合成生物学 ›› 2021, Vol. 2 ›› Issue (6): 1046-1060.DOI: 10.12211/2096-8280.2021-098
• 研究论文 • 上一篇
在大肠杆菌中从头生物合成3-苯丙醇
高虎涛, 王佳, 孙新晓, 申晓林, 袁其朋
- 北京化工大学,化工资源有效利用国家重点实验室,北京 100029
-
收稿日期:2021-10-21修回日期:2021-11-29出版日期:2021-12-31发布日期:2022-01-21 -
通讯作者:申晓林,袁其朋 -
作者简介:高虎涛 (1997—),男,硕士研究生。研究方向为芳香氨基酸衍生物的合成生物学和代谢工程。E-mail:2019201106@buct.edu.cn申晓林 (1984—),女,博士,副教授。研究方向为代谢工程及微生物合成生物学。E-mail:shenxl@mail.buct.edu.cn袁其朋 (1969—),男,博士,教授。研究方向为代谢工程及微生物合成生物学。E-mail:yuanqp@mail.buct.edu.cn -
基金资助:国家重点研发计划(2018YFA0901800)
De novo biosynthesis of 3-phenylpropanol in E. coli
GAO Hutao, WANG Jia, SUN Xinxiao, SHEN Xiaolin, YUAN Qipeng
- State Key Laboratory of Effective Utilization of Chemical Resources,Beijing University of Chemical Technology,Beijing 100029,China
-
Received:2021-10-21Revised:2021-11-29Online:2021-12-31Published:2022-01-21 -
Contact:SHEN Xiaolin, YUAN Qipeng
摘要:
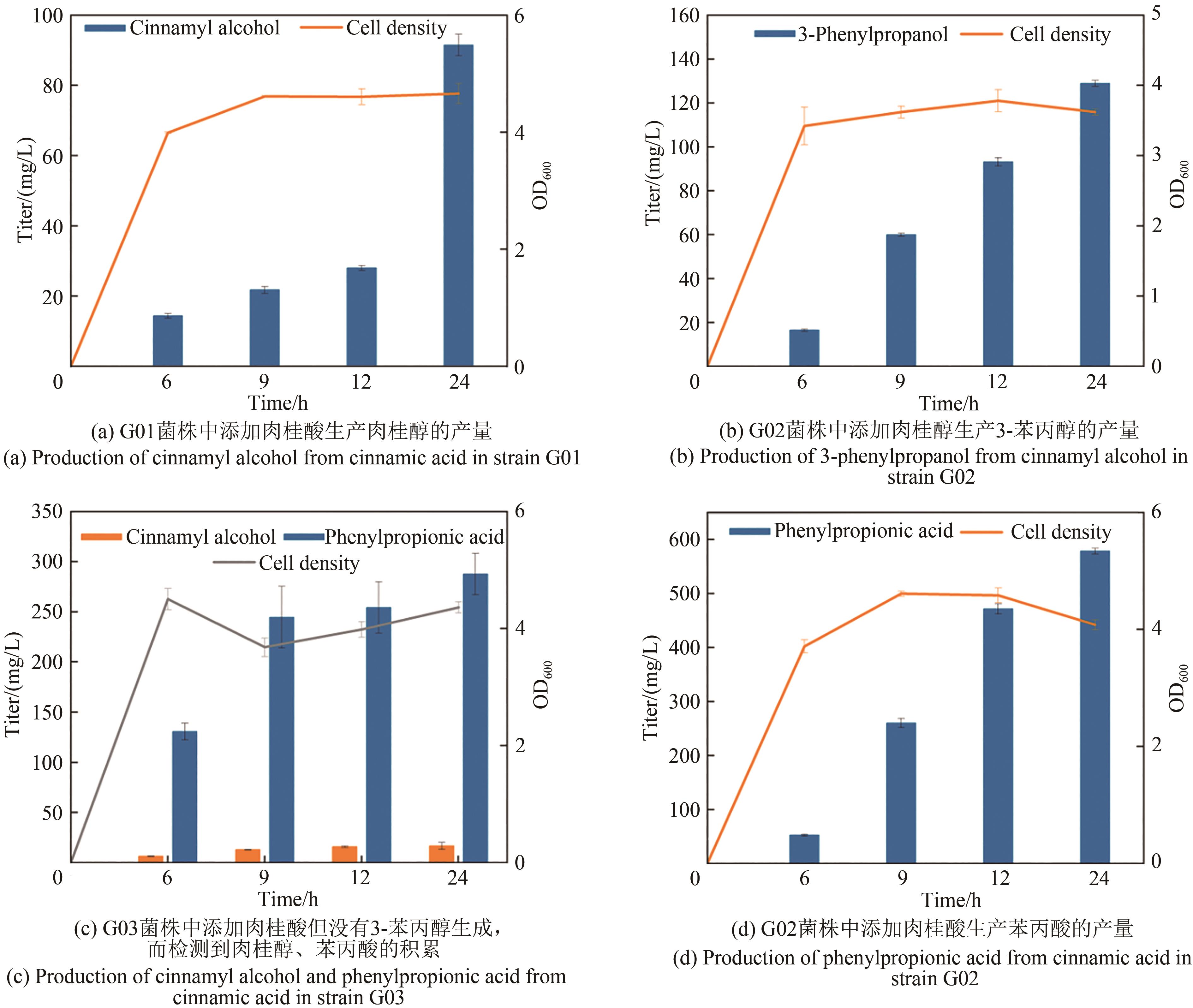
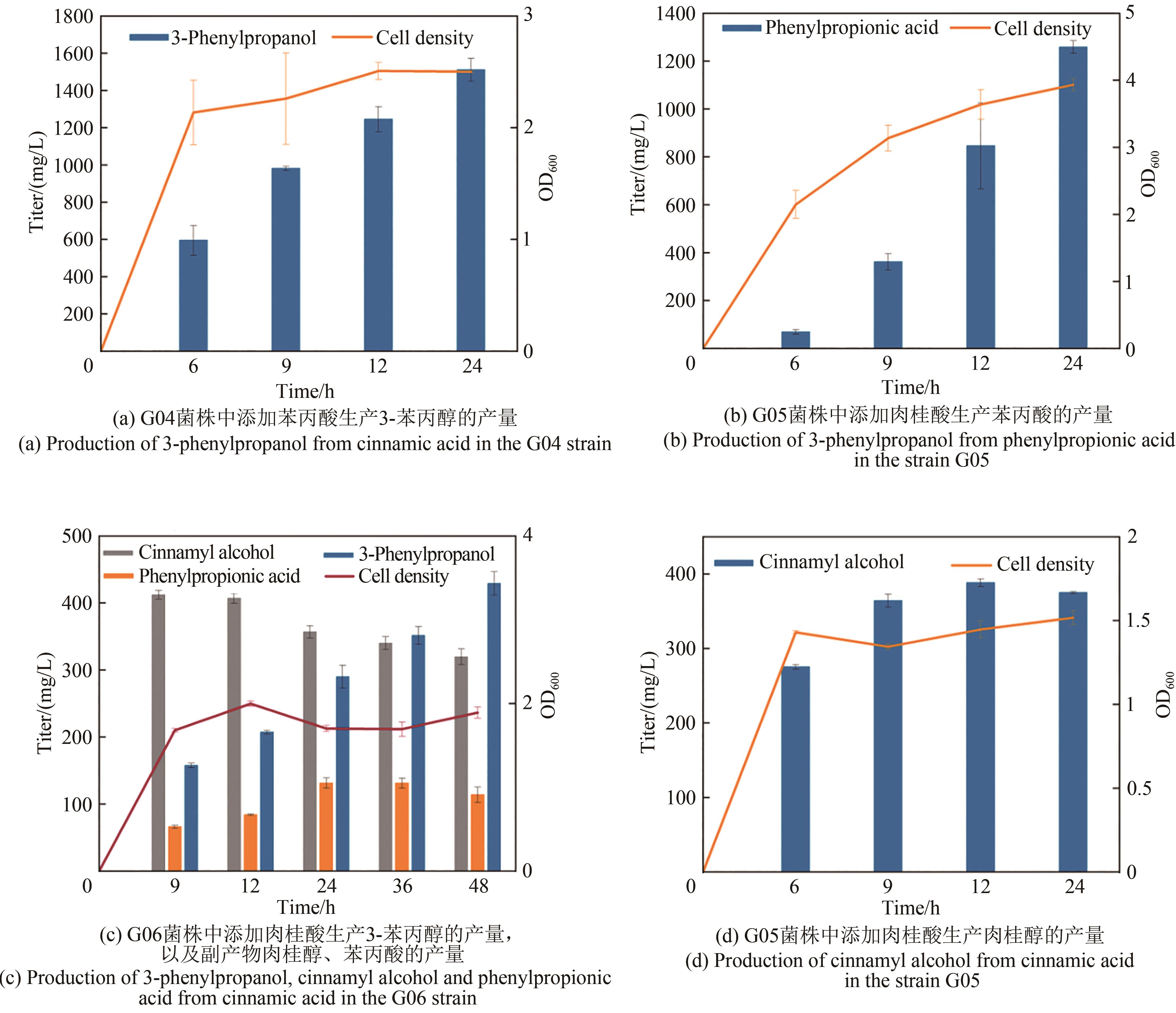
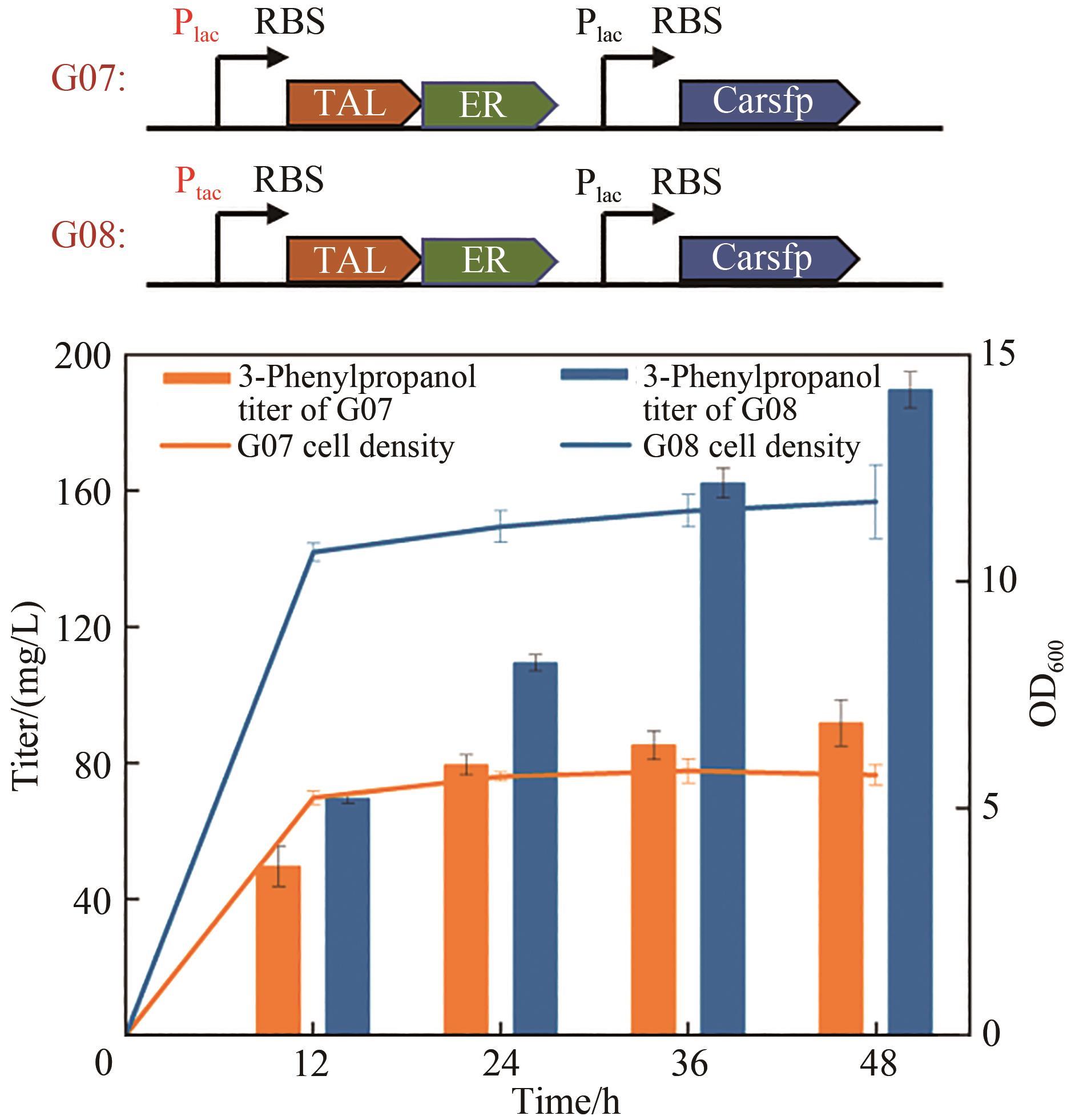
3-苯丙醇是一种具有芳香味的高价值香料,在医药、化妆品、食品等领域有广泛用途,是生产多种药品和化学品的重要前体。其目前的生产方法主要依赖于植物提取和化学合成,存在产物得率低、生产周期长和环境不友好等缺点。为解决这些问题,构建微生物细胞工厂利用可再生资源合成3-苯丙醇具有重要的意义。本研究通过将目标化合物与微生物自身代谢网络建立联系,基于底物或中间体与产物的结构类似性以及化合物间的基团转移关系,设计并构建了两条不同的3-苯丙醇的人工生物合成途径。其中,依赖羧酸还原酶的苯丙醇生物合成途径具有较高的生产效率。在大肠杆菌中实现了以甘油为碳源,从头生物合成3-苯丙醇,产量达91 mg/L。通过消除限速步骤,增加莽草酸途径碳通量以及敲除竞争途径等代谢工程策略的实施,将苯丙醇的产量提高到了841 mg/L,较初始菌株产量提高了9.2倍,为苯丙醇的绿色、可持续、大规模生产提供了基础。
中图分类号:
引用本文
高虎涛, 王佳, 孙新晓, 申晓林, 袁其朋. 在大肠杆菌中从头生物合成3-苯丙醇[J]. 合成生物学, 2021, 2(6): 1046-1060.
GAO Hutao, WANG Jia, SUN Xinxiao, SHEN Xiaolin, YUAN Qipeng. De novo biosynthesis of 3-phenylpropanol in E. coli[J]. Synthetic Biology Journal, 2021, 2(6): 1046-1060.
| Strains | Genotype | Source |
|---|---|---|
| Trans 5α | Lab Stock | |
| BW25113(F′) | Lab Stock | |
| BW25113(F′) ∆pykA ∆pykF | This study | |
| G01 | BW25113(F′) harboring pZE-CCR-4CL1 | This study |
| G02 | BW25113(F′) harboring pCS-ER | This study |
| G03 | BW25113(F′) harboring pZE-CCR-4CL1 and pCS-ER | This study |
| G04 | BW25113(F′) harboring pZE-ER | This study |
| G05 | BW25113(F′) harboring pCS-Carsfp | This study |
| G06 | BW25113(F′) harboring pZE-ER and pCS-Carsfp | This study |
| G07 | BW25113(F′) harboring pZE-RgTAL-ER and pCS-Carsfp | This study |
| G08 | BW25113(F′) harboring pZE-tac-RgTAL-ER-Carsfp | This study |
| G09 | BW25113(F′)∆pykA∆pykF harboring pZE-tac-RgTAL-ER-Carsfp | This study |
| G10 | BW25113(F′) harboring pCS-lac-APTA and pZE-tac-RgTAL-ER-Carsfp | This study |
| G11 | BW25113(F′) harboring pCS-tac-APTA and pZE-tac-RgTAL-ER-Carsfp | This study |
| G12 | BW25113(F′)∆pykA∆pykF harboring pCS-tac-APTA and pZE-tac-RgTAL-ER-Carsfp | This study |
表1 本实验所用到的菌株
Tab. 1 Strains used in this study
| Strains | Genotype | Source |
|---|---|---|
| Trans 5α | Lab Stock | |
| BW25113(F′) | Lab Stock | |
| BW25113(F′) ∆pykA ∆pykF | This study | |
| G01 | BW25113(F′) harboring pZE-CCR-4CL1 | This study |
| G02 | BW25113(F′) harboring pCS-ER | This study |
| G03 | BW25113(F′) harboring pZE-CCR-4CL1 and pCS-ER | This study |
| G04 | BW25113(F′) harboring pZE-ER | This study |
| G05 | BW25113(F′) harboring pCS-Carsfp | This study |
| G06 | BW25113(F′) harboring pZE-ER and pCS-Carsfp | This study |
| G07 | BW25113(F′) harboring pZE-RgTAL-ER and pCS-Carsfp | This study |
| G08 | BW25113(F′) harboring pZE-tac-RgTAL-ER-Carsfp | This study |
| G09 | BW25113(F′)∆pykA∆pykF harboring pZE-tac-RgTAL-ER-Carsfp | This study |
| G10 | BW25113(F′) harboring pCS-lac-APTA and pZE-tac-RgTAL-ER-Carsfp | This study |
| G11 | BW25113(F′) harboring pCS-tac-APTA and pZE-tac-RgTAL-ER-Carsfp | This study |
| G12 | BW25113(F′)∆pykA∆pykF harboring pCS-tac-APTA and pZE-tac-RgTAL-ER-Carsfp | This study |
| Plasmids | Description | Source |
|---|---|---|
| pZE12-luc | pLlacO-1; luc; ColE1 ori; Ampr | Lab Stock |
| pCS27 | pLlacO-1, P15A ori, Kanr | Lab Stock |
| pZE-CCR-4CL1 | pZE-luc carrying CCR from Leucaena leucocephala, and 4CL1 from Arabidopsis thaliana | This study |
| pZE-ER | pZE-luc carrying ER from Clostridium acetobutylicum | This study |
| pCS-ER | pCS27 carrying ER from C. acetobutylicum | This study |
| pCS-carsfp | pCS27 carrying Car from Mycobacterium marinum and Sfp from Bacillus subtilis | This study |
| pZE-RgTAL-ER | pZE-luc carrying TAL from Rhodobacter glutinis and ER from C.acetobutylicum | This study |
表2 本实验所用到的基础质粒
Tab. 2 Plasmids used in this study
| Plasmids | Description | Source |
|---|---|---|
| pZE12-luc | pLlacO-1; luc; ColE1 ori; Ampr | Lab Stock |
| pCS27 | pLlacO-1, P15A ori, Kanr | Lab Stock |
| pZE-CCR-4CL1 | pZE-luc carrying CCR from Leucaena leucocephala, and 4CL1 from Arabidopsis thaliana | This study |
| pZE-ER | pZE-luc carrying ER from Clostridium acetobutylicum | This study |
| pCS-ER | pCS27 carrying ER from C. acetobutylicum | This study |
| pCS-carsfp | pCS27 carrying Car from Mycobacterium marinum and Sfp from Bacillus subtilis | This study |
| pZE-RgTAL-ER | pZE-luc carrying TAL from Rhodobacter glutinis and ER from C.acetobutylicum | This study |
| Primer | Sequence 5′-3′ |
|---|---|
| CCR-4CL1-1-F-KpnI | gggaaaGGTACCatgcctgctgcggctccagc |
| CCR-4CL1-1-R-BamHI | gggaaaGGATCCttatttggtcggcagcggcaggtg |
| CCR-4CL1-2-F-BamHI | gggaaaGGATCCaggagatataccatggcgccacaagaacaagcagt |
| CCR-4CL1-2-R-XbaI | gggaaaTCTAGAttacaatccatttgctagttttgccctc |
| ER-KpnI-F | gggaaaGGTACCatgaacaaatacaagaaattatttgaaccaatcaaaattgg |
| ER-XbaI-F | gggaaaTCTAGAttatatatggtttgcaacttcaaaagcatccc |
| ER框-SpeI-F | gggaaaACTAGTaattgtgagcggataacaattgacattgtga |
| ER框-SacI-R | gggaaaGAGCTCacaacagataaaacgaaaggcccagtc |
| TAL框-SpeI-F | gggaaaACTAGTctcgagaattgtgagcggataacaattga |
| TAL框-SacI-R | gggaaaGAGCTCcgacaaacaacagataaaacgaaaggcc |
| Car-KpnI-F | gggaaaGGTACCatgtcacctatcacccgcgagg |
| Car-BamHI-R | gggaaaGGATCCtcacagcaagcccagcagac |
| sfp-BamHI-F | gggaaaGGATCCaggagatataccatgaagatttacggaatttatatgg |
| sfp-XbaI-R | GGGAAAtctagattataaaagctcttcgtacgagactattgtgat |
| AP-NheI-R | gggaaaGCTAGCttatttcttcagttcagccaggcttaacc |
| TA-NheI-F | gggaaaGCTAGCaggagatataccatgtcctcacgtaaagagcttg |
| APTA-XbaI-F | gggaaaTCTAGAatgacacaacctctttttctgatcggg |
| APTA-BamHI-R | gggaaaGGATCCttacccgcgacgcgctttta |
| pCS-tac-反-BamHI-F | gggaaaGGATCCgtcgccaatcacgcgtgaagagc |
| pCS-tac-反-XbaI-R | gggaaaCATATGttataaaagctcttcgtacgagacta |
| tac-Car框-F-SpeI | gggaaaACTAGTctcgagttgacaattaatcatcggctcg |
| tac-Car框-R-SacI | gggaaaGAGCTCcgacaaacaacagataaaacgaaaggcc |
表3 本实验所用到的引物
Tab. 3 Primers used in this study
| Primer | Sequence 5′-3′ |
|---|---|
| CCR-4CL1-1-F-KpnI | gggaaaGGTACCatgcctgctgcggctccagc |
| CCR-4CL1-1-R-BamHI | gggaaaGGATCCttatttggtcggcagcggcaggtg |
| CCR-4CL1-2-F-BamHI | gggaaaGGATCCaggagatataccatggcgccacaagaacaagcagt |
| CCR-4CL1-2-R-XbaI | gggaaaTCTAGAttacaatccatttgctagttttgccctc |
| ER-KpnI-F | gggaaaGGTACCatgaacaaatacaagaaattatttgaaccaatcaaaattgg |
| ER-XbaI-F | gggaaaTCTAGAttatatatggtttgcaacttcaaaagcatccc |
| ER框-SpeI-F | gggaaaACTAGTaattgtgagcggataacaattgacattgtga |
| ER框-SacI-R | gggaaaGAGCTCacaacagataaaacgaaaggcccagtc |
| TAL框-SpeI-F | gggaaaACTAGTctcgagaattgtgagcggataacaattga |
| TAL框-SacI-R | gggaaaGAGCTCcgacaaacaacagataaaacgaaaggcc |
| Car-KpnI-F | gggaaaGGTACCatgtcacctatcacccgcgagg |
| Car-BamHI-R | gggaaaGGATCCtcacagcaagcccagcagac |
| sfp-BamHI-F | gggaaaGGATCCaggagatataccatgaagatttacggaatttatatgg |
| sfp-XbaI-R | GGGAAAtctagattataaaagctcttcgtacgagactattgtgat |
| AP-NheI-R | gggaaaGCTAGCttatttcttcagttcagccaggcttaacc |
| TA-NheI-F | gggaaaGCTAGCaggagatataccatgtcctcacgtaaagagcttg |
| APTA-XbaI-F | gggaaaTCTAGAatgacacaacctctttttctgatcggg |
| APTA-BamHI-R | gggaaaGGATCCttacccgcgacgcgctttta |
| pCS-tac-反-BamHI-F | gggaaaGGATCCgtcgccaatcacgcgtgaagagc |
| pCS-tac-反-XbaI-R | gggaaaCATATGttataaaagctcttcgtacgagacta |
| tac-Car框-F-SpeI | gggaaaACTAGTctcgagttgacaattaatcatcggctcg |
| tac-Car框-R-SacI | gggaaaGAGCTCcgacaaacaacagataaaacgaaaggcc |
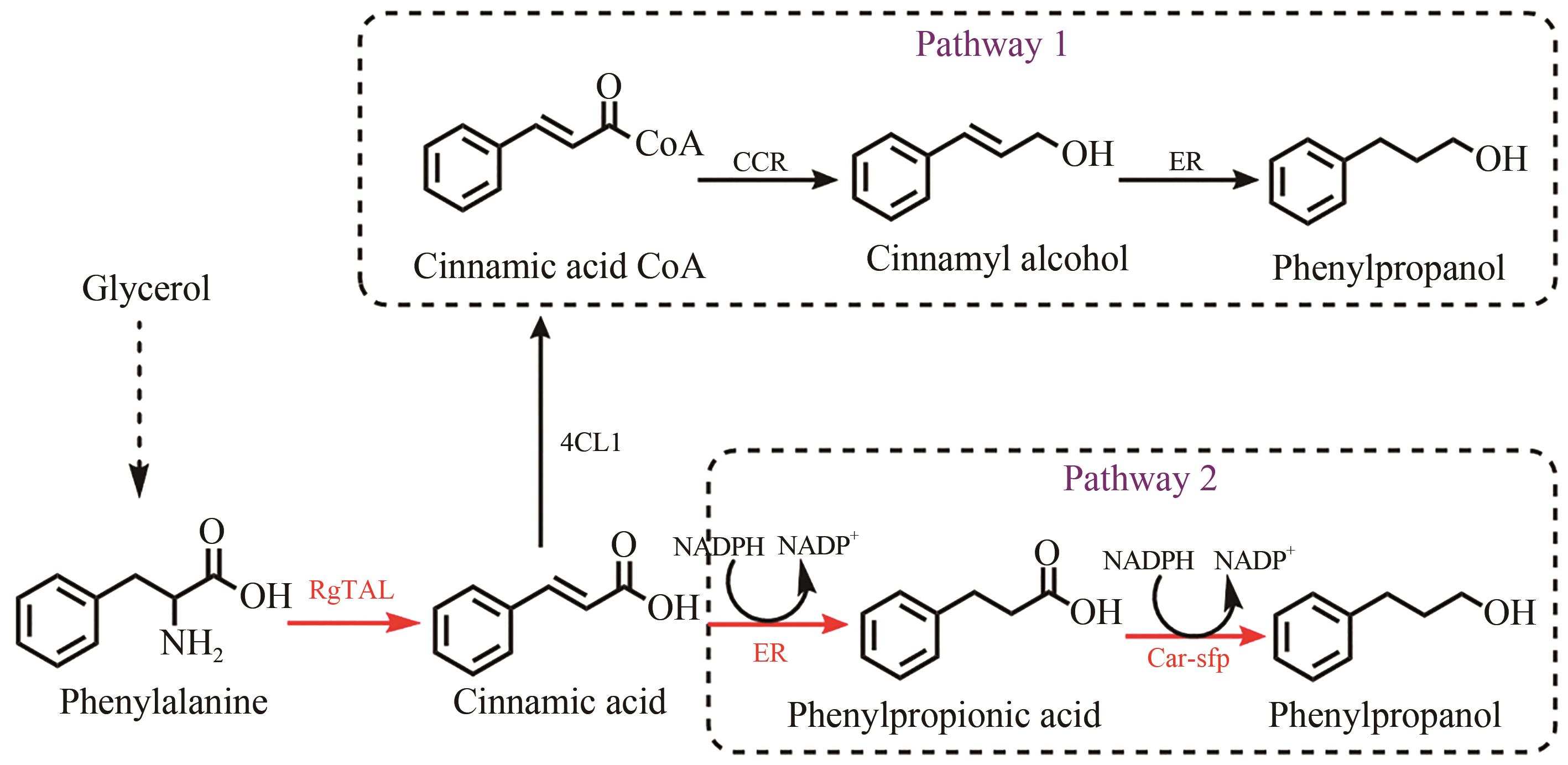
图1 3-苯丙醇合成途径的设计RgTALRgTAL—来自Rhodobacter glutinis的苯丙氨酸氨裂合酶;4CL1—来自Arabidopsis thaliana香豆酸辅酶A连接酶;CCR—来自Leucaena leucocephala的肉桂酰辅酶A还原酶;ER—来自Clostridium acetobutylicum的烯酸还原酶;Car—来自Mycobacterium marinum的羧酸还原酶;sfp—来自Bacillus subtilis的磷酸泛肽基转移酶
Fig. 1 Design of synthetic routes of 3-phenylpropanolRgTAL—phenylalanine ammonia lyase from Rhodobacter glutinis; 4CL1—from Arabidopsis thaliana coumarate-CoA ligase; CCR— cinnamoyl-CoA reductase from Leucaena leucocephala; ER—enoic acid reduction from Clostridium acetobutylicum Enzyme; Car—carboxylic acid reductase from Marine Mycobacterium; sfp—phosphoubiquitin transferase from Bacillus subtilis
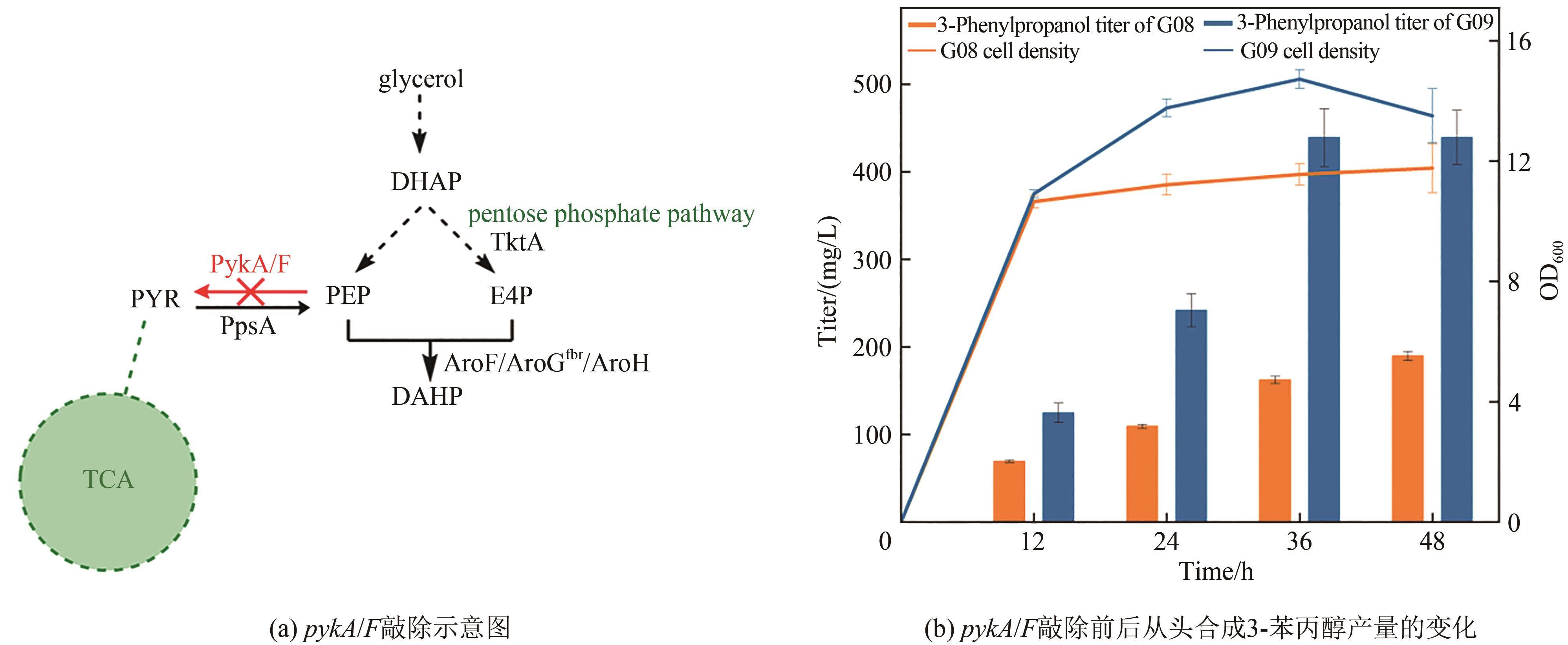
图5 pykA/F敲除对3-苯丙醇产量的影响DHAP—磷酸二羟丙酮;PYR—丙酮酸;PEP—磷酸烯醇式丙酮酸;E4P—D-赤藓糖-4-磷酸;DAHP—3-脱氧-D-阿拉伯糖庚酸七磷酸酯;TktA—转酮酶;PykA/F—丙酮酸激酶;PpsA—磷酸烯醇式丙酮酸合酶;AroF/AroGfbr/AroH—3-脱氧-D -阿拉伯糖庚酸七磷酸酯合酶
Fig. 5 Production of 3-phenylpropanol in pykA/F knockout strainDHAP—dihydroxyacetone phosphate; PYR—pyruvate; PEP—phosphoenolpyruvate; E4P—D-erythrose-4-phosphate; DAHP—3-deoxy-D-arabinoheptanoate heptaphosphate; TktA—Transketolase; PykA/F—pyruvate kinase; PpsA—phosphoenolpyruvate synthase; AroF/AroGfbr/AroH—3-deoxy-D-arabinoheptanoate heptaphosphate synthase
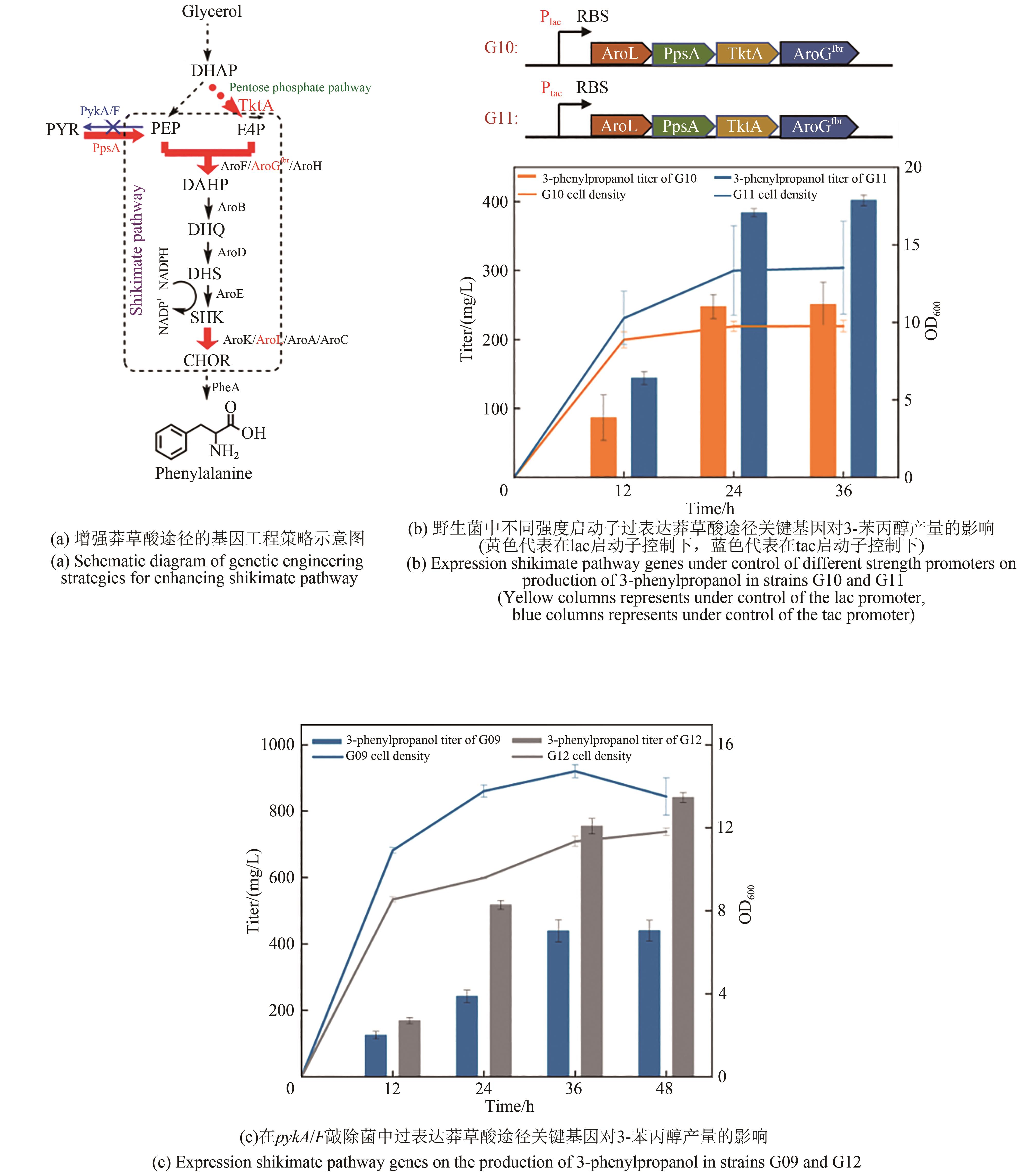
图6 增强莽草酸途径对3-苯丙醇产量的影响DHQ—3-脱氢奎尼酸;DHS—3-脱氢莽草酸;SHK—莽草酸;CHOR—分支酸;AroB—脱氢奎尼酸合成酶;AroD—脱氢奎尼酸脱水酶;AroE—莽草酸脱氢酶;AroK/AroL/AroA/AroC—脱氢莽草酸脱水酶;PheA—预苯酸脱水酶
Fig. 6 Strengthening of shikimate pathway on production of 3-phenylpropanolDHQ—3-dehydroquinic acid; DHS—3-dehydroshikimate; SHK—shikimic acid; CHOR—chorismate; AroB—dehydroquinic acid synthase; AroD—dehydroquinic acid dehydratase, AroE—Shikimate dehydrogenase; AroK/AroL/AroA/AroC—dehydroshikimate dehydratase; PheA—prephenate dehydrogenase
| 1 | BHATIA S P, WELLINGTON G A, COCCHIARA J, et al. Fragrance material review on 3-phenyl-1-propanol[J]. Food and Chemical Toxicology, 2011, 49: S246-S51. |
| 2 | SÁ A G A, MENESES A C D, ARAÚJO P H H D, et al. A review on enzymatic synthesis of aromatic esters used as flavor ingredients for food, cosmetics and pharmaceuticals industries[J]. Trends in Food Science & Technology, 2017, 69: 95-105. |
| 3 | LÜ J, YU C Q, GUO Y, et al. Gallstone disease and the risk of type 2 diabetes[J]. Scientific Reports, 2017, 7(1): 15853. |
| 4 | PANTEN J, SURBURG H. Flavors and fragrances (Ⅲ): Aromatic and heterocyclic compounds[J]. Ullmann's Encyclopedia of Industrial Chemistry, 2016: 1-45. |
| 5 | ZHOU Y Y, LI Z H, LIU Y B, et al. Regulating hydrogenation chemoselectivity of α, β-unsaturated aldehydes by combination of transfer and catalytic hydrogenation[J]. ChemSusChem, 2020, 13(7): 1746-1750. |
| 6 | MAO X Y, WANG Y. Cooperative carbon emission reduction through the Belt and Road Initiative[J]. Environmental Science and Pollution Research, 2021: 1-22. |
| 7 | YUAN X L, SHENG X R, CHEN L P, et al. Carbon footprint and embodied carbon transfer at the provincial level of the Yellow River Basin[J]. The Science of the Total Environment, 2022, 803: 149993. |
| 8 | KHALIL A S, COLLINS J J. Synthetic biology: applications come of age[J]. Nature Reviews Genetics, 2010, 11(5): 367-379. |
| 9 | RAN F A, HSU P D, WRIGHT J, et al. Genome engineering using the CRISPR-Cas9 system[J]. Nature Protocols, 2013, 8(11): 2281-308. |
| 10 | FRANGOUL H, ALTSHULER D, CAPPELLINI M D, et al. CRISPR-Cas9 gene editing for sickle cell disease and β-thalassemia[J]. The New England Journal of Medicine, 2021, 384(3): 252-260. |
| 11 | BIAN X Y, HUANG F, STEWART F A, et al. Direct cloning, genetic engineering, and heterologous expression of the syringolin biosynthetic gene cluster in E. coli through Red/ET recombineering[J]. ChemBioChem, 2012, 13(13): 1946-1952. |
| 12 | CORREA A, OPPEZZO P. Tuning different expression parameters to achieve soluble recombinant proteins in E. coli: advantages of high-throughput screening[J]. Biotechnology Journal, 2011, 6(6): 715-730. |
| 13 | WANG Y, LI Q G, ZHENG P, et al. Evolving the L-lysine high-producing strain of Escherichia coli using a newly developed high-throughput screening method[J]. Journal of Industrial Microbiology & Biotechnology, 2016, 43(9): 1227-1235. |
| 14 | LEE S Y, KIM H U, PARK J H, et al. Metabolic engineering of microorganisms: general strategies and drug production[J]. Drug Discovery Today, 2009, 14(1/2): 78-88. |
| 15 | SHEN X L, CHEN X, WANG J, et al. Design and construction of an artificial pathway for biosynthesis of acetaminophen in Escherichia coli [J]. Metabolic Engineering, 2021, 68: 26-33. |
| 16 | FENG J C, YANG C, ZHAO Z H, et al. Application of cell-free protein synthesis system for the biosynthesis of L-theanine[J]. ACS Synthetic Biology, 2021, 10(3): 620-631. |
| 17 | BOO Y C. p-Coumaric acid as an active ingredient in cosmetics: a review focusing on its antimelanogenic effects[J]. Antioxidants, 2019, 8(8): 275. |
| 18 | SHANG Y Z, WEI W P, ZHANG P, et al. Engineering Yarrowia lipolytica for enhanced production of arbutin[J]. Journal of Agricultural and Food Chemistry, 2020, 68(5): 1364-1372. |
| 19 | YIM H, HASELBECK R, NIU W, et al. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol[J]. Nature Chemical Biology, 2011, 7(7): 445-452. |
| 20 | ATSUMI S, LIAO J C. Metabolic engineering for advanced biofuels production from Escherichia coli [J]. Current Opinion in Biotechnology, 2008, 19(5): 414-419. |
| 21 | DZIGA D, LISZNIANSKA M, WLADYKA B. Bioreactor study employing bacteria with enhanced activity toward cyanobacterial toxins microcystins[J]. Toxins, 2014, 6(8): 2379-2392. |
| 22 | CHEN Z Y, SUN X X, LI Y, et al. Metabolic engineering of Escherichia coli for microbial synthesis of monolignols[J]. Metabolic Engineering, 2017, 39: 102-109. |
| 23 | SUN J, LIN Y H, SHEN X L, et al. Aerobic biosynthesis of hydrocinnamic acids in Escherichia coli with a strictly oxygen-sensitive enoate reductase[J]. Metabolic Engineering, 2016, 35: 75-82. |
| 24 | AKHTAR M K, TURNER N J, JONES P R. Carboxylic acid reductase is a versatile enzyme for the conversion of fatty acids into fuels and chemical commodities[J]. Proceedings of the National Academy of Sciences of the United States of America, 2013, 110(1): 87-92. |
| 25 | WANG J, LI C Y, ZOU Y S, et al. Bacterial synthesis of C3-C5 diols via extending amino acid catabolism[J]. Proceedings of the National Academy of Sciences of the United States of America, 2020, 117(32): 19159-19167. |
| 26 | VENKITASUBRAMANIAN P, DANIELS L, ROSAZZA J P. Reduction of carboxylic acids by Nocardia aldehyde oxidoreductase requires a phosphopantetheinylated enzyme[J]. JBC, 2007, 282(1): 478-485. |
| 27 | HALL M, STUECKLER C, HAUER B, et al. Asymmetric bioreduction of activated C=C bonds using Zymomonas mobilis NCR enoate reductase and old yellow enzymes OYE 1-3 from yeasts[J]. European Journal of Organic Chemistry, 2008, 2008(9): 1511-1516. |
| 28 | WANG S Y, ZHANG S W, XIAO A F, et al. Metabolic engineering of Escherichia coli for the biosynthesis of various phenylpropanoid derivatives[J]. Metabolic Engineering, 2015, 29: 153-159. |
| 29 | MAEDA H, DUDAREVA N. The shikimate pathway and aromatic amino acid biosynthesis in plants[J]. Annual Review of Plant Biology, 2012, 63: 73-105. |
| 30 | ROBERTS C W, ROBERTS F, LYONS R E, et al. The shikimate pathway and its branches in apicomplexan parasites[J]. The Journal of Infectious Diseases, 2002, 185(S1): S25-S36. |
| 31 | XU J Z, YU H B, HAN M, et al. Metabolic engineering of glucose uptake systems in Corynebacterium glutamicum for improving the efficiency of L-lysine production[J]. Journal of Industrial Microbiology & Biotechnology, 2019, 46(7): 937-949. |
| 32 | LINDNER S N, KNEBEL S, PALLERLA S R, et al. Cg2091 encodes a polyphosphate/ATP-dependent glucokinase of Corynebacterium glutamicum [J]. Applied Microbiology and Biotechnology, 2010, 87(2): 703-713. |
| 33 | LINDNER S N, SEIBOLD G M, HENRICH A, et al. Phosphotransferase system-independent glucose utilization in Corynebacterium glutamicum by inositol permeases and glucokinases[J]. Applied and Environmental Microbiology, 2011, 77(11): 3571-3581. |
| 34 | VOGT M, HAAS S, KLAFFL S, et al. Pushing product formation to its limit: metabolic engineering of Corynebacterium glutamicum for L-leucine overproduction[J]. Metabolic Engineering, 2014, 22: 40-52. |
| 35 | KHAMDUANG M, PACKDIBAMRUNG K, CHUTMANOP J, et al. Production of l-phenylalanine from glycerol by a recombinant Escherichia coli [J]. Journal of Industrial Microbiology & Biotechnology, 2009, 36(10): 1267-1274. |
| 36 | AHN J O, LEE H W, SAHA R, et al. Exploring the effects of carbon sources on the metabolic capacity for shikimic acid production in Escherichia coli using in silico metabolic predictions[J]. Journal of Microbiology and Biotechnology, 2008, 18(11): 1773-1784. |
| 37 | LEE S J, LEE D Y, KIM T Y, et al. Metabolic engineering of Escherichia coli for enhanced production of succinic acid, based on genome comparison and in silico gene knockout simulation[J]. Applied and Environmental Microbiology, 2005, 71(12): 7880-7887. |
| 38 | CHAVADI S, WOOFF E, COLDHAM N G, et al. Global effects of inactivation of the pyruvate kinase gene in the Mycobacterium tuberculosis complex[J]. Journal of Bacteriology, 2009, 191(24): 7545-7553. |
| 39 | POTTS A H, VAKULSKAS C A, PANNURI A, et al. Global role of the bacterial post-transcriptional regulator CsrA revealed by integrated transcriptomics[J]. Nature Communications, 2017, 8(1): 1596. |
| 40 | TATARKO M, ROMEO T. Disruption of a global regulatory gene to enhance central carbon flux into phenylalanine biosynthesis in Escherichia coli [J]. Current Microbiology, 2001, 43(1): 26-32. |
| 41 | SUZUKI K, BABITZKE P, KUSHNER S R, et al. Identification of a novel regulatory protein (CsrD) that targets the global regulatory RNAs CsrB and CsrC for degradation by RNase E[J]. Genes & Development, 2006, 20(18): 2605-2617. |
| 42 | YAKANDAWALA N, ROMEO T, FRIESEN A D, et al. Metabolic engineering of Escherichia coli to enhance phenylalanine production[J]. Applied Microbiology and Biotechnology, 2008, 78(2): 283-291. |
| 43 | JIANG M, ZHANG H R. Engineering the shikimate pathway for biosynthesis of molecules with pharmaceutical activities in E. coli [J]. Current Opinion in Biotechnology, 2016, 42: 1-6. |
| 44 | LI Z, DING D Q, WANG H Y, et al. Engineering Escherichia coli to improve tryptophan production via genetic manipulation of precursor and cofactor pathways[J]. Synthetic and Systems Biotechnology, 2020, 5(3): 200-205. |
| 45 | KNOP D R, DRATHS K M, CHANDRAN S S, et al. Hydroaromatic equilibration during biosynthesis of shikimic acid[J]. Journal of the American Chemical Society, 2001, 123(42): 10173-10182. |
| 46 | LU J-L, LIAO J C. Metabolic engineering and control analysis for production of aromatics: role of transaldolase[J]. Biotechnology and Bioengineering, 1997, 53(2): 132-138. |
| 47 | CHANDRAN S S, YI J, DRATHS K M, et al. Phosphoenolpyruvate availability and the biosynthesis of shikimic acid[J]. Biotechnology Progress, 2003, 19(3): 808-814. |
| 48 | ZHOU H Y, LIAO X Y, WANG T W, et al. Enhanced L-phenylalanine biosynthesis by co-expression of pheAfbr and aroFwt [J]. Bioresource Technology, 2010, 101(11): 4151-4156. |
| 49 | MCCANDLISS R J, POLING M D, HERRMANN K M. 3-Deoxy-D-arabino-heptulosonate 7-phosphate synthase. Purification and molecular characterization of the phenylalanine-sensitive isoenzyme from Escherichia coli [J]. JBC, 1978, 253(12): 4259-4265. |
| 50 | ELY B, PITTARD J. Aromatic amino acid biosynthesis: regulation of shikimate kinase in Escherichia coli K-12[J]. Journal of Bacteriology, 1979, 138(3): 933-43. |
| 51 | DEFEYTER R C, PITTARD J. Purification and properties of shikimate kinase Ⅱ from Escherichia coli K-12[J]. Journal of Bacteriology, 1986, 165(1): 331-333. |
| 52 | KIM B, BINKLEY R, KIM H U, et al. Metabolic engineering of Escherichia coli for the enhanced production of L-tyrosine[J]. Biotechnology and Bioengineering, 2018, 115(10): 2554-2564. |
| 53 | PARSONS C V, HARRIS D M M, PATTEN C L. Regulation of indole-3-acetic acid biosynthesis by branched-chain amino acids in Enterobacter cloacae UW5 [J]. FEMS Microbiology Letters, 2015, 362(18): fnv153. |
| 54 | GOTTARDI M, KNUDSEN J D, PRADO L, et al. De novo biosynthesis of trans-cinnamic acid derivatives in Saccharomyces cerevisiae [J]. Applied Microbiology and Biotechnology, 2017, 101(12): 4883-4893. |
| 55 | LIU Z N, ZHANG X, LEI D W, et al. Metabolic engineering of Escherichia coli for de novo production of 3-phenylpropanol via retrobiosynthesis approach[J]. Microbial Cell Factories, 2021, 20(1): 121. |
| 56 | KAUP B, BRINGER-MEYER S, SAHM H. Metabolic engineering of Escherichia coli: construction of an efficient biocatalyst for D-mannitol formation in a whole-cell biotransformation[J]. Applied Microbiology and Biotechnology, 2004, 64(3): 333-339. |
| 57 | WANG J, YANG Y P, ZHANG R H, et al. Microbial production of branched-chain dicarboxylate 2-methylsuccinic acid via enoate reductase-mediated bioreduction[J]. Metabolic Engineering, 2018, 45: 1-10. |
| [1] | 郭姝媛, 张倩楠, 姑丽克孜·买买提热夏提, 杨一群, 于涛. 液体生物燃料合成与炼制的研究进展[J]. 合成生物学, 2025, 6(1): 18-44. |
| [2] | 邵明威, 孙思勉, 杨时茂, 陈国强. 基于极端微生物的生物制造[J]. 合成生物学, 2024, 5(6): 1419-1436. |
| [3] | 赵亮, 李振帅, 付丽平, 吕明, 王士安, 张全, 刘立成, 李福利, 刘自勇. 生物转化一碳化合物原料产油脂与单细胞蛋白研究进展[J]. 合成生物学, 2024, 5(6): 1300-1318. |
| [4] | 竺方欢, 岑雪聪, 陈振. 微生物合成二元醇研究进展[J]. 合成生物学, 2024, 5(6): 1367-1385. |
| [5] | 禹伟, 高教琪, 周雍进. 一碳生物转化合成有机酸的研究进展[J]. 合成生物学, 2024, 5(5): 1169-1188. |
| [6] | 陈锡玮, 张华然, 邹懿. 真菌源非核糖体肽类药物生物合成及代谢工程[J]. 合成生物学, 2024, 5(3): 571-592. |
| [7] | 谢皇, 郑义蕾, 苏依婷, 阮静怡, 李永泉. 放线菌聚酮类化合物生物合成体系重构研究进展[J]. 合成生物学, 2024, 5(3): 612-630. |
| [8] | 惠真, 唐啸宇. CRISPR/Cas9编辑系统在微生物天然产物研究中的应用[J]. 合成生物学, 2024, 5(3): 658-671. |
| [9] | 赵静宇, 张健, 祁庆生, 王倩. 基于细菌双组分系统的生物传感器的研究进展[J]. 合成生物学, 2024, 5(1): 38-52. |
| [10] | 孙绘梨, 崔金玉, 栾国栋, 吕雪峰. 面向高效光驱固碳产醇的蓝细菌合成生物技术研究进展[J]. 合成生物学, 2023, 4(6): 1161-1177. |
| [11] | 晏雄鹰, 王振, 娄吉芸, 张皓瑜, 黄星宇, 王霞, 杨世辉. 生物燃料高效生产微生物细胞工厂构建研究进展[J]. 合成生物学, 2023, 4(6): 1082-1121. |
| [12] | 程真真, 张健, 高聪, 刘立明, 陈修来. 代谢工程改造微生物利用甲酸研究进展[J]. 合成生物学, 2023, 4(4): 756-778. |
| [13] | 刘家宇, 杨智晗, 杨蕾, 朱丽英, 朱政明, 江凌. 合成生物技术驱动酪丁酸梭菌细胞工厂开发的研究进展[J]. 合成生物学, 2022, 3(6): 1174-1200. |
| [14] | 郭姝媛, 吴良焕, 刘香健, 王博, 于涛. 微生物中一碳代谢网络构建的进展与挑战[J]. 合成生物学, 2022, 3(1): 116-137. |
| [15] | 陈久洲, 王钰, 蒲伟, 郑平, 孙际宾. 5-氨基乙酰丙酸生物合成技术的发展及展望[J]. 合成生物学, 2021, 2(6): 1000-1016. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||