J, HUO Y X, et al. Pathway-adapted biosensor for high-throughput screening of O-methyltransferase and its application in vanillin synthesis[J]. ACS Synthetic Biology, 2024, 13(9): 2873-2886. |
| [89] | OGAWA Y, SAITO Y, YAMAGUCHI H, et al. Engineering the substrate specificity of toluene degrading enzyme XylM using biosensor XylS and machine learning[J]. ACS Synthetic Biology, 2023, 12(2): 572-582. |
| [90] | BAHLS M O, PLATZ L, MORGADO G, et al. Directed evolution of biofuel-responsive biosensors for automated optimization of branched-chain alcohol biosynthesis[J]. Metabolic Engineering, 2022, 69: 98-111. |
| [91] | BAUMANN P T, MOLIN M DAL, ARING H, et al. Beyond rational-biosensor-guided isolation of 100 independently evolved bacterial strain variants and comparative analysis of their genomes[J]. BMC Biology, 2023, 21(1): 183. |
| [92] | YANG H N, HE Y C, ZHOU S H, et al. Dynamic regulation and cofactor engineering of Escherichia coli to enhance production of glycolate from corn stover hydrolysate[J]. Bioresource Technology, 2024, 398: 130531. |
| [93] | CHEN Z Y, YU S Z, LIU J, et al. Concentration recognition-based auto-dynamic regulation system (CRUISE) enabling efficient production of higher alcohols[J]. Advanced Science, 2024, 11(23): e2310215. |
| [94] | ZHOU S H, ZHANG Q Y, YUAN M W, et al. Static and dynamic regulation of precursor supply pathways to enhance raspberry ketone synthesis from glucose in Escherichia coli [J]. Journal of Agricultural and Food Chemistry, 2024, 72(42): 23411-23421. |
| [95] | YU H, LI F, WANG Y X, et al. Electro-controlled distribution of reducing equivalents to boost isobutanol biosynthesis in microbial electro-fermentation of S. oneidensis[J]. Joule, 2025, 9(1): 101773. |
| [96] | ZHAO X X, WU Y L, FENG T Y, et al. Dynamic upregulation of the rate-limiting enzyme for valerolactam biosynthesis in Corynebacterium glutamicum [J]. Metabolic Engineering, 2023, 77: 89-99. |
| [97] | SUN H H, ZHAO H M, ANG E L. A new biosensor for stilbenes and a cannabinoid enabled by genome mining of a transcriptional regulator[J]. ACS Synthetic Biology, 2020, 9(4): 698-705. |
| [98] | HANKO E K R, JOOSAB NOOR MAHOMED T A, STONEY R A, et al. TFBMiner: a user-friendly command line tool for the rapid mining of transcription factor-based biosensors[J]. ACS Synthetic Biology, 2023, 12(5): 1497-1507. |
| [99] | EKAS H M, WANG B, SILVERMAN A D, et al. An automated cell-free workflow for transcription factor engineering[J]. ACS Synthetic Biology, 2024, 13(10): 3389-3399. |
| [100] | LIU K, ZHANG Y S, LIU K, et al. De novo design of a transcription factor for a progesterone biosensor[J]. Biosensors and Bioelectronics, 2022, 203: 113897. |
| [101] | ZHAO M, HU M K, HAN R M, et al. Dynamics design of a non-natural transcription factor responding to androst-4-ene-3, 17-dione[J]. Synthetic and Systems Biotechnology, 2024, 9(3): 436-444. |
| [102] | RICHTER M F, ZHAO K T, ETON E, et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity[J]. Nature Biotechnology, 2020, 38(7): 883-891. |
| [103] | JUMPER J, EVANS R, PRITZEL A, et al. Highly accurate protein structure prediction with AlphaFold[J]. Nature, 2021, 596(7873): 583-589. |
| [104] | RUBE H T, RASTOGI C, FENG S Q, et al. Prediction of protein-ligand binding affinity from sequencing data with interpretable machine learning[J]. Nature Biotechnology, 2022, 40(10): 1520-1527. |
| [105] | ZHANG P C, WANG H C, XU H W, et al. Deep flanking sequence engineering for efficient promoter design using DeepSEED[J]. Nature Communications, 2023, 14: 6309. |
| [106] | BAYER T, HÄNEL L, HUSARCIKOVA J, et al. In vivo detection of low molecular weight platform chemicals and environmental contaminants by genetically encoded biosensors[J]. ACS Omega, 2023, 8(26): 23227-23239. |
| [107] | TELLECHEA-LUZARDO J, STIEBRITZ M T, CARBONELL P. Transcription factor-based biosensors for screening and dynamic regulation[J]. Frontiers in Bioengineering and Biotechnology, 2023, 11: 1118702. |
| [108] | NEUBAUER P, JUNNE S. Scale-down simulators for metabolic analysis of large-scale bioprocesses[J]. Current Opinion in Biotechnology, 2010, 21(1): 114-121. |
| [109] | 王晟, 王泽琛, 陈威华, 等. 基于人工智能和计算生物学的合成生物学元件设计[J]. 合成生物学, 2023, 4(3): 422-443. |
| WANG S, WANG Z C, CHEN W H, et al. Design of synthetic biology components based on artificial intelligence and computational biology[J]. Synthetic Biology Journal, 2023, 4(3): 422-443. | |
| [110] | ZHANG C, LIU H, LI X J, et al. Modularized synthetic biology enabled intelligent biosensors[J]. Trends in Biotechnology, 2023, 41(8): 1055-1065. |
| [111] | BOADA Y, VIGNONI A, PICÓ J, et al. Extended metabolic biosensor design for dynamic pathway regulation of cell factories[J]. iScience, 2020, 23(7): 101305. |
| [112] | ABRAMSON J, ADLER J, DUNGER J, et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3[J]. Nature, 2024, 630(8016): 493-500. |
| [113] | CHAN C T Y, KENNEDY V, KINSHUK S. A domain swapping strategy to create modular transcriptional regulators for novel topology in genetic network[J]. Biotechnology Advances, 2024, 72: 108345. |
| [114] | DEMEESTER W, DE BAETS J, DUCHI D, et al. MoBioS: modular platform technology for high-throughput construction and characterization of tunable transcriptional biological sensors[J]. Biosensors, 2023, 13(6): 590. |
| [115] | LEE H, XIE T, KANG B, et al. Plug-and-play protein biosensors using aptamer-regulated in vitro transcription[J]. Nature Communications, 2024, 15: 7973. |
| [1] | 鲁锦畅, 武耀康, 吕雪芹, 刘龙, 陈坚, 刘延峰. 神经酰胺类鞘脂的绿色生物制造[J]. 合成生物学, 2025, 6(2): 422-444. |
| [2] | 郭肖杰, 剪兴金, 王立言, 张翀, 邢新会. 合成生物学表型测试生物反应器及其装备化研究进展[J]. 合成生物学, 2024, 5(1): 16-37. |
| [3] | 晏雄鹰, 王振, 娄吉芸, 张皓瑜, 黄星宇, 王霞, 杨世辉. 生物燃料高效生产微生物细胞工厂构建研究进展[J]. 合成生物学, 2023, 4(6): 1082-1121. |
| [4] | 刘欢, 崔球. 原位电离质谱技术在微生物菌株筛选中的应用进展[J]. 合成生物学, 2023, 4(5): 980-999. |
| [5] | 吴玉洁, 刘欣欣, 刘健慧, 杨开广, 随志刚, 张丽华, 张玉奎. 基于高通量液相色谱质谱技术的菌株筛选与关键分子定量分析研究进展[J]. 合成生物学, 2023, 4(5): 1000-1019. |
| [6] | 秦伟彤, 杨广宇. 微液滴高通量筛选方法的研究与应用进展[J]. 合成生物学, 2023, 4(5): 966-979. |
| [7] | 孙梦楚, 陆亮宇, 申晓林, 孙新晓, 王佳, 袁其朋. 基于荧光检测的高通量筛选技术和装备助力细胞工厂构建[J]. 合成生物学, 2023, 4(5): 947-965. |
| [8] | 赵国淼, 杨鑫, 张媛, 王靖, 谭剑, 魏超, 周娜娜, 李凡, 王小艳. 生物设施平台及其工业应用[J]. 合成生物学, 2023, 4(5): 892-903. |
| [9] | 高纤云, 牛灵雪, 见妮, 管宁子. 微生物合成生物学在疾病诊疗上的应用进展[J]. 合成生物学, 2023, 4(2): 263-282. |
| [10] | 涂然, 李世新, 李昊霓, 王猛. 液滴微流控技术在微生物工程菌株选育中的应用进展[J]. 合成生物学, 2023, 4(1): 165-184. |
| [11] | 任师超, 孙秋艳, 冯旭东, 李春. 微生物细胞工厂合成五环三萜皂苷类化合物[J]. 合成生物学, 2022, 3(1): 168-183. |
| [12] | 熊亮斌, 宋璐, 赵云秋, 刘坤, 刘勇军, 王风清, 魏东芝. 甾体化合物绿色生物制造:从生物转化到微生物从头合成[J]. 合成生物学, 2021, 2(6): 942-963. |
| [13] | 郭亮, 高聪, 柳亚迪, 陈修来, 刘立明. 大肠杆菌生产饲用氨基酸的研究进展[J]. 合成生物学, 2021, 2(6): 964-981. |
| [14] | 陈久洲, 王钰, 蒲伟, 郑平, 孙际宾. 5-氨基乙酰丙酸生物合成技术的发展及展望[J]. 合成生物学, 2021, 2(6): 1000-1016. |
| [15] | 袁姚梦, 邢新会, 张翀. 微生物细胞工厂的设计构建:从诱变育种到全基因组定制化创制[J]. 合成生物学, 2020, 1(6): 656-673. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||