合成生物学 ›› 2023, Vol. 4 ›› Issue (4): 756-778.DOI: 10.12211/2096-8280.2023-032
代谢工程改造微生物利用甲酸研究进展
程真真1,2, 张健1,2, 高聪1,2, 刘立明1,2, 陈修来1,2
- 1.江南大学,食品科学与技术国家重点实验室,江苏 无锡 214122
2.江南大学,食品安全国际合作联合实验室,江苏 无锡 214122
-
收稿日期:2023-04-17修回日期:2023-05-22出版日期:2023-08-31发布日期:2023-09-14 -
通讯作者:陈修来 -
作者简介:程真真 (2001—),女,学士。研究方向为微生物固碳途径的设计、构建与应用。E-mail:zzch0929@163.com陈修来 (1985—),男,博士,教授。研究方向为微生物代谢工程与合成生物学。E-mail:xlchen@jiangnan.edu.cn -
基金资助:国家重点研发计划(2021YFC2103500);国家自然科学基金(22122806);江苏省自然科学基金(BK20211529);江南大学基本科研计划面上培育项目(JUSRP22031)
Progress in metabolic engineering of microorganisms for the utilization of formate
CHENG Zhenzhen1,2, ZHANG Jian1,2, GAO Cong1,2, LIU Liming1,2, CHEN Xiulai1,2
- 1.State Key Laboratory of Food Science and Technology,Jiangnan University,Wuxi 214122,Jiangsu,China
2.International Joint Laboratory on Food Safety,Jiangnan University,Wuxi 214122,Jiangsu,China
-
Received:2023-04-17Revised:2023-05-22Online:2023-08-31Published:2023-09-14 -
Contact:CHEN Xiulai
摘要:
微生物利用甲酸生产高附加值产品,是实现碳资源回收利用与绿色产业发展的重要策略之一。然而,在微生物利用甲酸过程中存在甲酸利用效率偏低、细胞生长速率缓慢、目标代谢物产量不高等问题。为了解决上述问题,本文从甲酸利用的微生物、代谢路径与代谢工程策略三个方面,系统总结分析了代谢工程改造微生物利用甲酸的研究进展。在甲酸利用微生物方面,概述了天然甲酸利用微生物的代谢特点以及模式微生物的代谢工程改造潜能;在甲酸利用代谢路径方面,梳理了天然的甲酸利用路径、重构与优化的甲酸利用路径与人工的甲酸利用路径的关键步骤、能量/还原力消耗与特点;在甲酸利用代谢工程策略方面,阐述了提高甲酸同化效率与改善甲酸利用微生物细胞生长的关键方法。最后,从甲酸利用的微生物、代谢路径与代谢工程策略三个方面,展望了微生物利用甲酸的发展方向,为甲酸生物经济的发展奠定了基础。
中图分类号:
引用本文
程真真, 张健, 高聪, 刘立明, 陈修来. 代谢工程改造微生物利用甲酸研究进展[J]. 合成生物学, 2023, 4(4): 756-778.
CHENG Zhenzhen, ZHANG Jian, GAO Cong, LIU Liming, CHEN Xiulai. Progress in metabolic engineering of microorganisms for the utilization of formate[J]. Synthetic Biology Journal, 2023, 4(4): 756-778.
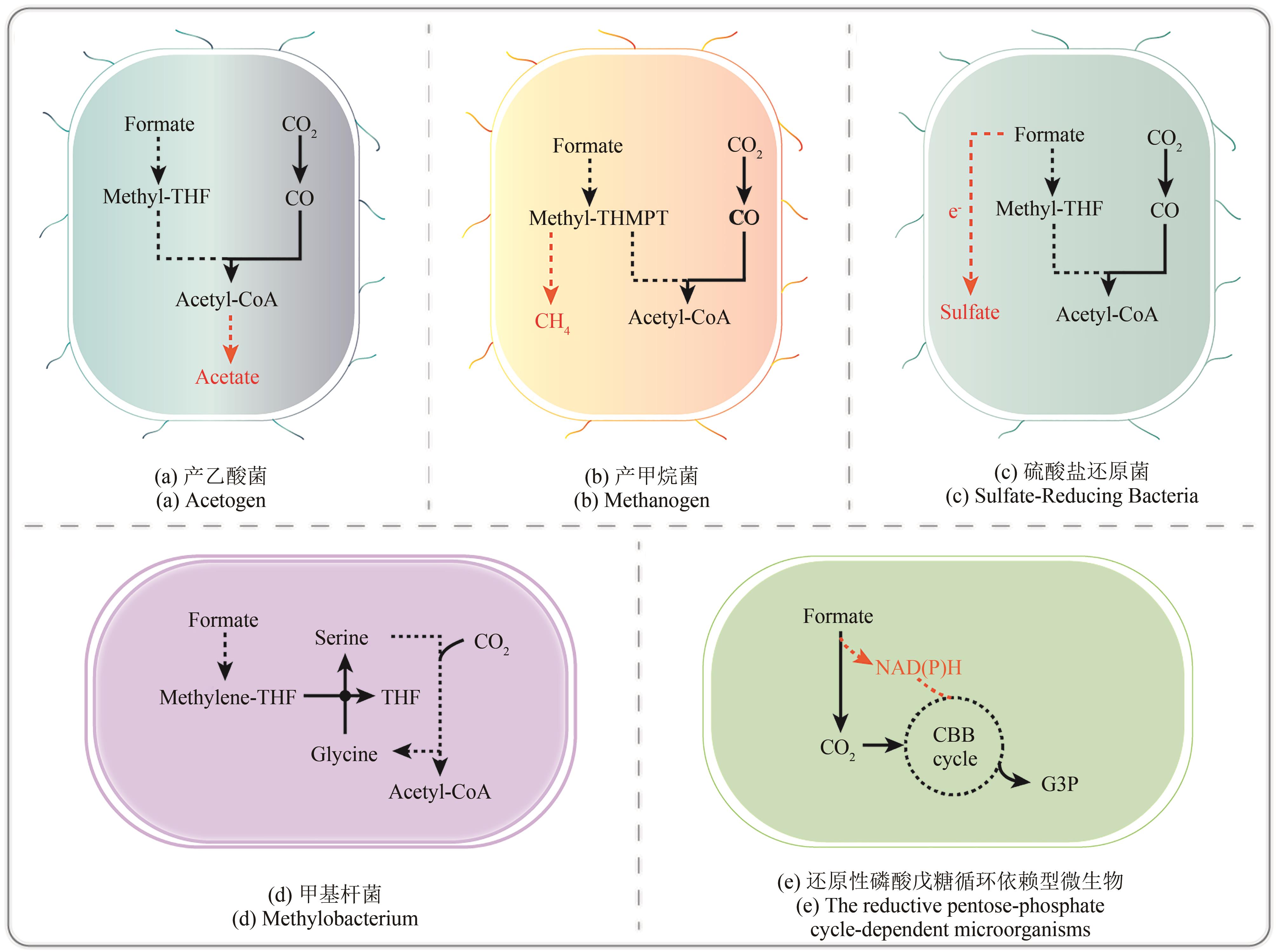
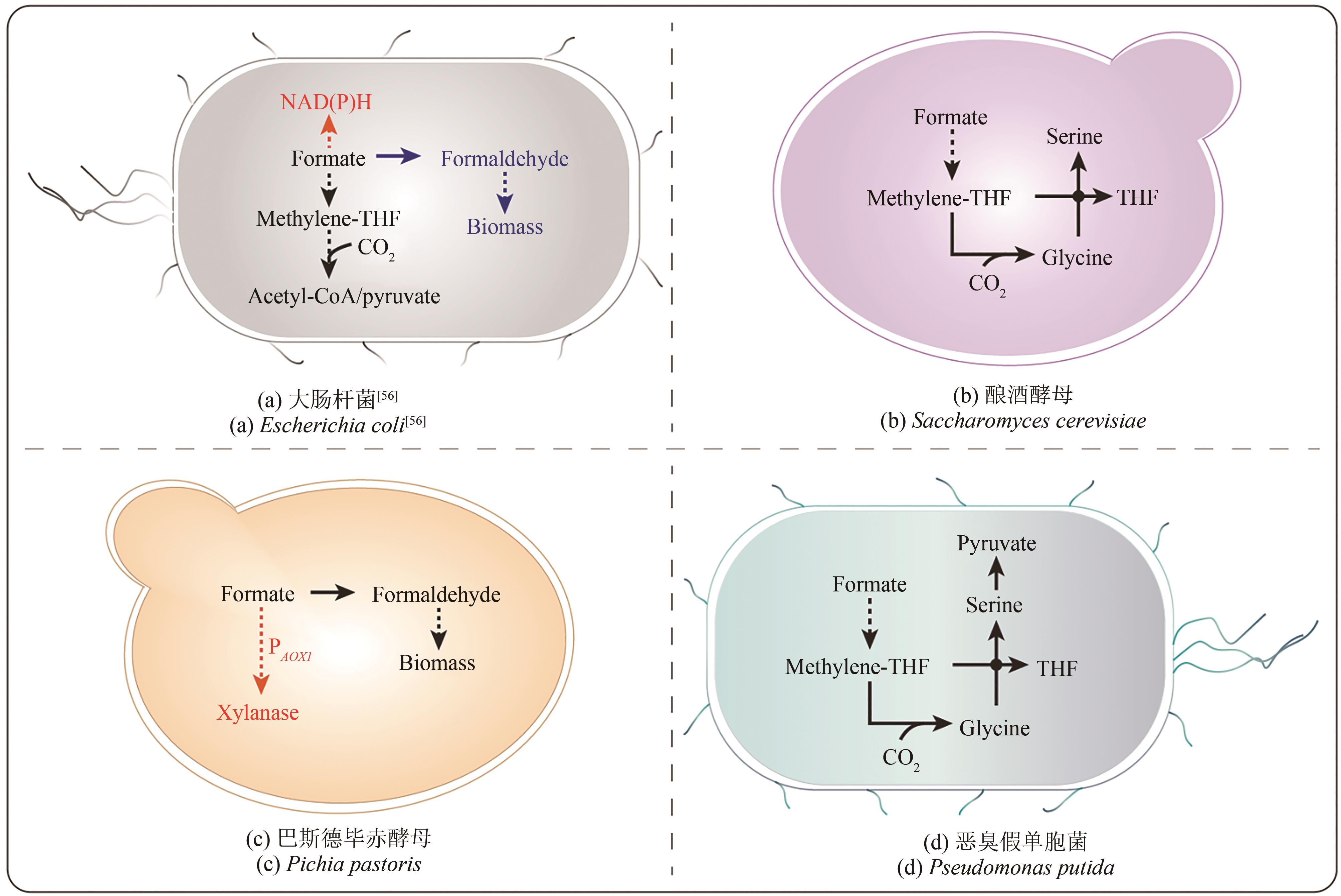
图1 天然甲酸利用微生物CBB cycle—Calvin-Benson-Bassham循环;THF—四氢叶酸;THMPT—四氢甲基蝶呤
Fig. 1 Natural formate-utilizing microorganismsCBB cycle—Calvin-Benson-Bassham cycle; THF—tetrahydrofolate; THMPT—tetrahydromethanopterin
| 微生物种类 | 需氧类型 | 模式菌株 | 天然甲酸 代谢路径 | 应用优劣势 | 参考 文献 |
|---|---|---|---|---|---|
| 产乙酸菌 | 专性厌氧 | 伍氏醋酸杆菌 | 还原性乙酰辅酶A途径 | 优势:能够天然利用甲酸和CO2合成乙酸、乙醇等化合物 劣势:在厌氧条件下生长,不利于工业应用;代谢研究尚不完善,代谢改造工具少 | [ |
| 产甲烷菌 | 专性厌氧 | 氢营养型海沼甲烷球菌 | 还原性乙酰辅酶A途径 | 优势:能够天然利用甲酸和CO2合成甲烷,在清洁能源产业方面具有应用潜能 劣势:在厌氧条件下生长,不利于工业应用;代谢研究尚不完善,代谢改造工具少 | [ |
| 硫酸盐还原菌 | 专型厌氧 | 普通脱硫弧菌 | 还原性乙酰辅酶A途径 | 优势:具有非常高的氢化酶活性,能够天然利用甲酸产氢,在清洁能源产业方面具有应用潜能 劣势:厌氧条件下生长,不利于工业应用;代谢研究尚不完善,代谢改造工具少 | [ |
| 甲基杆菌 | 好氧或 兼性厌氧 | 扭脱甲基杆菌AM1 | 丝氨酸循环 | 优势:能够天然利用甲酸、CO2与甲醇等一碳底物进行细胞生长;模式菌株的研究与改造相对充分,具有一定的改造潜能 劣势:所具有的甲酸代谢路径的能量利用效率偏低,不利于工业应用 | [ |
| CBB循环 依赖型微生物 | 好氧 | 钩虫贪铜菌H16 | 还原性磷酸戊糖循环 | 优势:具有甲酸脱氢酶,能够利用甲酸作为唯一碳源和能源;模式菌株的研究与改造相对充分,具有一定的应用潜能 劣势:甲酸的氧化供能存在能量浪费,因此微生物的细胞生长速率与工业生产潜能均偏低 | [ |
表1 天然甲酸利用微生物
Table 1 Natural formate-utilizing microorganisms
| 微生物种类 | 需氧类型 | 模式菌株 | 天然甲酸 代谢路径 | 应用优劣势 | 参考 文献 |
|---|---|---|---|---|---|
| 产乙酸菌 | 专性厌氧 | 伍氏醋酸杆菌 | 还原性乙酰辅酶A途径 | 优势:能够天然利用甲酸和CO2合成乙酸、乙醇等化合物 劣势:在厌氧条件下生长,不利于工业应用;代谢研究尚不完善,代谢改造工具少 | [ |
| 产甲烷菌 | 专性厌氧 | 氢营养型海沼甲烷球菌 | 还原性乙酰辅酶A途径 | 优势:能够天然利用甲酸和CO2合成甲烷,在清洁能源产业方面具有应用潜能 劣势:在厌氧条件下生长,不利于工业应用;代谢研究尚不完善,代谢改造工具少 | [ |
| 硫酸盐还原菌 | 专型厌氧 | 普通脱硫弧菌 | 还原性乙酰辅酶A途径 | 优势:具有非常高的氢化酶活性,能够天然利用甲酸产氢,在清洁能源产业方面具有应用潜能 劣势:厌氧条件下生长,不利于工业应用;代谢研究尚不完善,代谢改造工具少 | [ |
| 甲基杆菌 | 好氧或 兼性厌氧 | 扭脱甲基杆菌AM1 | 丝氨酸循环 | 优势:能够天然利用甲酸、CO2与甲醇等一碳底物进行细胞生长;模式菌株的研究与改造相对充分,具有一定的改造潜能 劣势:所具有的甲酸代谢路径的能量利用效率偏低,不利于工业应用 | [ |
| CBB循环 依赖型微生物 | 好氧 | 钩虫贪铜菌H16 | 还原性磷酸戊糖循环 | 优势:具有甲酸脱氢酶,能够利用甲酸作为唯一碳源和能源;模式菌株的研究与改造相对充分,具有一定的应用潜能 劣势:甲酸的氧化供能存在能量浪费,因此微生物的细胞生长速率与工业生产潜能均偏低 | [ |
| 微生物 | 非天然甲酸代谢路径 | 主要的应用优势 | 改造潜能 | 参考文献 |
|---|---|---|---|---|
| 大肠杆菌 | 重构的卡尔文循环; 改良的丝氨酸循环; 高丝氨酸循环; rTHF-rgcv途径及关键模块 合成乙酰辅酶A途径等 | 生长周期短; 遗传背景清晰; 代谢工程改造工具种类丰富且高效 | 设计并构建更加高效的甲酸利用路径; 通过实验室适应性进化与培养条件优化等策略提高菌株对甲酸的耐受性 | [ |
| 酿酒酵母 | rTHF-rgcv途径 的关键模块 | 遗传背景清晰; 分子遗传操作工具与技术成熟且高效; 具有内源性甲酸脱氢酶,对甲酸的耐受性较高 | 通过代谢改造进一步提高rTHF-rgcv途径对甲酸的利用效率,开发以甲酸为唯一碳源和能源的工程菌株 | [ |
| 毕赤酵母 | 以甲酸作为P AOX1 的诱导物 | 能够利用甲醇作为唯一碳源和能源进行细胞生长; 在生物医药产业和工业酶生产方面具有巨大潜力 | 通过代谢改造进一步提高菌株对甲酸的利用效率; 基于菌株的代谢网络,设计并构建更加高效的甲酸利用路径 | [ |
| 恶臭假单胞菌 | rTHF-rgcv途径 | 能够编码多种天然甲酸脱氢酶; 具有灵活的代谢机制,能够抵抗氧化应激和多种有毒化合物 | 开发更高效的分子遗传操作工具,进一步完善菌株的甲酸代谢网络 | [ |
表2 代谢工程改造的微生物
Table 2 Metabolically engineered microorganisms
| 微生物 | 非天然甲酸代谢路径 | 主要的应用优势 | 改造潜能 | 参考文献 |
|---|---|---|---|---|
| 大肠杆菌 | 重构的卡尔文循环; 改良的丝氨酸循环; 高丝氨酸循环; rTHF-rgcv途径及关键模块 合成乙酰辅酶A途径等 | 生长周期短; 遗传背景清晰; 代谢工程改造工具种类丰富且高效 | 设计并构建更加高效的甲酸利用路径; 通过实验室适应性进化与培养条件优化等策略提高菌株对甲酸的耐受性 | [ |
| 酿酒酵母 | rTHF-rgcv途径 的关键模块 | 遗传背景清晰; 分子遗传操作工具与技术成熟且高效; 具有内源性甲酸脱氢酶,对甲酸的耐受性较高 | 通过代谢改造进一步提高rTHF-rgcv途径对甲酸的利用效率,开发以甲酸为唯一碳源和能源的工程菌株 | [ |
| 毕赤酵母 | 以甲酸作为P AOX1 的诱导物 | 能够利用甲醇作为唯一碳源和能源进行细胞生长; 在生物医药产业和工业酶生产方面具有巨大潜力 | 通过代谢改造进一步提高菌株对甲酸的利用效率; 基于菌株的代谢网络,设计并构建更加高效的甲酸利用路径 | [ |
| 恶臭假单胞菌 | rTHF-rgcv途径 | 能够编码多种天然甲酸脱氢酶; 具有灵活的代谢机制,能够抵抗氧化应激和多种有毒化合物 | 开发更高效的分子遗传操作工具,进一步完善菌株的甲酸代谢网络 | [ |
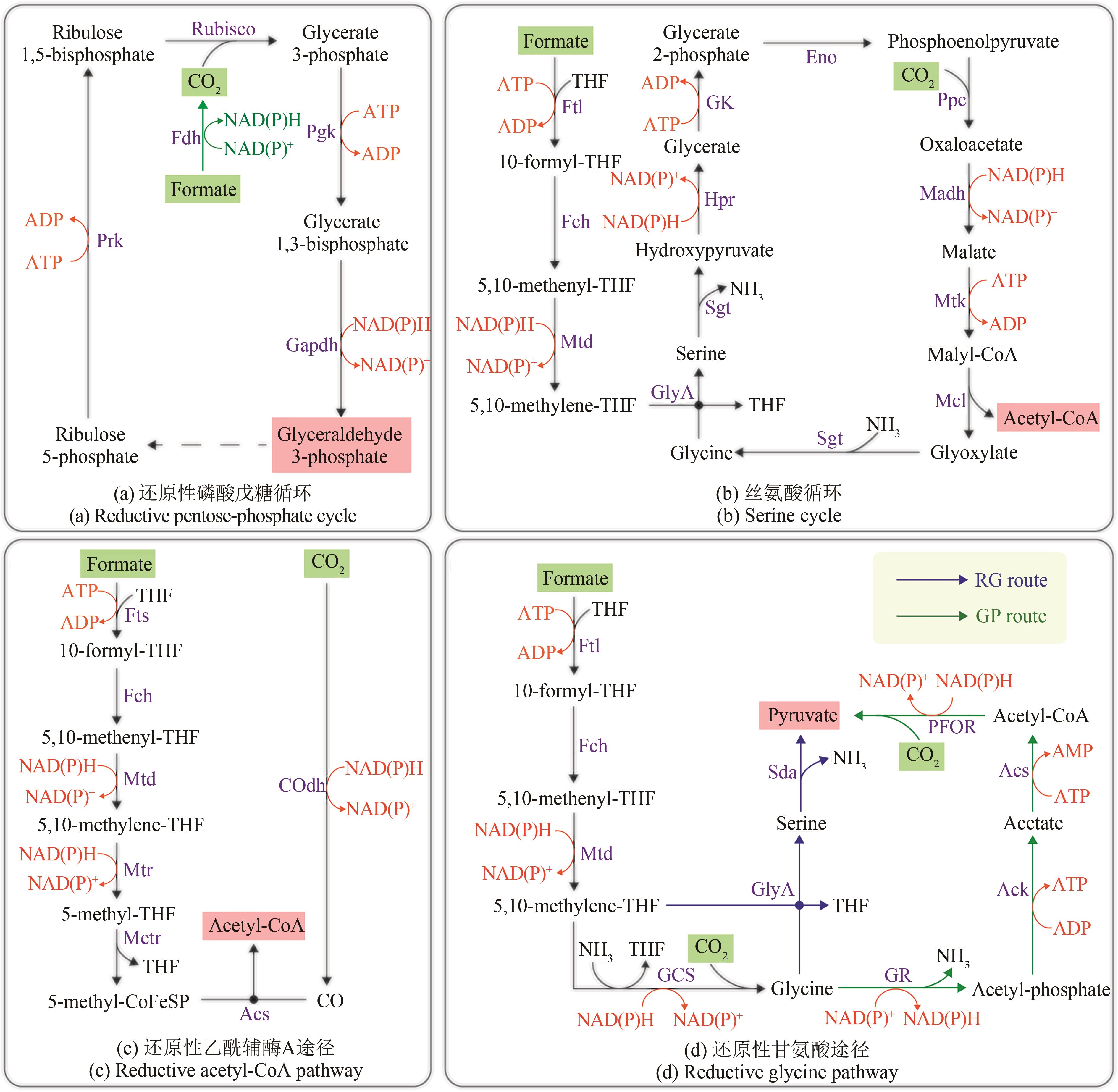
图3 天然甲酸利用路径(绿色底色的化合物为路径底物,粉色底色的化合物为路径产物)Ack—乙酸激酶;Acs—乙酰辅酶A合成酶;COdh—CO脱氢酶;CoFeSP—钴铁硫蛋白;Eno—烯醇化酶;Fch—次甲基四氢叶酸环水解酶;Fdh—甲酸脱氢酶;Ftl—甲酸四氢叶酸连接酶;Fts—甲酰四氢叶酸合成酶;Gapdh—3-磷酸甘油醛脱氢酶;GCS—甘氨酸裂解系统;Gk—甘油酸激酶;GlyA—丝氨酸羟甲基转移酶;GR—甘氨酸还原酶复合体;Hpr—羟基丙酮还原酶;Madh—苹果酸脱氢酶;Mcl—苹果酰辅酶A裂解酶;Metr—甲基转移酶;Mtd—亚甲基四氢叶酸脱氢酶;Mtk—苹果酸裂解酶;Mtr—亚甲基四氢叶酸还原酶;PFOR—丙酮酸铁氧还蛋白氧化还原酶;Pgk—磷酸甘油酸激酶;Ppc—PEP羧化酶;Prk—磷酸核酮糖激酶;Rubisco—核酮糖-1,5-二磷酸羧化酶/加氧酶;Sda—丝氨酸脱氨酶;Sgt—丝氨酸-乙醛酸转氨酶;THF—四氢叶酸
Fig. 3 Natural formate-utilizing pathways(compounds with a green background are pathway substrates; compounds with a pink background are pathway products) Ack—Acetate kinase; Acs—acetyl-CoA synthase; COdh—CO dehydrogenase; CoFeSP—corrinoid ironsulfur protein; Eno—enolase; Fch—methenyl-tetrahydrofolate cyclohydrolase; Fdh—formate dehydrogenase; Ftl—formate-tetrahydrofolate ligase; Fts—formyl-tetrahydrofolate synthetase; Gapdh—glyceraldehyde-3-phosphate dehydrogenase; GCS—glycine cleavage system; Gk—glycerate kinase; GlyA—serine hydroxymethyltransferase; GR—Glycine reductase complex; Hpr—hydroxypyruvate reductase; Madh—malate dehydrogenase; Mcl—malyl-CoA lyase; Metr—methyltransferase; Mtd—methylene-tetrahydrofolate dehydrogenase; Mtk—malate thiokinase; Mtr—methylene-tetrahydrofolate reductase; PFOR—Pyruvate ferredoxin oxidoreductase; Pgk—Phosphoglycerate kinase; Ppc—phosphoenolpyruvate carboxylase; Prk—phosphoribulo kinase; Rubisco—Ribulose-1,5-bisphosphate carboxylase/oxygenase; Sda—serine deaminase; Sgt—serine-glyoxylate transaminase; THF—tetrahydrofolate
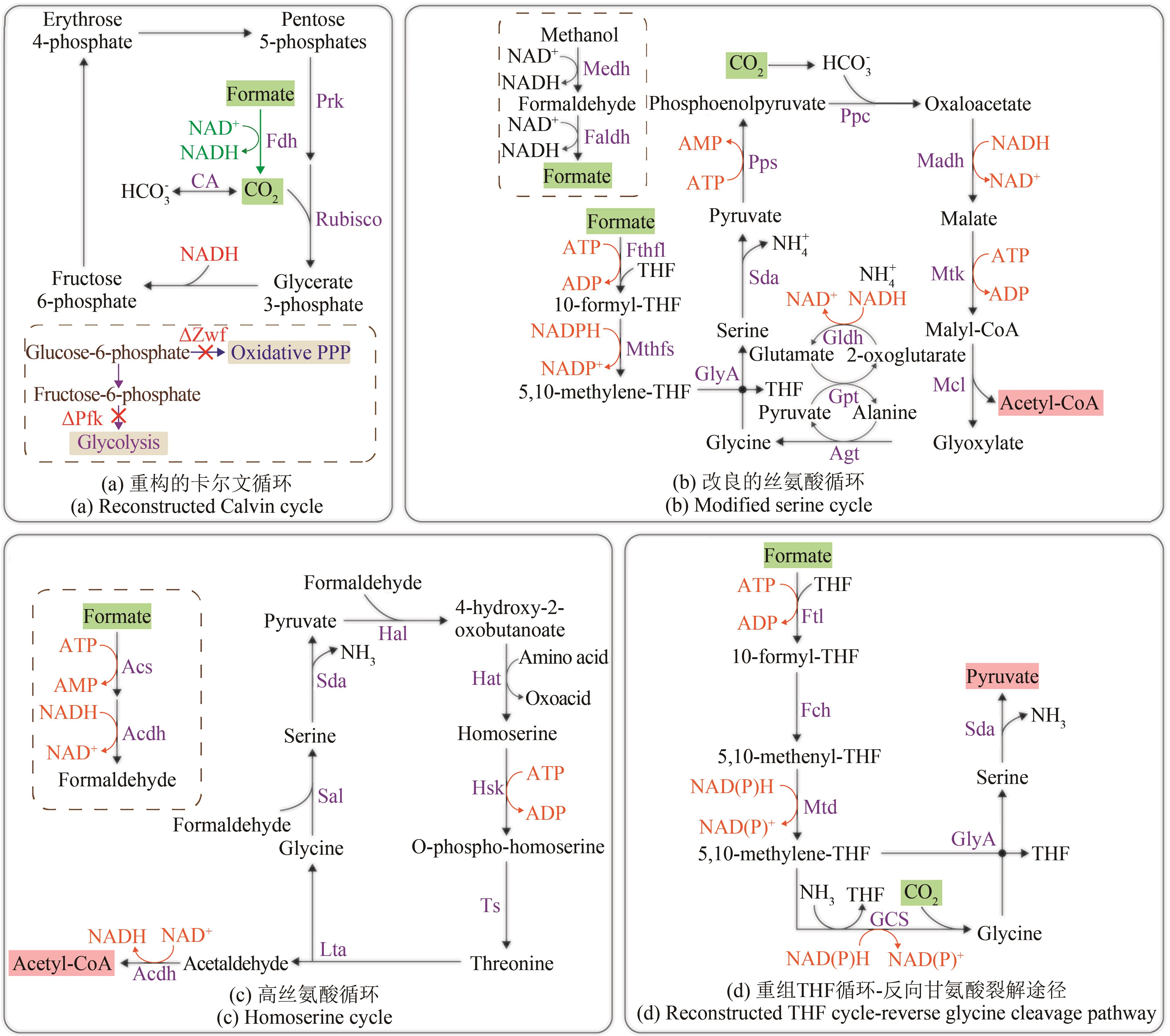
图4 重构优化甲酸利用路径(绿色底色的化合物为路径底物,粉色底色的化合物为路径产物)Acdh—乙醛脱氢酶;Acs—乙酰辅酶A合成酶;Agt—丙氨酸-乙醛酸转氨酶;CA—碳酸酐酶;Faldh—甲醛脱氢酶;Fch—次甲基四氢叶酸环水解酶;Fdh—甲酸脱氢酶;Fthfl—甲酸四氢叶酸连接酶;Ftl—甲酸四氢叶酸连接酶;GCS—甘氨酸裂解系统;Gldh—谷氨酸脱氢酶;GlyA—丝氨酸羟甲基转移酶;Gpt—谷氨酸-丙酮酸转氨酶;Hal—HOB醛缩酶;Hat—HOB转氨酶;Hsk—高丝氨酸激酶;Lta—苏氨酸醛缩酶;Madh—苹果酸脱氢酶;Mcl—苹果酰辅酶A裂解酶;Medh—甲醇脱氢酶;Mtd—亚甲基四氢叶酸脱氢酶;Mthfs—5,10-亚甲基四氢叶酸合成酶;Mtk—苹果酸硫激酶;Oxidative PPP—氧化戊糖磷酸途径;Pfk—磷酸果糖激酶;Ppc—PEP羧化酶;Pps—PEP合成酶;Prk—磷酸核酮糖激酶;Rubisco—核酮糖-1,5-二磷酸羧化酶;Sal—丝氨酸醛缩酶;Sda—丝氨酸脱氨酶;THF—四氢叶酸;Ts—苏氨酸合成酶;Zwf—6-磷酸葡萄糖脱氢酶
Fig. 4 Reconstruction and optimization of formate-utilizing pathways(compounds with a green background are pathway substrates; compounds with a pink background are pathway products) Acdh—acetaldehyde dehydrogenase; Acs—acetyl-CoA synthase; Agt—alanine-glyoxylate transaminase; CA—carbonic anhydrase; Faldh—formaldehyde dehydrogenase; Fch—methenyl-tetrahydrofolate cyclohydrolase; Fdh—formate dehydrogenase; Fthfl—formate-tetrahydrofolate ligase; Ftl—formate-tetrahydrofolate ligase; GCS—glycine cleavage system; Gldh—glutamate dehydrogenase; GlyA—serine hydroxymethyltransferase; Gpt—glutamate-pyruvate transaminase; Hal—4-hydroxy-2-oxobutanoate aldolase; Hat—4-hydroxy-2-oxobutanoate aminotransferase; Hsk—homoserine kinase; Lta—threonine aldolase; Madh—malate dehydrogenase; Mcl—malyl-CoA lyase; Medh—methanol dehydrogenase; Mtd—methylene-tetrahydrofolate dehydrogenase; Mthfs—5,10-methylene-tetrahydrofolate synthase; Mtk—malate thiokinase; Oxidative PPP—Oxidative pentose-phosphate pathway; Pfk—phosphofructokinase; Ppc—phosphoenolpyruvate carboxylase; Pps—phosphoenolpyruvate synthase; Prk—phosphoribulo kinase; Rubisco—ribulose-1,5-bisphosphate carboxylase; Sal—serine aldolase; Sda—serine deaminase; THF—tetrahydrofolate; Ts—threonine synthase; Zwf—glucose-6-phosphate dehydrogenase
| 菌株 | 路径酶来源 | 参考文献 |
|---|---|---|
| Escherichia coli | Ftl、Fch、Mtd(源自Clostridium ljungdahalii) GCS、GlyA、Sda(内源酶) | [ |
| Escherichia coli | Ftl、Fch、Mtd(源自Methylobacterium extorquens AM1) GCS、GlyA(内源酶) | [ |
| Escherichia coli | Ftl、Fch、Mtd(源自Methylobacterium extorquens CM4) GCS、GlyA、Sda(内源酶) Fdh(源自Candida boidinii) | [ |
| Escherichia coli | Ftl(源自Clostridium kluyveri) FolD(Fch/ Mtd)、GCS、GlyA(内源酶) | [ |
| Escherichia coli | Ftl、Fch、Mtd(源自Methylobacterium extorquens AM1) GCS、GlyA、Sda(内源酶) Fdh(源自Pseudomonas sp.) | [ |
Escherichia coli (Bang等[ | Ftl、Fch、Mtd(源自Methylobacterium extorquens CM4) GCS、GlyA、Sda(内源酶) Fdh(源自Candida boidinii、Arabidopsis thaliana) | [ |
Escherichia coli (Döring等[ | Ftl(源自Clostridium kluyveri) FolD(Fch/ Mtd)、GCS、GlyA(内源酶) | [ |
Escherichia coli (Kim等[ | Ftl、Fch、Mtd(源自Methylobacterium extorquens AM1) GCS、GlyA、Sda(内源酶) Fdh(源自Pseudomonas sp. ) | [ |
| Saccharomyces cerevisiae | MIS1(Ftl/ Fch/ Mtd)、GCS(内源酶) Fdh(内源酶) | [ |
| Cupriavidus necator | Ftl、Fch、Mtd(源自Methylobacterium extorquens AM1) GCS、GlyA、Sda(内源酶) Fdh(内源酶) | [ |
| Clostridium pasteurianum | GCS(源自Gottschalkia acidurici) Ftl、Fch、Mtd、GlyA、Sda(内源酶) | [ |
| Pseudomonas putida | Ftl、Fch、Mtd(源自Methylobacterium extorquens AM1) GCS、GlyA、Sda(内源酶) Fdh(内源酶) | [ |
表3 rTHF-rgcv途径的路径酶来源
Table 3 Source of pathway enzymes of the rTHF-rgcv pathway
| 菌株 | 路径酶来源 | 参考文献 |
|---|---|---|
| Escherichia coli | Ftl、Fch、Mtd(源自Clostridium ljungdahalii) GCS、GlyA、Sda(内源酶) | [ |
| Escherichia coli | Ftl、Fch、Mtd(源自Methylobacterium extorquens AM1) GCS、GlyA(内源酶) | [ |
| Escherichia coli | Ftl、Fch、Mtd(源自Methylobacterium extorquens CM4) GCS、GlyA、Sda(内源酶) Fdh(源自Candida boidinii) | [ |
| Escherichia coli | Ftl(源自Clostridium kluyveri) FolD(Fch/ Mtd)、GCS、GlyA(内源酶) | [ |
| Escherichia coli | Ftl、Fch、Mtd(源自Methylobacterium extorquens AM1) GCS、GlyA、Sda(内源酶) Fdh(源自Pseudomonas sp.) | [ |
Escherichia coli (Bang等[ | Ftl、Fch、Mtd(源自Methylobacterium extorquens CM4) GCS、GlyA、Sda(内源酶) Fdh(源自Candida boidinii、Arabidopsis thaliana) | [ |
Escherichia coli (Döring等[ | Ftl(源自Clostridium kluyveri) FolD(Fch/ Mtd)、GCS、GlyA(内源酶) | [ |
Escherichia coli (Kim等[ | Ftl、Fch、Mtd(源自Methylobacterium extorquens AM1) GCS、GlyA、Sda(内源酶) Fdh(源自Pseudomonas sp. ) | [ |
| Saccharomyces cerevisiae | MIS1(Ftl/ Fch/ Mtd)、GCS(内源酶) Fdh(内源酶) | [ |
| Cupriavidus necator | Ftl、Fch、Mtd(源自Methylobacterium extorquens AM1) GCS、GlyA、Sda(内源酶) Fdh(内源酶) | [ |
| Clostridium pasteurianum | GCS(源自Gottschalkia acidurici) Ftl、Fch、Mtd、GlyA、Sda(内源酶) | [ |
| Pseudomonas putida | Ftl、Fch、Mtd(源自Methylobacterium extorquens AM1) GCS、GlyA、Sda(内源酶) Fdh(内源酶) | [ |
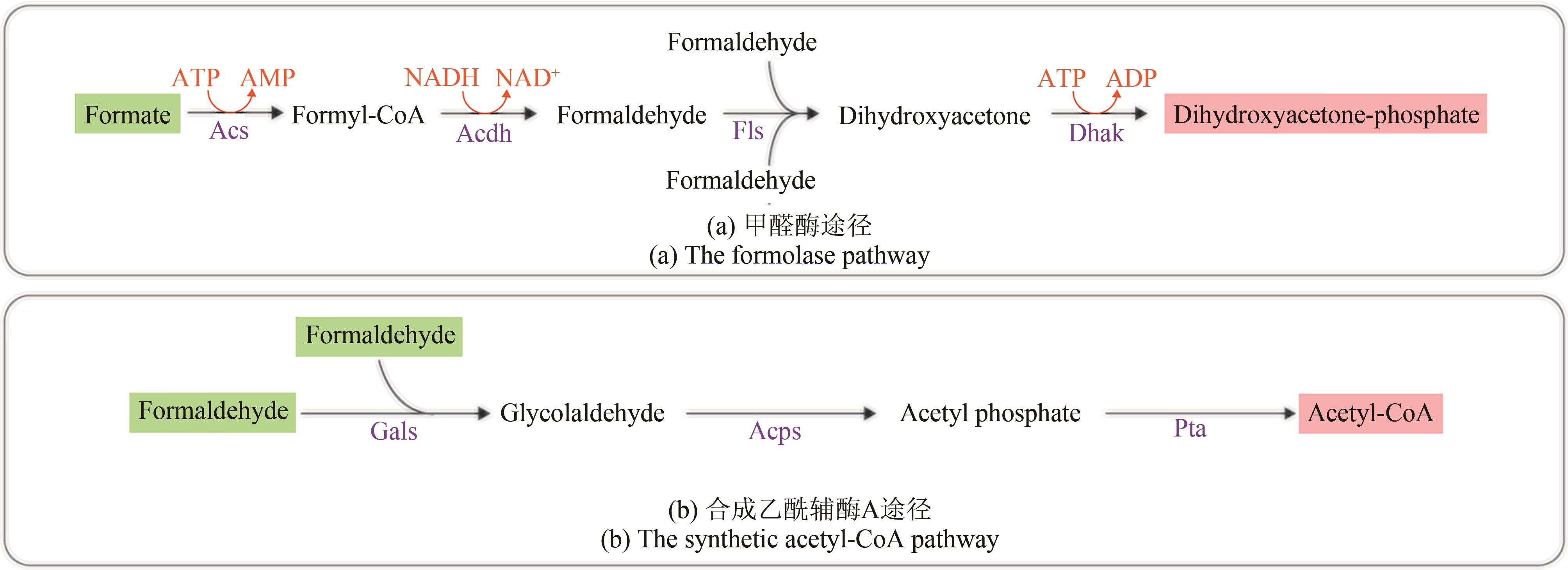
图5 人工设计甲酸利用路径Acdh—乙醛脱氢酶;Acs—乙酰辅酶A合成酶;Acps—乙酰磷酸合成酶;Dhak—二羟基丙酮激酶;Fls—甲醛酶;Gals—乙醇醛合成酶;Pta—磷酸转乙酰酶(绿色底色的化合物为路径底物,粉色底色的化合物为路径产物)
Fig. 5 Artificial formate-utilizing pathwaysAcdh—acetaldehyde dehydrogenase; Acs—acetyl-CoA synthase; Acps—acetyl phosphate synthase; Dhak—dihydroxyacetone kinase; Fls—formolase; Gals—glycolaldehyde synthase; Pta—phosphate acetyltransferase (compounds with a green background are pathway substrates, compounds with a pink background are pathway products)

图6 提高甲酸同化效率的关键方法FDH—甲酸脱氢酶;Hint—H蛋白的氨甲基化形式;Hox—H蛋白的氧化形式;Hred—H蛋白的还原形式;TCA cycle—三羧酸循环;THF—四氢叶酸
Fig. 6 Key methods to improve the efficiency of formate assimilationFDH—formate dehydrogenase; Hint—aminomethylated form of H protein; Hox—oxidized form of H protein; Hred—reduced form of H protein; TCA cycle—tricarboxylic acid cycle; THF—tetrahydrofolate;
| 1 | ESCOBAR J C, LORA E S, VENTURINI O J, et al. Biofuels: environment, technology and food security[J]. Renewable & Sustainable Energy Reviews, 2009, 13(6/7): 1275-1287. |
| 2 | ZHOU Y J, KERKHOVEN E J, NIELSEN J. Barriers and opportunities in bio-based production of hydrocarbons[J]. Nature Energy, 2018, 3(11): 925-935. |
| 3 | LV X Q, YU W W, ZHANG C Y, et al. C1-based biomanufacturing: advances, challenges and perspectives[J]. Bioresource Technology, 2023, 367: 128259. |
| 4 | ZHANG C Q, OTTENHEIM C, WEINGARTEN M, et al. Microbial utilization of next-generation feedstocks for the biomanufacturing of value-added chemicals and food ingredients[J]. Frontiers in Bioengineering and Biotechnology, 2022, 10: 874612. |
| 5 | SAKARIKA M, GANIGUÉ R, RABAEY K. Methylotrophs: from C1 compounds to food[J]. Current Opinion in Biotechnology, 2022, 75: 102685. |
| 6 | FLAIZ M, LUDWIG G, BENGELSDORF F R, et al. Production of the biocommodities butanol and acetone from methanol with fluorescent FAST-tagged proteins using metabolically engineered strains of Eubacterium limosum [J]. Biotechnology for Biofuels, 2021, 14(1): 117. |
| 7 | OKOYE-CHINE C G, OTUN K, SHIBA N, et al. Conversion of carbon dioxide into fuels—a review[J]. Journal of CO2 Utilization, 2022, 62: 102099. |
| 8 | YOON J H, CHANG W J, OH S H, et al. Metabolic engineering of Methylorubrum extorquens AM1 for poly (3-hydroxybutyrate-co-3-hydroxyvalerate) production using formate[J]. International Journal of Biological Macromolecules, 2021, 177: 284-293. |
| 9 | RAY S, JIN J O, CHOI I, et al. Recent trends of biotechnological production of polyhydroxyalkanoates from C1 carbon sources[J]. Frontiers in Bioengineering and Biotechnology, 2023, 10: 907500. |
| 10 | LIU Y Q, BAI C X, XU Q, et al. Improved methanol-derived lovastatin production through enhancement of the biosynthetic pathway and intracellular lovastatin efflux in methylotrophic yeast[J].Bioresources and Bioprocessing, 2018, 5(1): 22. |
| 11 | LIU Y Q, TU X H, XU Q, et al. Engineered monoculture and co-culture of methylotrophic yeast for de novo production of monacolin J and lovastatin from methanol[J]. Metabolic Engineering, 2018, 45: 189-199. |
| 12 | YASIN M, JEONG Y S, PARK S Y, et al. Microbial synthesis gas utilization and ways to resolve kinetic and mass-transfer limitations[J]. Bioresource Technology, 2015, 177: 361-374. |
| 13 | COTTON C A, CLAASSENS N J, BENITO-VAQUERIZO S, et al. Renewable methanol and formate as microbial feedstocks[J]. Current Opinion in Biotechnology, 2020, 62: 168-180. |
| 14 | JIANG W, HERNÁNDEZ VILLAMOR D, PENG H D, et al. Metabolic engineering strategies to enable microbial utilization of C1 feedstocks[J]. Nature Chemical Biology, 2021, 17(8): 845-855. |
| 15 | THIJS B, RONGÉ J, MARTENS J A. Matching emerging formic acid synthesis processes with application requirements[J]. Green Chemistry, 2022, 24(6): 2287-2295. |
| 16 | CLAASSENS N J, BORDANABA-FLORIT G, COTTON C A R, et al. Replacing the Calvin cycle with the reductive glycine pathway in Cupriavidus necator [J]. Metabolic Engineering, 2020, 62: 30-41. |
| 17 | KIM S, LINDNER S N, ASLAN S, et al. Growth of E. coli on formate and methanol via the reductive glycine pathway[J]. Nature Chemical Biology, 2020, 16(5): 538-545. |
| 18 | BANG J, HWANG C H, AHN J H, et al. Escherichia coli is engineered to grow on CO2 and formic acid[J]. Nature Microbiology, 2020, 5(12): 1459-1463. |
| 19 | GONZALEZ DE LA CRUZ J, MACHENS F, MESSERSCHMIDT K, et al. Core catalysis of the reductive glycine pathway demonstrated in yeast[J]. ACS Synthetic Biology, 2019, 8(5): 911-917. |
| 20 | DRAKE H L, KÜSEL K, MATTHIES C. Acetogenic prokaryotes[M/OL]//The Prokaryotes. Berlin, Heidelberg: Springer Berlin Heidelberg, 2013: 3-60 [2023-03-01]. . |
| 21 | MOON J, DÖNIG J, KRAMER S, et al. Formate metabolism in the acetogenic bacterium Acetobacterium woodii [J]. Environmental Microbiology, 2021, 23(8): 4214-4227. |
| 22 | NEUENDORF C S, VIGNOLLE G A, DERNTL C, et al. A quantitative metabolic analysis reveals Acetobacterium woodii as a flexible and robust host for formate-based bioproduction[J]. Metabolic Engineering, 2021, 68: 68-85. |
| 23 | REEVE J N. Molecular biology of methanogens[J]. Annual Review of Microbiology, 1992, 46: 165-191. |
| 24 | LUPA B, HENDRICKSON E L, LEIGH J A, et al. Formate-dependent H2 production by the mesophilic methanogen Methanococcus maripaludis [J]. Applied and Environmental Microbiology, 2008, 74(21): 6584-6590. |
| 25 | SARMIENTO F B, LEIGH J A, WHITMAN W B. Genetic systems for hydrogenotrophic methanogens[M/OL]//Methods in Methane Metabolism, Part A: Methods in Enzymology. Amsterdam: Elsevier, 2011: 43-73 [2023-03-01]. . |
| 26 | GOYAL N, ZHOU Z, KARIMI I A. Metabolic processes of Methanococcus maripaludis and potential applications[J]. Microbial Cell Factories, 2016, 15(1): 107. |
| 27 | MARTINS M, PEREIRA I A C. Sulfate-reducing bacteria as new microorganisms for biological hydrogen production[J]. International Journal of Hydrogen Energy, 2013, 38(28): 12294-12301. |
| 28 | RITTMANN S K M R, LEE H S, LIM J K, et al. One-carbon substrate-based biohydrogen production: microbes, mechanism, and productivity[J]. Biotechnology Advances, 2015, 33(1): 165-177. |
| 29 | MARTINS M, MOURATO C, PEREIRA I A C. Desulfovibrio vulgaris growth coupled to formate-driven H2 production[J]. Environmental Science & Technology, 2015, 49(24): 14655-14662. |
| 30 | SINGH R P, SINGH R N, SRIVASTAVA M K, et al. Structure prediction and analysis of MxaF from obligate, facultative and restricted facultative methylobacterium[J]. Bioinformation, 2012, 8(21): 1042-1046. |
| 31 | CROWTHER G J, KOSÁLY G, LIDSTROM M E. Formate as the main branch point for methylotrophic metabolism in Methylobacterium extorquens AM1[J]. Journal of Bacteriology, 2008, 190(14): 5057-5062. |
| 32 | CUI L Y, YANG J, LIANG W F, et al. Sodium formate redirects carbon flux and enhances heterologous mevalonate production in Methylobacterium extorquens AM1[J]. Biotechnology Journal, 2023, 18(2): 2200402. |
| 33 | POHLMANN A, FRICKE W F, REINECKE F, et al. Genome sequence of the bioplastic-producing "Knallgas" bacterium Ralstonia eutropha H16[J]. Nature Biotechnology, 2006, 24(10): 1257-1262. |
| 34 | CRAMM R. Genomic view of energy metabolism in Ralstonia eutropha H16[J]. Journal of Molecular Microbiology and Biotechnology, 2008, 16(1/2): 38-52. |
| 35 | PAN H J, WANG J, WU H L, et al. Synthetic biology toolkit for engineering Cupriviadus necator H16 as a platform for CO2 valorization[J].Biotechnology for Biofuels, 2021, 14(1): 212. |
| 36 | CALVEY C H, SANCHEZ I N V, WHITE A M, et al. Improving growth of Cupriavidus necator H16 on formate using adaptive laboratory evolution-informed engineering[J]. Metabolic Engineering, 2023, 75: 78-90. |
| 37 | SCHUCHMANN K, MÜLLER V. Autotrophy at the thermodynamic limit of life: a model for energy conservation in acetogenic bacteria[J]. Nature Reviews Microbiology, 2014, 12(12): 809-821. |
| 38 | LIU Y C, WHITMAN W B. Metabolic, phylogenetic, and ecological diversity of the methanogenic Archaea[J]. Annals of the New York Academy of Sciences, 2008, 1125(1): 171-189. |
| 39 | 冷欢, 杨清, 黄钢锋, 等. 氢营养型产甲烷代谢途径研究进展[J]. 微生物学报, 2020, 60(10): 2136-2160. |
| LENG H, YANG Q, HUANG G F, et al. Recent advances in hydrogenotrophic methanogenesis[J]. Acta Microbiologica Sinica, 2020, 60(10): 2136-2160. | |
| 40 | THAUER R K, KASTER A K, SEEDORF H, et al. Methanogenic Archaea: ecologically relevant differences in energy conservation[J]. Nature Reviews Microbiology, 2008, 6(8): 579-591. |
| 41 | COSTA K C, WONG P M, WANG T S, et al. Protein complexing in a methanogen suggests electron bifurcation and electron delivery from formate to heterodisulfide reductase[J]. Proceedings of the National Academy of Sciences of the United States of America, 2010, 107(24): 11050-11055. |
| 42 | LI J, ZHANG L Y, XU Q, et al. CRISPR-Cas9 toolkit for genome editing in an autotrophic CO2-fixing methanogenic archaeon[J]. Microbiology Spectrum, 2022, 10(4): 01165-22. |
| 43 | BAO J C, DE DIOS MATEOS E, SCHELLER S. Efficient CRISPR/Cas12a-based genome-editing toolbox for metabolic engineering in Methanococcus maripaludis [J]. ACS Synthetic Biology, 2022, 11(7): 2496-2503. |
| 44 | HEIDELBERG J F, SESHADRI R, HAVEMAN S A, et al. The genome sequence of the anaerobic, sulfate-reducing bacterium Desulfovibrio vulgaris hildenborough[J]. Nature Biotechnology, 2004, 22(5): 554-559. |
| 45 | JANSEN K, THAUER R K, WIDDEL F, et al. Carbon assimilation pathways in sulfate reducing bacteria. Formate, carbon dioxide, carbon monoxide, and acetate assimilation by Desulfovibrio baarsii [J]. Archives of Microbiology, 1984, 138(3): 257-262. |
| 46 | SÁNCHEZ-ANDREA I, ALVES GUEDES I, HORNUNG B, et al. The reductive glycine pathway allows autotrophic growth of Desulfovibrio desulfuricans [J]. Nature Communications, 2020, 11: 5090. |
| 47 | SUN H, SPRING S, LAPIDUS A, et al. Complete genome sequence of Desulfarculus baarsii type strain (2st14T)[J]. Standards in Genomic Sciences, 2010, 3(3): 276-284. |
| 48 | MARTINS M, MOURATO C, MORAIS-SILVA F O, et al. Electron transfer pathways of formate-driven H2 production in Desulfovibrio [J]. Applied Microbiology and Biotechnology, 2016, 100(18): 8135-8146. |
| 49 | ZHANG W M, SONG M, YANG Q, et al. Current advance in bioconversion of methanol to chemicals[J]. Biotechnology for Biofuels, 2018, 11: 260. |
| 50 | KIM S M, LEE S H, KIM I K, et al. Structural insight into a molecular mechanism of methenyltetrahydrofolate cyclohydrolase from Methylobacterium extorquens AM1[J]. International Journal of Biological Macromolecules, 2022, 202: 234-240. |
| 51 | TONG S, ZHAO L Z, ZHU D L, et al. From formic acid to single-cell protein: genome-scale revealing the metabolic network of Paracoccus communis MA5[J].Bioresources and Bioprocessing, 2022, 9(1): 55. |
| 52 | PICKERING B S, ORESNIK I J. Formate-dependent autotrophic growth in Sinorhizobium meliloti [J]. Journal of Bacteriology, 2008, 190(19): 6409-6418. |
| 53 | SOROKIN D Y, LÜCKER S, VEJMELKOVA D, et al. Nitrification expanded: discovery, physiology and genomics of a nitrite-oxidizing bacterium from the Phylum chloroflexi [J]. The ISME Journal, 2012, 6(12): 2245-2256. |
| 54 | GROSTERN A, ALVAREZ-COHEN L. RubisCO-based CO2 fixation and C1 metabolism in the actinobacterium Pseudonocardia dioxanivorans CB1190[J]. Environmental Microbiology, 2013, 15(11): 3040-3053. |
| 55 | ORLOVA M V, TARLACHKOV S V, DUBININA G A, et al. Genomic insights into metabolic versatility of a lithotrophic sulfur-oxidizing diazotrophic Alphaproteobacterium Azospirillum thiophilum [J]. FEMS Microbiology Ecology, 2016, 92(12): fiw199. |
| 56 | BANG J, AHN J H, LEE J A, et al. Synthetic formatotrophs for one-carbon biorefinery[J]. Advanced Science, 2021, 8(12): 2100199. |
| 57 | GLEIZER S, BEN-NISSAN R, BAR-ON Y M, et al. Conversion of Escherichia coli to generate all biomass carbon from CO2 [J]. Cell, 2019, 179(6): 1255-1263.e12. |
| 58 | YU H, LIAO J C. A modified serine cycle in Escherichia coli coverts methanol and CO2 to two-carbon compounds[J]. Nature Communications, 2018, 9: 3992. |
| 59 | HE H, HOPER R, DODENHOFT M, et al. An optimized methanol assimilation pathway relying on promiscuous formaldehyde-condensing aldolases in E. coli [J]. Metabolic Engineering, 2020, 60: 1-13. |
| 60 | YISHAI O, GOLDBACH L, TENENBOIM H, et al. Engineered assimilation of exogenous and endogenous formate in Escherichia coli [J]. ACS Synthetic Biology, 2017, 6(9): 1722-1731. |
| 61 | TASHIRO Y, HIRANO S, MATSON M M, et al. Electrical-biological hybrid system for CO2 reduction[J]. Metabolic Engineering, 2018, 47: 211-218. |
| 62 | YISHAI O, BOUZON M, DÖRING V, et al. In vivo assimilation of one-carbon via a synthetic reductive glycine pathway in Escherichia coli [J]. ACS Synthetic Biology, 2018, 7(9): 2023-2028. |
| 63 | BANG J, LEE S Y. Assimilation of formic acid and CO2 by engineered Escherichia coli equipped with reconstructed one-carbon assimilation pathways[J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(40): E9271-E9279. |
| 64 | DÖRING V, DARII E, YISHAI O, et al. Implementation of a reductive route of one-carbon assimilation in Escherichia coli through directed evolution[J]. ACS Synthetic Biology, 2018, 7(9): 2029-2036. |
| 65 | DELMAS V A, PERCHAT N, MONET O, et al. Genetic and biocatalytic basis of formate dependent growth of Escherichia coli strains evolved in continuous culture[J]. Metabolic Engineering, 2022, 72: 200-214. |
| 66 | KIM S, GIRALDO N, RAINALDI V, et al. Optimizing E. coli as a formatotrophic platform for bioproduction via the reductive glycine pathway[J]. Frontiers in Bioengineering and Biotechnology, 2023, 11: 1091899. |
| 67 | LU X Y, LIU Y W, YANG Y Q, et al. Constructing a synthetic pathway for acetyl-coenzyme A from one-carbon through enzyme design[J]. Nature Communications, 2019, 10: 1378. |
| 68 | LIU B, LI H J, ZHOU H L, et al. Enhancing xylanase expression by Komagataella phaffii by formate as carbon source and inducer[J].Applied Microbiology and Biotechnology, 2022, 106(23): 7819-7829. |
| 69 | LIU B, ZHAO Y X, ZHOU H L, et al. Enhancing xylanase expression of Komagataella phaffii induced by formate through Mit1 co-expression[J].Bioprocess and Biosystems Engineering, 2022, 45(9): 1515-1525. |
| 70 | TURLIN J, DRONSELLA B, DE MARIA A, et al. Integrated rational and evolutionary engineering of genome-reduced Pseudomonas putida strains promotes synthetic formate assimilation[J]. Metabolic Engineering, 2022, 74: 191-205. |
| 71 | BAR-EVEN A, NOOR E, FLAMHOLZ A, et al. Design and analysis of metabolic pathways supporting formatotrophic growth for electricity-dependent cultivation of microbes[J]. Biochimica et Biophysica Acta (BBA)-Bioenergetics, 2013, 1827(8/9): 1039-1047. |
| 72 | BAR-EVEN A, FLAMHOLZ A, NOOR E, et al. Thermodynamic constraints shape the structure of carbon fixation pathways[J]. Biochimica et Biophysica Acta (BBA) - Bioenergetics, 2012, 1817(9): 1646-1659. |
| 73 | KIM S J, YOON J, IM D K, et al. Adaptively evolved Escherichia coli for improved ability of formate utilization as a carbon source in sugar-free conditions[J].Biotechnology for Biofuels, 2019, 12:207. |
| 74 | NITSCHKE W, RUSSELL M J. Beating the acetyl coenzyme A-pathway to the origin of life[J]. Philosophical Transactions of the Royal Society of London Series B, Biological Sciences, 2013, 368(1622): 20120258. |
| 75 | YISHAI O, LINDNER S N, GONZALEZ DE LA CRUZ J, et al. The formate bio-economy[J]. Current Opinion in Chemical Biology, 2016, 35: 1-9. |
| 76 | LIU Z H, WANG K, CHEN Y, et al. Third-generation biorefineries as the means to produce fuels and chemicals from CO2 [J]. Nature Catalysis, 2020, 3(3): 274-288. |
| 77 | SONG Y, LEE J S, SHIN J, et al. Functional cooperation of the glycine synthase-reductase and Wood-Ljungdahl pathways for autotrophic growth of Clostridium drakei [J]. Proceedings of the National Academy of Sciences of the United States of America, 2020, 117(13): 7516-7523. |
| 78 | MAO W, YUAN Q Q, QI H G, et al. Recent progress in metabolic engineering of microbial formate assimilation[J].Applied Microbiology and Biotechnology, 2020, 104(16): 6905-6917. |
| 79 | ZHANG H, LI Y C, NIE J L, et al. Structure-based dynamic analysis of the glycine cleavage system suggests key residues for control of a key reaction step[J]. Communications Biology, 2020, 3: 756. |
| 80 | XU Y Y, REN J, WANG W, et al. Improvement of glycine biosynthesis from one-carbon compounds and ammonia catalyzed by the glycine cleavage system in vitro [J]. Engineering in Life Sciences, 2022, 22(1): 40-53. |
| 81 | ERB T J, KELLER P, VORHOLT J A. Escherichia coli in auto(trophic) mode[J]. Cell, 2019, 179(6): 1244-1245. |
| 82 | ZHANG Y X, LI F L, DONG J, et al. Recent advances in designing efficient electrocatalysts for electrochemical carbon dioxide reduction to formic acid/formate[J]. Journal of Electroanalytical Chemistry, 2023, 928: 117018. |
| 83 | JUNQUEIRA J R C, DAS D, CATHRIN BRIX A, et al. Simultaneous anodic and cathodic formate production in a paired electrolyzer by CO2 reduction and glycerol oxidation[J]. ChemSusChem, 2023: e202300667. |
| 84 | WANG T, CHEN J D, REN X Y, et al. Halogen-incorporated Sn catalysts for selective electrochemical CO2 reduction to formate[J]. Angewandte Chemie International Edition, 2023, 62(10): e202211174. |
| 85 | BAR-EVEN A. Formate assimilation: the metabolic architecture of natural and synthetic pathways[J]. Biochemistry, 2016, 55(28): 3851-3863. |
| 86 | CHEN N H, DJOKO K Y, VEYRIER F J, et al. Formaldehyde stress responses in bacterial pathogens[J]. Frontiers in Microbiology, 2016, 7: 257. |
| 87 | ALKIM C, FARIAS D, FREDONNET J, et al. Toxic effect and inability of L-homoserine to be a nitrogen source for growth of Escherichia coli resolved by a combination of in vivo evolution engineering and omics analyses[J]. Frontiers in Microbiology, 2022, 13: 1051425. |
| 88 | KLEIN V J, IRLA M, GIL LÓPEZ M, et al. Unravelling formaldehyde metabolism in bacteria: road towards synthetic methylotrophy[J]. Microorganisms, 2022, 10(2): 220. |
| 89 | HONG Y, ARBTER P, WANG W, et al. Introduction of glycine synthase enables uptake of exogenous formate and strongly impacts the metabolism in Clostridium pasteurianum [J]. Biotechnology and Bioengineering, 2021, 118(3): 1366-1380. |
| 90 | KHANA D B, CALLAGHAN M M, AMADOR-NOGUEZ D. Novel computational and experimental approaches for investigating the thermodynamics of metabolic networks[J]. Current Opinion in Microbiology, 2022, 66: 21-31. |
| 91 | CHEN K, ARNOLD F H. Engineering new catalytic activities in enzymes[J]. Nature Catalysis, 2020, 3(3): 203-213. |
| 92 | KIM D H, NOH M H, PARK M H, et al. Enzyme activity engineering based on sequence co-evolution analysis[J]. Metabolic Engineering, 2022, 74: 49-60. |
| 93 | SIEGEL J B, SMITH A L, POUST S, et al. Computational protein design enables a novel one-carbon assimilation pathway[J]. Proceedings of the National Academy of Sciences of the United States of America, 2015, 112(12): 3704-3709. |
| 94 | HU G P, LI Z H, MA D L, et al. Light-driven CO2 sequestration in Escherichia coli to achieve theoretical yield of chemicals[J]. Nature Catalysis, 2021, 4(5): 395-406. |
| 95 | HU G P, GUO L, GAO C, et al. Synergistic metabolism of glucose and formate increases the yield of short-chain organic acids in Escherichia coli [J]. ACS Synthetic Biology, 2022, 11(1): 135-143. |
| 96 | CLAASSENS N J, HE H, BAR-EVEN A. Synthetic methanol and formate assimilation via modular engineering and selection strategies[J]. Current Issues in Molecular Biology, 2019, 33(1): 237-248. |
| 97 | DRAGOSITS M, MATTANOVICH D. Adaptive laboratory evolution—principles and applications for biotechnology[J]. Microbial Cell Factories, 2013, 12: 64. |
| 98 | AHN J H, BANG J, KIM W J, et al. Formic acid as a secondary substrate for succinic acid production by metabolically engineered Mannheimia succiniciproducens [J]. Biotechnology and Bioengineering, 2017, 114(12): 2837-2847. |
| 99 | SINGH A, MOESTEDT J, BERG A, et al. Microbiological surveillance of biogas plants: targeting acetogenic community[J]. Frontiers in Microbiology, 2021, 12: 700256. |
| 100 | SUDA K, SAKAMOTO S, IGUCHI A, et al. Novel quantitative method for individual isotopomer of organic acids from 13C tracer experiments determines carbon flow in acetogenesis[J]. Talanta, 2023, 257: 124328. |
| 101 | KIRST H, FERLEZ B H, LINDNER S N, et al. Toward a glycyl radical enzyme containing synthetic bacterial microcompartment to produce pyruvate from formate and acetate[J]. Proceedings of the National Academy of Sciences of the United States of America, 2022, 119(8): e2116871119. |
| 102 | NA D, YOO S M, CHUNG H, et al. Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs[J]. Nature Biotechnology, 2013, 31(2): 170-174. |
| 103 | PONATH F, HÖR J, VOGEL J. An overview of gene regulation in bacteria by small RNAs derived from mRNA 3′ ends[J]. FEMS Microbiology Reviews, 2022, 46(5): fuac017. |
| 104 | WENK S, SCHANN K, HE H, et al. An "energy-auxotroph" Escherichia coli provides an in vivo platform for assessing NADH regeneration systems[J]. Biotechnology and Bioengineering, 2020, 117(11): 3422-3434. |
| [1] | 郭姝媛, 张倩楠, 姑丽克孜·买买提热夏提, 杨一群, 于涛. 液体生物燃料合成与炼制的研究进展[J]. 合成生物学, 2025, 6(1): 18-44. |
| [2] | 邵明威, 孙思勉, 杨时茂, 陈国强. 基于极端微生物的生物制造[J]. 合成生物学, 2024, 5(6): 1419-1436. |
| [3] | 赵亮, 李振帅, 付丽平, 吕明, 王士安, 张全, 刘立成, 李福利, 刘自勇. 生物转化一碳化合物原料产油脂与单细胞蛋白研究进展[J]. 合成生物学, 2024, 5(6): 1300-1318. |
| [4] | 竺方欢, 岑雪聪, 陈振. 微生物合成二元醇研究进展[J]. 合成生物学, 2024, 5(6): 1367-1385. |
| [5] | 陈雨, 张康, 邱以婧, 程彩云, 殷晶晶, 宋天顺, 谢婧婧. 微生物电合成技术转化二氧化碳研究进展[J]. 合成生物学, 2024, 5(5): 1142-1168. |
| [6] | 禹伟, 高教琪, 周雍进. 一碳生物转化合成有机酸的研究进展[J]. 合成生物学, 2024, 5(5): 1169-1188. |
| [7] | 陈锡玮, 张华然, 邹懿. 真菌源非核糖体肽类药物生物合成及代谢工程[J]. 合成生物学, 2024, 5(3): 571-592. |
| [8] | 虞旭昶, 吴辉, 李雷. 文库构建与基因簇靶向筛选驱动的微生物天然产物高效发现[J]. 合成生物学, 2024, 5(3): 492-506. |
| [9] | 谢皇, 郑义蕾, 苏依婷, 阮静怡, 李永泉. 放线菌聚酮类化合物生物合成体系重构研究进展[J]. 合成生物学, 2024, 5(3): 612-630. |
| [10] | 雷茹, 陶慧, 刘天罡. 基因组深度挖掘驱动微生物萜类化合物高效发现[J]. 合成生物学, 2024, 5(3): 507-526. |
| [11] | 惠真, 唐啸宇. CRISPR/Cas9编辑系统在微生物天然产物研究中的应用[J]. 合成生物学, 2024, 5(3): 658-671. |
| [12] | 赵静宇, 张健, 祁庆生, 王倩. 基于细菌双组分系统的生物传感器的研究进展[J]. 合成生物学, 2024, 5(1): 38-52. |
| [13] | 周强, 周大伟, 孙敬翔, 王靖楠, 姜万奎, 章文明, 蒋羽佳, 信丰学, 姜岷. 微生物发酵法合成虾青素的研究进展[J]. 合成生物学, 2024, 5(1): 126-143. |
| [14] | 高春辉, 杨宁, 王创, 侯时季, 严建兵, 郑金水, 李锦, 吴尘聊, 蔡鹏. 有涌现性功能的合成菌群在作物育种上的应用前景[J]. 合成生物学, 2024, 5(1): 144-153. |
| [15] | 朱华伟, 李寅. 生物光伏:环境友好的新型太阳能利用技术[J]. 合成生物学, 2023, 4(6): 1259-1280. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||