合成生物学 ›› 2021, Vol. 2 ›› Issue (4): 509-527.DOI: 10.12211/2096-8280.2021-041
纪念王义翘教授:解脂耶氏酵母替代植物油脂的技术瓶颈及展望
徐鹏1,2
- 1.广东以色列理工学院化学工程系,广东 汕头 515063
2.马里兰大学巴尔的摩分校化学、生化与环境工程系,马里兰州 巴尔的摩 21250,美国
-
收稿日期:2021-04-03修回日期:2021-05-12出版日期:2021-08-31发布日期:2021-09-10 -
作者简介:徐鹏 (1980—),男,工学博士,于2020年获得“生物技术与生物工程王义翘教授奖” (Biotechnology & Bioengineering Daniel I.C. Wang Award)。研究方向主要涉及代谢工程、合成生物学、生化工程、生物分子绿色制造、智能控制和生化反应网络建模等。E-mail:peng.xu@gtiit.edu.cn
In memory of Prof. Daniel I.C. Wang: Engineering Yarrowia lipolytica for the production of plant-based lipids: technical constraints and perspectives for a sustainable cellular agriculture economy
XU Peng1,2
- 1.Department of Chemical Engineering,Guangdong-Technion,Israel Institute of Technology,Shantou 515063,Guangdong,China
2.Department of Chemical,Biochemical and Environmental Engineering,University of Maryland,Baltimore County,Baltimore,MD 21250,USA
-
Received:2021-04-03Revised:2021-05-12Online:2021-08-31Published:2021-09-10
摘要:
构建细胞农业经济是解决资源短缺、降低温室气体排放、延缓全球变暖和实现可持续性经济发展的重要手段之一。微生物细胞工厂具有易于遗传改造、便于工程放大和高效利用可再生资源的优势,成为了现代生物制造的重要组成部分。王义翘教授是现代生化工程技术的开创者和奠基人,本人及同事(乔康健博士、胡鹏博士、周康博士等人)在Stephanopoulos实验室所从事的构建油脂细胞工厂的工作,受益于王先生在MIT所创建的生物工程技术中心。植物油脂具有2000多亿美元的年均市场需求,本文从植物油脂需求激增所造成的负面环境效应出发,分析了目前植物油脂的市场供应现状。产油酵母细胞工厂具有替代植物源油脂的巨大潜能。本文围绕产油酵母的代谢工程遗传改造策略,总结了如何提高碳源转化率、油脂产量、油脂生产速率和菌体生长适用性;进一步归纳了构建高效解脂耶氏酵母细胞工厂的主要技术瓶颈,其中包括产油菌株的高通量筛选和表型鉴定技术、产油酵母的代谢调控机制和发酵动力学模型等。作者进一步探讨了以蔗糖作为原材料,生产植物源功能性油脂的经济可行性和技术可行性。作者预测解脂耶氏酵母具有极大的潜力,可以解决当前高附加值油脂(比如用于巧克力生产的可可脂,潜在市场为500亿美元)的市场需求。开发产油酵母微生物资源,提供健康的功能性油脂,将会解决一系列能源、健康食品和环境资源等问题,促进我们迈向低碳性和可持续性的经济运转模式。
中图分类号:
引用本文
徐鹏. 纪念王义翘教授:解脂耶氏酵母替代植物油脂的技术瓶颈及展望[J]. 合成生物学, 2021, 2(4): 509-527.
XU Peng. In memory of Prof. Daniel I.C. Wang: Engineering Yarrowia lipolytica for the production of plant-based lipids: technical constraints and perspectives for a sustainable cellular agriculture economy[J]. Synthetic Biology Journal, 2021, 2(4): 509-527.
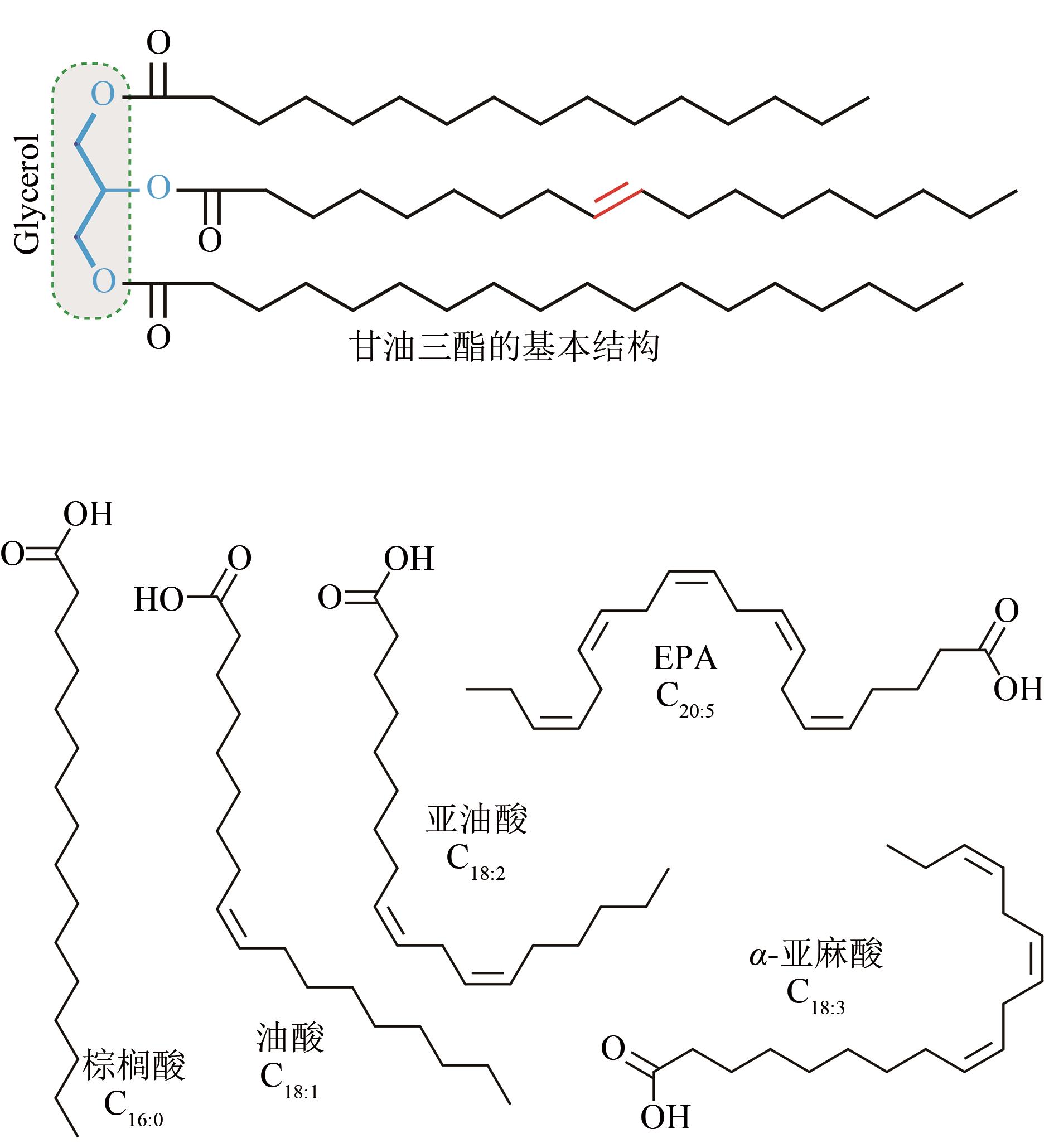
图1 油脂的分子结构及重要脂肪酸分子
Fig. 1 The basic structure of triglyceride and the four important fatty acids: palmitic acid (C16∶0), oleic acid (C18∶1), linoleic acid (C18∶2), eicosapentaenoic acid (EPA, C20∶5) and α-linolenic acid (ALA, C18∶3).
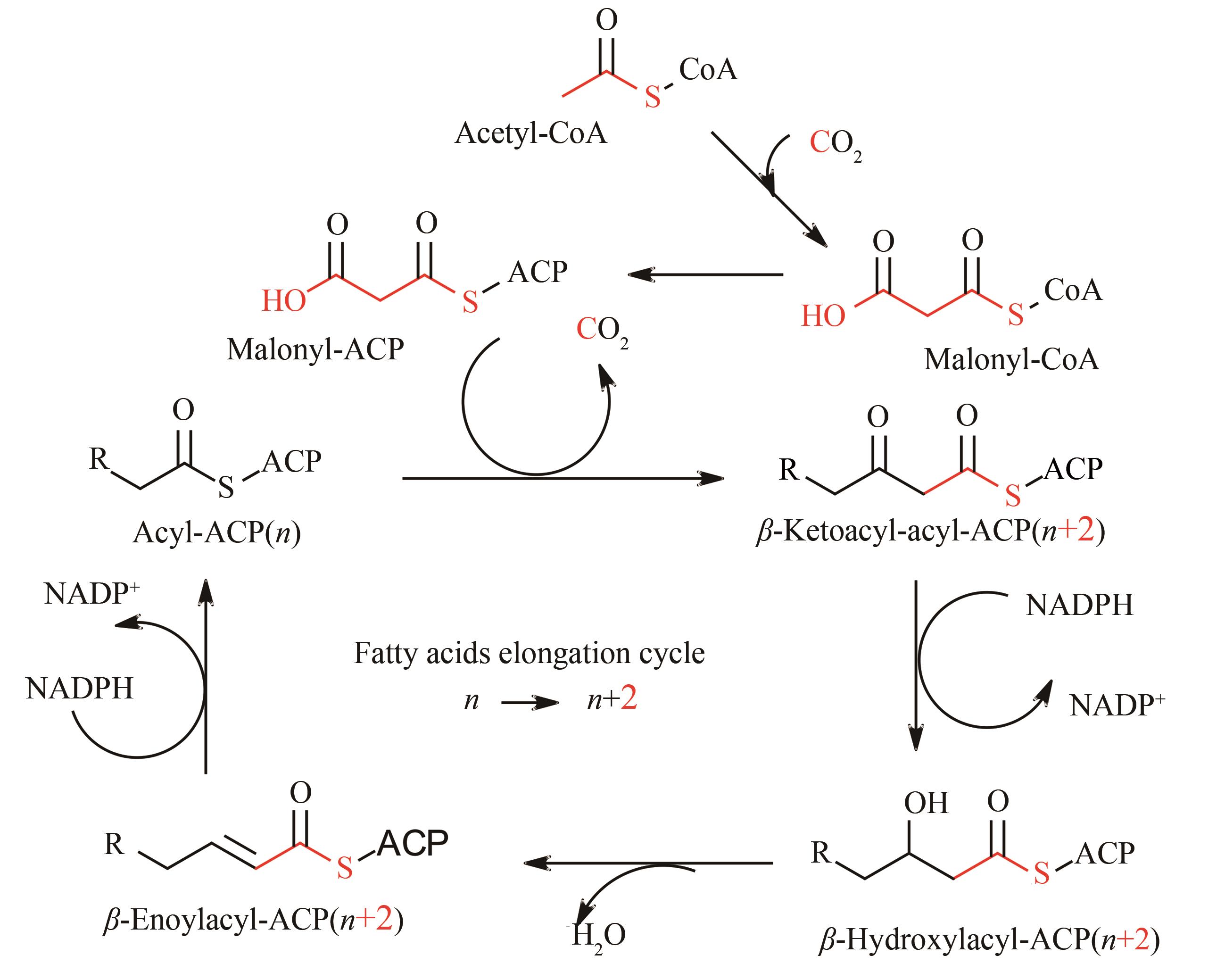
图3 脂肪酸碳链延伸过程需要还原性辅因子NADPH(n表示碳链长度)
Fig. 3 The enzymatic elongation cycle of fatty acids needs reducing equivalents NADPH(n is the chain-length of carbon backbones)
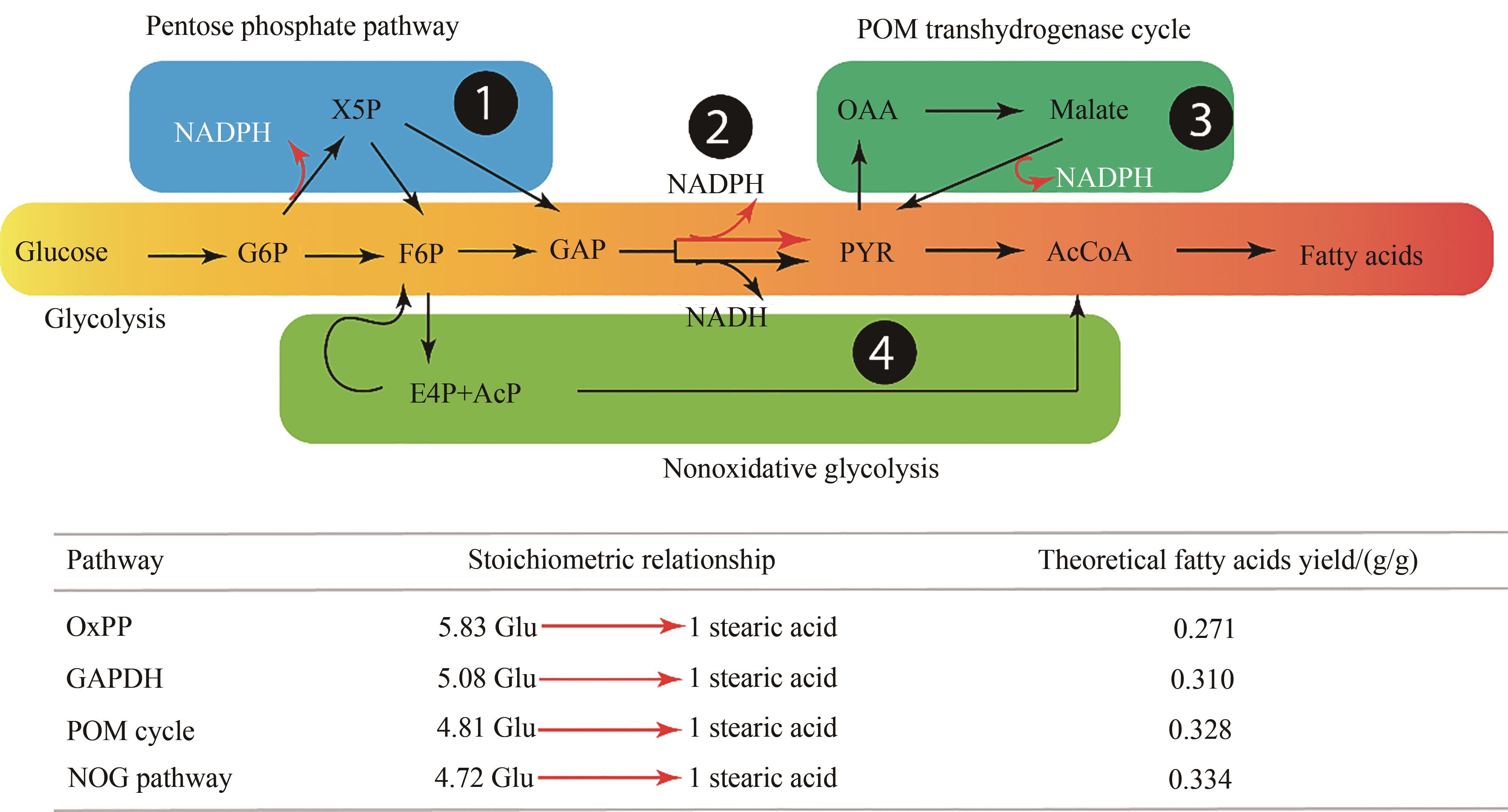
图4 不同还原性辅因子替代途径对脂肪酸合成效率的影响[56](图示中包含了以下四种途径:①磷酸戊糖途径;②NADPH-特异性的3-磷酸甘油醛脱氢酶;③丙酮酸-草酰乙酸-苹果酸转氢反应;④非氧化性糖酵解途径)
Fig. 4 Carbon conversion efficiency from NADPH for fatty acids synthesis pathways [56](OxPP—oxidative pentose phosphate pathway; GAPHD—NADPH-specific glyceraldehyde-3-phosphate dehydrogenase; POM cycle—pyruvate-oxaloacetate-malate transhydrogenase cycle; NOG—non-oxidative glycolytic pathway)
| 方程编号 Equation No. | 方程 Equations | 描述 Description |
|---|---|---|
| (1) | 细胞比生长速率 Specific growth rate | |
| (2) | 非油脂生物量积累 Oil-free cell growth | |
| (3) | 油脂积累 Lipid accumulation | |
| (4) | 底物消耗 Substrate consumption | |
| (5) | 总生物量 Total biomass | |
| (6) | 动态含油量 Oil content | |
| (7) | 动态过程得率 Process yield | |
| (8) | 总的生产强度 Overall productivity |
表1 产油酵母胞内积累油脂的分批发酵动力学描述
Tab. 1 Fermentation kinetics of the oil accumulation process in oleaginous yeast in batch culture
| 方程编号 Equation No. | 方程 Equations | 描述 Description |
|---|---|---|
| (1) | 细胞比生长速率 Specific growth rate | |
| (2) | 非油脂生物量积累 Oil-free cell growth | |
| (3) | 油脂积累 Lipid accumulation | |
| (4) | 底物消耗 Substrate consumption | |
| (5) | 总生物量 Total biomass | |
| (6) | 动态含油量 Oil content | |
| (7) | 动态过程得率 Process yield | |
| (8) | 总的生产强度 Overall productivity |
| 油料作物 Oil crop | 产量/[t/(hm2·a)] Production | 含油量/% Oil content | 最大产油得率/[t/(hm2·a)] Maximal oil yield |
|---|---|---|---|
| 黄豆 Soybean | 3.1 | 20 | 0.62 |
| 葵花籽 Sunflower seed | 1.7 | 50 | 0.85 |
| 油菜籽 Canola | 2.0 | 45 | 0.90 |
| 可可果 Cocoa bean | 0.8 | 50 | 0.40 |
| 棕榈果 Oil palm | 9.1 | 41 | 3.69 |
表2 世界主要油料作物的单位面积产量及产油得率
Tab. 2 The unit area output and maximal oil yield from major oil crops.
| 油料作物 Oil crop | 产量/[t/(hm2·a)] Production | 含油量/% Oil content | 最大产油得率/[t/(hm2·a)] Maximal oil yield |
|---|---|---|---|
| 黄豆 Soybean | 3.1 | 20 | 0.62 |
| 葵花籽 Sunflower seed | 1.7 | 50 | 0.85 |
| 油菜籽 Canola | 2.0 | 45 | 0.90 |
| 可可果 Cocoa bean | 0.8 | 50 | 0.40 |
| 棕榈果 Oil palm | 9.1 | 41 | 3.69 |
| 糖料作物 Sugar crop | 产量/[t/(hm2·a)] Production | 含糖量/% Sugar content | 最大产油得率①/[t/(hm2·a)] Maximal oil yield |
|---|---|---|---|
| 玉米 Corn | 10.7 (dry, 干重) | 74 (dry, 干重) | 2.15 |
| 红薯 Sweet potato | 25 (wet, 湿重) | 20 (wet, 湿重) | 1.36 |
| 甘蔗 Sugarcane | 80.9 (wet, 湿重) | 14 (wet, 湿重) | 3.07 |
表3 以糖料作物为原料经解脂耶氏酵母转化后的产油得率
Tab. 3 The maximal oil yield from Y. lipolytica converting sugar feedstock from corn, sweet potato or sugarcane.
| 糖料作物 Sugar crop | 产量/[t/(hm2·a)] Production | 含糖量/% Sugar content | 最大产油得率①/[t/(hm2·a)] Maximal oil yield |
|---|---|---|---|
| 玉米 Corn | 10.7 (dry, 干重) | 74 (dry, 干重) | 2.15 |
| 红薯 Sweet potato | 25 (wet, 湿重) | 20 (wet, 湿重) | 1.36 |
| 甘蔗 Sugarcane | 80.9 (wet, 湿重) | 14 (wet, 湿重) | 3.07 |
| 1 | AFEYAN N B, COONEY C L. Professor Daniel I C. Wang. a legacy of education, innovation, publication, and leadership [J]. Biotechnology and Bioengineering, 2020, 117(12): 3615-3627. |
| 2 | XUE Z, SHARPE P L, HONG S P, et al. Production of omega-3 eicosapentaenoic acid by metabolic engineering of Yarrowia lipolytica [J]. Nature Biotechnology, 2013, 31(8): 734-740. |
| 3 | SIMOPOULOS A P. The importance of the ratio of omega-6/omega-3 essential fatty acids [J]. Biomedicine & Pharmacotherapy, 2002, 56(8): 365-379. |
| 4 | VIJAY V, PIMM S L, JENKINS C N, et al. The impacts of oil palm on recent deforestation and biodiversity loss [J]. PLoS ONE, 2016, 11(7): e0159668. |
| 5 | CARLSON K M, HEILMAYR R, GIBBS H K, et al. Effect of oil palm sustainability certification on deforestation and fire in Indonesia [J]. Proceedings of the National Academy of Sciences, of the United States of America, 2018, 115(1): 121. |
| 6 | LEDESMA-AMARO R, NICAUD J M. Yarrowia lipolytica as a biotechnological chassis to produce usual and unusual fatty acids [J]. Progress in Lipid Research, 2016, 61: 40-50. |
| 7 | XU P, GU Q, WANG W, et al. Modular optimization of multi-gene pathways for fatty acids production in E. coli [J]. Nature Communications, 2013, 4: 1409. |
| 8 | HU P, CHAKRABORTY S, KUMAR A, et al. Integrated bioprocess for conversion of gaseous substrates to liquids [J]. Proceedings of the National Academy of Sciences of the United States of America, 2016, 113(14): 3773-3778. |
| 9 | XU J, LIU N, QIAO K, et al. Application of metabolic controls for the maximization of lipid production in semicontinuous fermentation [J]. Proceedings of the National Academy of Sciences of the United States of America, 2017, 114(27): E5308-E5316. |
| 10 | MA J, GU Y, MARSAFARI M, et al. Synthetic biology, systems biology, and metabolic engineering of Yarrowia lipolytica toward a sustainable biorefinery platform [J]. Journal of Industrial Microbiology and Biotechnology, 2020, 47(9/10): 845-862. |
| 11 | SCHWARTZ C M, HUSSAIN M S, BLENNER M, et al. Synthetic RNA polymerase III promoters facilitate high-efficiency CRISPR-Cas9-mediated genome editing in Yarrowia lipolytica [J]. ACS Synthetic Biology, 2016, 5(4): 356-359. |
| 12 | EDWARDS H, YANG Z, XU P. Characterization of Met25 as a color associated genetic marker in Yarrowia lipolytica [J]. Metabolic Engineering Communications, 2020, 11: e00147. |
| 13 | YANG Z, EDWARDS H, XU P. CRISPR-Cas12a/Cpf1-assisted precise, efficient and multiplexed genome-editing in Yarrowia lipolytica [J]. Metabolic Engineering Communications, 2020, 10: e00112. |
| 14 | YANG Z, BLENNER M. Genome editing systems across yeast species [J]. Current Opinion in Biotechnology, 2020, 66: 255-266. |
| 15 | YANG Z, XU P. Implementing CRISPR-Cas12a for efficient genome editing in Yarrowia lipolytica [M]// WHEELDON I, BLENNER M. Yarrowia lipolytica: methods and protocols. New York: Springer, 2021: 111-121. |
| 16 | CELIŃSKA E, LEDESMA-AMARO R, LARROUDE M, et al. Golden Gate Assembly system dedicated to complex pathway manipulation in Yarrowia lipolytica [J]. Microbial Biotechnology, 2017, 10(2): 450-455. |
| 17 | EGERMEIER M, SAUER M, MARX H. Golden Gate-based metabolic engineering strategy for wild-type strains of Yarrowia lipolytica [J]. FEMS Microbiology Letters, 2019, 366(4): fnz022. |
| 18 | LARROUDE M, PARK Y K, SOUDIER P, et al. A modular Golden Gate toolkit for Yarrowia lipolytica synthetic biology [J]. Microbial Biotechnology, 2019, 12(6): 1249-1259. |
| 19 | TONG Y, ZHOU J, ZHANG L, et al. A golden-gate based cloning toolkit to build violacein pathway libraries in Yarrowia lipolytica [J]. ACS Synthetic Biology, 2021, 10(1): 115-124. |
| 20 | WONG L, HOLDRIDGE B, ENGEL J, et al. Genetic tools for streamlined and accelerated pathway engineering in Yarrowia Lipolytica [M]// SANTOS C N S and AJIKUMAR P K. Microbial metabolic engineering: methods and protocols. New York: Springer, 2019, 155-177. |
| 21 | WONG L, ENGEL J, JIN E, et al. YaliBricks, a versatile genetic toolkit for streamlined and rapid pathway engineering in Yarrowia lipolytica [J]. Metabolic Engineering Communications, 2017, 5(): 68-77. |
| 22 | LIU H, MARSAFARI M, DENG L, et al. Understanding lipogenesis by dynamically profiling transcriptional activity of lipogenic promoters in Yarrowia lipolytica [J]. Applied Microbiology and Biotechnology, 2019, 103(7): 3167-3179. |
| 23 | GUO Z-P, BORSENBERGER V, CROUX C, et al. An artificial chromosome ylAC enables efficient assembly of multiple genes in Yarrowia lipolytica for biomanufacturing [J]. Communications Biology, 2020, 3(1): 199. |
| 24 | LV Y, EDWARDS H, ZHOU J, et al. Combining 26S rDNA and the Cre-loxP system for iterative gene integration and efficient marker curation in Yarrowia lipolytica [J]. ACS Synthetic Biology, 2019, 8(3): 568-576. |
| 25 | LIU J, WU X, YAO M, et al. Chassis engineering for microbial production of chemicals: from natural microbes to synthetic organisms [J]. Current Opinion in Biotechnology, 2020, 66: 105-112. |
| 26 | LIU Y, SU A, LI J, et al. Towards next-generation model microorganism chassis for biomanufacturing [J]. Applied Microbiology and Biotechnology, 2020, 104(21): 9095-9108. |
| 27 | PENDRILL F, PERSSON U M, GODAR J, et al. Agricultural and forestry trade drives large share of tropical deforestation emissions [J]. Global Environmental Change, 2019, 56: 1-10. |
| 28 | SCHROTH G, LÄDERACH P, MARTINEZ-VALLE A I, et al. Vulnerability to climate change of cocoa in West Africa: patterns, opportunities and limits to adaptation [J]. Science of the Total Environment, 2016, 556: 231-241. |
| 29 | MARKHAM K A, PALMER C M, CHWATKO M, et al. Rewiring Yarrowia lipolytica toward triacetic acid lactone for materials generation [J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(9): 2096. |
| 30 | LIU H, MARSAFARI M, WANG F, et al. Engineering acetyl-CoA metabolic shortcut for eco-friendly production of polyketides triacetic acid lactone in Yarrowia lipolytica [J]. Metabolic Engineering, 2019, 56: 60-68. |
| 31 | QIU X, XU P, ZHAO X, et al. Combining genetically-encoded biosensors with high throughput strain screening to maximize erythritol production in Yarrowia lipolytica [J]. Metabolic Engineering, 2020, 60: 66-76. |
| 32 | XU P, QIAO K, AHN W S, et al. Engineering Yarrowia lipolytica as a platform for synthesis of drop-in transportation fuels and oleochemicals [J]. Proceedings of the National Academy of Sciences of the United States of America, 2016, 113(39): 10848-10853. |
| 33 | SUN W, YANG Z, XU P. Engineering Yarrowia lipolytica for production of fatty alcohols with YaliBrick vectors[M]// WHEELDON I, BLENNER M.Yarrowia lipolytica: methods and protocols. New York: Springer, 2021: 159-173. |
| 34 | LI J, MA Y, LIU N,et al. Synthesis of high-titer alka(e)nes in Yarrowia lipolytica is enabled by a discovered mechanism [J]. Nature Communications, 2020, 11(1): 6198. |
| 35 | XUE Z, SHARPE P L, HONG S P, et al. Production of omega-3 eicosapentaenoic acid by metabolic engineering of Yarrowia lipolytica [J]. Nature Biotechnology, 2013, 31: 734-740. |
| 36 | LV Y, QIAN S, DU G, et al. Coupling feedback genetic circuits with growth phenotype for dynamic population control and intelligent bioproduction [J]. Metabolic Engineering, 2019, 54: 109-116. |
| 37 | LV Y, MARSAFARI M, KOFFAS M, et al. Optimizing oleaginous yeast cell factories for flavonoids and hydroxylated flavonoids biosynthesis[J]. ACS Synthetic Biology, 2019, 8(11): 2514-2523. |
| 38 | PALMER C M, MILLER K K, NGUYEN A, et al. Engineering 4-coumaroyl-CoA derived polyketide production in Yarrowia lipolytica through a β-oxidation mediated strategy [J]. Metabolic Engineering, 2020, 57: 174-181. |
| 39 | LIU H, WANG F, DENG L, et al. Genetic and bioprocess engineering to improve squalene production in Yarrowia lipolytica [J]. Bioresource Technology, 2020, 317: 123991. |
| 40 | HUANG Y Y, JIAN X X, LV Y B, et al. Enhanced squalene biosynthesis in Yarrowia lipolytica based on metabolically engineered acetyl-CoA metabolism [J]. Journal of Biotechnology, 2018, 281: 106-114. |
| 41 | TANG W-Y, WANG D-P, TIAN Y, et al. Metabolic engineering of Yarrowia lipolytica for improving squalene production [J]. Bioresource Technology, 2021, 323: 124652. |
| 42 | MARSAFARI M, XU P. Debottlenecking mevalonate pathway for antimalarial drug precursor amorphadiene biosynthesis in Yarrowia lipolytica [J]. Metabolic Engineering Communications, 2020, 10: e00121. |
| 43 | SÁEZ-SÁEZ J, WANG G, MARELLA E R, et al. Engineering the oleaginous yeast Yarrowia lipolytica for high-level resveratrol production [J]. Metabolic Engineering, 2020, 62: 51-61. |
| 44 | GU Y, MA J, ZHU Y, et al. Engineering Yarrowia lipolytica as a chassis for de novo synthesis of five aromatic-derived natural products and chemicals[J]. ACS Synthetic Biology, 2020, 9(8): 2096-2106. |
| 45 | GU Y, MA J, ZHU Y, et al. Refactoring ehrlich pathway for high-yield 2-phenylethanol production in Yarrowia lipolytica [J]. ACS Synthetic Biology, 2020, 9(3): 623-633. |
| 46 | SOARES G P A, SOUZA K S T, VILELA L F, et al. γ-Decalactone production by Yarrowia lipolytica and Lindnera saturnus in crude glycerol [J]. Preparative Biochemistry & Biotechnology, 2017, 47(6): 633-637. |
| 47 | BRAGA A, BELO I. Biotechnological production of γ-decalactone, a peach like aroma, by Yarrowia lipolytica [J]. World Journal of Microbiology and Biotechnology, 2016, 32(10): 169. |
| 48 | MARELLA E R, DAHLIN J, DAM M I, et al. A single-host fermentation process for the production of flavor lactones from non-hydroxylated fatty acids [J]. Metabolic Engineering, 2020, 61: 427-436. |
| 49 | MA J, GU Y, XU P. A roadmap to engineering antiviral natural products synthesis in microbes [J]. Current Opinion in Biotechnology, 2020, 66: 140-149. |
| 50 | MARSAFARI M, SAMIZADEH H, RABIEI B, et al. Biotechnological production of flavonoids: an update on plant metabolic engineering, microbial host selection, and genetically encoded biosensors [J]. Biotechnology Journal, 2020, 15(8): 1900432. |
| 51 | 汪庆卓, 黄和. 合成生物技术驱动天然的真核油脂细胞工厂开发[J]. 合成生物学, 2021, doi:10.12211/2096-8280.2020-090 . |
| WANG Q Z, HUANG H. Synthetic biotechnology drives the development of natural eukaryotic lipid cell factories [J]. Synthetic Biology Journal, 2021, doi: 10.12211/2096-8280.2020-090 . | |
| 52 | MUHAMMAD A, FENG X D, RASOOL A, et al. Production of plant natural products through engineered Yarrowia lipolytica [J]. Biotechnology Advances, 2020, 43: 107555. |
| 53 | WASYLENKO T M, AHN W S, STEPHANOPOULOS G. The oxidative pentose phosphate pathway is the primary source of NADPH for lipid overproduction from glucose in Yarrowia lipolytica [J]. Metabolic Engineering, 2015, 30: 27-39. |
| 54 | RATLEDGE C. The role of malic enzyme as the provider of NADPH in oleaginous microorganisms: a reappraisal and unsolved problems [J]. Biotechnology Letters, 2014, 36(8): 1557-1568. |
| 55 | WYNN J P, HAMID A B A, RATLEDGE C. The role of malic enzyme in the regulation of lipid accumulation in filamentous fungi [J]. Microbiology, 1999, 145(8): 1911-1917. |
| 56 | QIAO K, WASYLENKO T M, ZHOU K, et al. Lipid production in Yarrowia lipolytica is maximized by engineering cytosolic redox metabolism [J]. Nature Biotechnology, 2017, 35(2): 173-177. |
| 57 | JAYAKODY L N, JIN Y S. In-depth understanding of molecular mechanisms of aldehyde toxicity to engineer robust Saccharomyces cerevisiae [J]. Applied Microbiology and Biotechnology, 2021, 105(7): 2675-2692. |
| 58 | XU P, QIAO K, STEPHANOPOULOS G. Engineering oxidative stress defense pathways to build a robust lipid production platform in Yarrowia lipolytica [J]. Biotechnology and Bioengineering, 2017, 114(7): 1521-1530. |
| 59 | GU Y, XU P. Synthetic yeast brews neuroactive compounds [J]. Nature Chemical Biology, 2021, 17(1): 8-9. |
| 60 | SHUIB S, IBRAHIM I, MACKEEN M M, et al. First evidence for a multienzyme complex of lipid biosynthesis pathway enzymes in Cunninghamella bainieri [J]. Scientific Reports, 2018, 8(1): 3077. |
| 61 | RATLEDGE C. Regulation of lipid accumulation in oleaginous micro-organisms [J]. Biochemical Society Transactions, 2002, 30(6): 1047. |
| 62 | ZHOU Y J, BUIJS N A, ZHU Z, et al. Production of fatty acid-derived oleochemicals and biofuels by synthetic yeast cell factories [J]. Nature Communications, 2016, 7(1): 11709. |
| 63 | YU T, ZHOU Y J, WENNING L, et al. Metabolic engineering of Saccharomyces cerevisiae for production of very long chain fatty acid-derived chemicals [J]. Nature Communications, 2017, 8(1): 15587. |
| 64 | BELD J, LEE D J, BURKART M D. Fatty acid biosynthesis revisited: structure elucidation and metabolic engineering [J]. Molecular BioSystems, 2015, 11(1): 38-59. |
| 65 | ZHU Z, HU Y, TEIXEIRA P G, et al. Multidimensional engineering of Saccharomyces cerevisiae for efficient synthesis of medium-chain fatty acids [J]. Nature Catalysis, 2020, 3(1): 64-74. |
| 66 | GAJEWSKI J, PAVLOVIC R, FISCHER M, et al. Engineering fungal de novo fatty acid synthesis for short chain fatty acid production [J]. Nature Communications, 2017, 8(1): 14650. |
| 67 | SINGH K, GRAF B, LINDEN A, et al. Discovery of a regulatory subunit of the yeast fatty acid synthase [J]. Cell, 2020, 180(6): 1130-1143. |
| 68 | BLAZECK J, HILL A, LIU L, et al. Harnessing Yarrowia lipolytica lipogenesis to create a platform for lipid and biofuel production [J]. Nature Communications, 2014, 5: 3131. |
| 69 | ZHANG X K, NIE M Y, CHEN J, et al. Multicopy integrants of crt genes and co-expression of AMP deaminase improve lycopene production in Yarrowia lipolytica [J]. Journal of Biotechnology, 2019, 289: 46-54. |
| 70 | SILVERMAN A M, QIAO K, XU P, et al. Functional overexpression and characterization of lipogenesis-related genes in the oleaginous yeast Yarrowia lipolytica [J]. Applied Microbiology and Biotechnology, 2016, 100(8): 3781-3798. |
| 71 | CHANG L, TANG X, LU H, et al. Role of adenosine monophosphate deaminase during fatty acid accumulation in oleaginous fungus Mortierella alpina [J]. Journal of Agricultural and Food Chemistry, 2019, 67(34): 9551-9559. |
| 72 | SEIP J, JACKSON R, HE H, et al. Snf1 is a regulator of lipid accumulation in Yarrowia lipolytica [J]. Applied and Environmental Microbiology, 2013, 79(23): 7360-7370. |
| 73 | POMRANING K R, KIM Y M, NICORA C D, et al. Multi-omics analysis reveals regulators of the response to nitrogen limitation in Yarrowia lipolytica [J]. BMC Genomics, 2016, 17(1): 138. |
| 74 | KERKHOVEN E J, POMRANING K R, BAKER S E, et al. Regulation of amino-acid metabolism controls flux to lipid accumulation in Yarrowia lipolytica [J]. npj Systems Biology and Applications, 2016, 2: 16005. |
| 75 | BLAZECK J, HILL A, LIU L, et al. Harnessing Yarrowia lipolytica lipogenesis to create a platform for lipid and biofuel production [J]. Nature Communications, 2014, 5: 3131. |
| 76 | POMRANING K R, BREDEWEG E L, BAKER S E. Regulation of nitrogen metabolism by GATA zinc finger transcription factors in Yarrowia lipolytica [J]. mSphere, 2017, 2(1): e00038-17. |
| 77 | XU P. Analytical solution for a hybrid Logistic-Monod cell growth model in batch and continuous stirred tank reactor culture [J]. Biotechnology and Bioengineering, 2020, 117(3): 873-878. |
| 78 | ROEHR M, ZEHENTGRUBER O, KUBICEK C P. Kinetics of biomass formation and citric acid production by Aspergillus niger on pilot plant scale [J]. Biotechnology and Bioengineering, 1981, 23(11): 2433-2445. |
| 79 | CAI D, ZHU C, CHEN S. Microbial production of nattokinase: current progress, challenge and prospect [J]. World Journal of Microbiology and Biotechnology, 2017, 33(5): p. 84. |
| 80 | XU P, DING Z, QIAN Z, et al. Improved production of mycelial biomass and ganoderic acid by submerged culture of Ganoderma lucidum SB97 using complex media [J]. Enzyme and Microbial Technology, 2008, 42(4): 325-331. |
| 81 | SURENDHIRAN D, VIJAY M, SIVAPRAKASH B, et al. Kinetic modeling of microalgal growth and lipid synthesis for biodiesel production [J]. 3 Biotech, 2015, 5(5): 663-669. |
| 82 | PAPANIKOLAOU S, AGGELIS G. Modeling lipid accumulation and degradation in Yarrowia lipolytica cultivated on industrial fats [J]. Current Microbiology, 2003, 46(6): 0398-0402. |
| 83 | DULERMO R, GAMBOA-MELÉNDEZ H, DULERMO T, et al. The fatty acid transport protein Fat1p is involved in the export of fatty acids from lipid bodies in Yarrowia lipolytica [J]. FEMS Yeast Research, 2014, 14(6): 883-896. |
| 84 | RUMIN J, BONNEFOND H, SAINT-JEAN B, et al. The use of fluorescent Nile red and BODIPY for lipid measurement in microalgae [J]. Biotechnology for Biofuels, 2015, 8(1): 42. |
| 85 | FRIEDLANDER J, TSAKRAKLIDES V, KAMINENI A, et al. Engineering of a high lipid producing Yarrowia lipolytica strain [J]. Biotechnology for Biofuels, 2016, 9(1): 77. |
| 86 | MENG Y, YAO C, XUE S, et al. Application of Fourier transform infrared (FT-IR) spectroscopy in determination of microalgal compositions [J]. Bioresource Technology, 2014, 151: 347-354. |
| 87 | CHMIELARZ M, SAMPELS S, BLOMQVIST J, et al. FT-NIR: a tool for rapid intracellular lipid quantification in oleaginous yeasts [J]. Biotechnology for Biofuels, 2019, 12(1): 169. |
| 88 | NĚMCOVÁ A, GONOVÁ D, SAMEK O, et al. The use of raman spectroscopy to monitor metabolic changes in stressed Metschnikowia sp. yeasts [J]. Microorganisms, 2021, 9(2): 277. |
| 89 | BOWMAN E K, ALPER H S. Microdroplet-assisted screening of biomolecule production for metabolic engineering applications [J]. Trends in Biotechnology, 2020, 38(7): 701-714. |
| 90 | CAO W, CHENG S, YANG J, et al. Large-scale lipid analysis with C ̿ C location and sn-position isomer resolving power [J]. Nature Communications, 2020. 11(1): 375. |
| 91 | WANG M, WEI Y, JI B, et al. Advances in metabolic engineering of saccharomyces cerevisiae for cocoa butter equivalent production [J]. Frontiers in Bioengineering and Biotechnology, 2020, 8: 1194. |
| 92 | WEI Y, SIEWERS V, NIELSEN J. Cocoa butter-like lipid production ability of non-oleaginous and oleaginous yeasts under nitrogen-limited culture conditions [J]. Applied Microbiology and Biotechnology, 2017, 101(9): 3577-3585. |
| 93 | WEI Y, BERGENHOLM D, GOSSING M, et al. Expression of cocoa genes in Saccharomyces cerevisiae improves cocoa butter production [J]. Microbial Cell Factories, 2018, 17(1): 11. |
| 94 | XIONG D, ZHANG H Y, XIE Y F, et al. Conversion of mutton fat to cocoa butter equivalent by increasing the unsaturated fatty acids at the sn-2 position of triacylglycerol through fermentation by Yarrowia Lipolytica [J]. American Journal of Biochemistry and Biotechnology, 2015, 11(2): 57-65. |
| 95 | ZHAO L N, LI B, XIONG D, et al. Cocoa-butter-equivalent production from Yarrowia lipolytica by optimization of fermentation technology [J]. American Journal of Biochemistry and Biotechnology, 2016, 12(4): 196-205. |
| 96 | METZ J G, ROESSLER P, FACCIOTTI D, et al. Production of polyunsaturated fatty acids by polyketide synthases in both prokaryotes and eukaryotes [J]. Science, 2001, 293(5528): 290. |
| [1] | 郭姝媛, 张倩楠, 姑丽克孜·买买提热夏提, 杨一群, 于涛. 液体生物燃料合成与炼制的研究进展[J]. 合成生物学, 2025, 6(1): 18-44. |
| [2] | 应汉杰, 柳东, 王振宇, 沈涛, 庄伟, 朱晨杰. 工业生物制造与“碳中和”目标探讨[J]. 合成生物学, 2025, 6(1): 1-7. |
| [3] | 邵明威, 孙思勉, 杨时茂, 陈国强. 基于极端微生物的生物制造[J]. 合成生物学, 2024, 5(6): 1419-1436. |
| [4] | 赵亮, 李振帅, 付丽平, 吕明, 王士安, 张全, 刘立成, 李福利, 刘自勇. 生物转化一碳化合物原料产油脂与单细胞蛋白研究进展[J]. 合成生物学, 2024, 5(6): 1300-1318. |
| [5] | 竺方欢, 岑雪聪, 陈振. 微生物合成二元醇研究进展[J]. 合成生物学, 2024, 5(6): 1367-1385. |
| [6] | 禹伟, 高教琪, 周雍进. 一碳生物转化合成有机酸的研究进展[J]. 合成生物学, 2024, 5(5): 1169-1188. |
| [7] | 陈锡玮, 张华然, 邹懿. 真菌源非核糖体肽类药物生物合成及代谢工程[J]. 合成生物学, 2024, 5(3): 571-592. |
| [8] | 谢皇, 郑义蕾, 苏依婷, 阮静怡, 李永泉. 放线菌聚酮类化合物生物合成体系重构研究进展[J]. 合成生物学, 2024, 5(3): 612-630. |
| [9] | 惠真, 唐啸宇. CRISPR/Cas9编辑系统在微生物天然产物研究中的应用[J]. 合成生物学, 2024, 5(3): 658-671. |
| [10] | 赵静宇, 张健, 祁庆生, 王倩. 基于细菌双组分系统的生物传感器的研究进展[J]. 合成生物学, 2024, 5(1): 38-52. |
| [11] | 孙绘梨, 崔金玉, 栾国栋, 吕雪峰. 面向高效光驱固碳产醇的蓝细菌合成生物技术研究进展[J]. 合成生物学, 2023, 4(6): 1161-1177. |
| [12] | 晏雄鹰, 王振, 娄吉芸, 张皓瑜, 黄星宇, 王霞, 杨世辉. 生物燃料高效生产微生物细胞工厂构建研究进展[J]. 合成生物学, 2023, 4(6): 1082-1121. |
| [13] | 孙梦楚, 陆亮宇, 申晓林, 孙新晓, 王佳, 袁其朋. 基于荧光检测的高通量筛选技术和装备助力细胞工厂构建[J]. 合成生物学, 2023, 4(5): 947-965. |
| [14] | 刁志钿, 王喜先, 孙晴, 徐健, 马波. 单细胞拉曼光谱测试分选装备研制及应用进展[J]. 合成生物学, 2023, 4(5): 1020-1035. |
| [15] | 吴玉洁, 刘欣欣, 刘健慧, 杨开广, 随志刚, 张丽华, 张玉奎. 基于高通量液相色谱质谱技术的菌株筛选与关键分子定量分析研究进展[J]. 合成生物学, 2023, 4(5): 1000-1019. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||