Biocontainment strategies of engineered bacteria
GAN Mudan, ZUO Jingrui, CAO Youzhi
- Functional Nano and Soft Materials (FUNSOM),Soochow University,Suzhou 215123,Jiangsui,China
-
Received:2025-02-14Revised:2025-04-27Published:2025-05-06 -
Contact:GAN Mudan
工程菌的生物安全防控策略
甘牡丹, 左静蕊, 曹友志
- 苏州大学,功能纳米与软物质研究院,江苏 苏州 215123
-
通讯作者:甘牡丹 -
作者简介:甘牡丹 (1987—),女,硕士,实验师。研究方向为基因工程改造病毒、工程菌和病原微生物的生物安全评价和实验室管理。E-mail:gmd@suda.edu.cn
CLC Number:
Cite this article
GAN Mudan, ZUO Jingrui, CAO Youzhi. Biocontainment strategies of engineered bacteria[J]. Synthetic Biology Journal, DOI: 10.12211/2096-8280.2025-010.
甘牡丹, 左静蕊, 曹友志. 工程菌的生物安全防控策略[J]. 合成生物学, DOI: 10.12211/2096-8280.2025-010.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2025-010
| 生物安全防控策略 | 实例 |
|---|---|
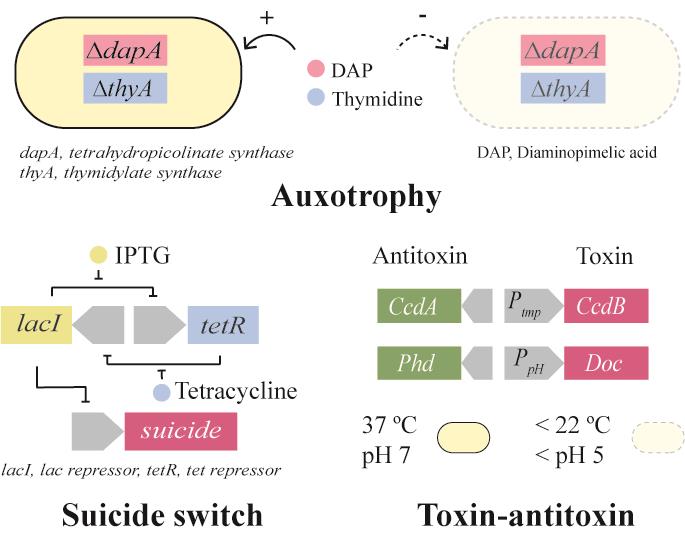
| 营养缺陷 | dapA基因敲除的营养缺陷[ |
| 自杀基因调控开关 | ATc调控的Deadman自杀基因开关;LacI-GalR融合转录因子的Passcode自杀基因开关[ |
| 毒素-抗毒素 | 温度响应的CcdB-CcdA毒素-抗毒素分子对[ |
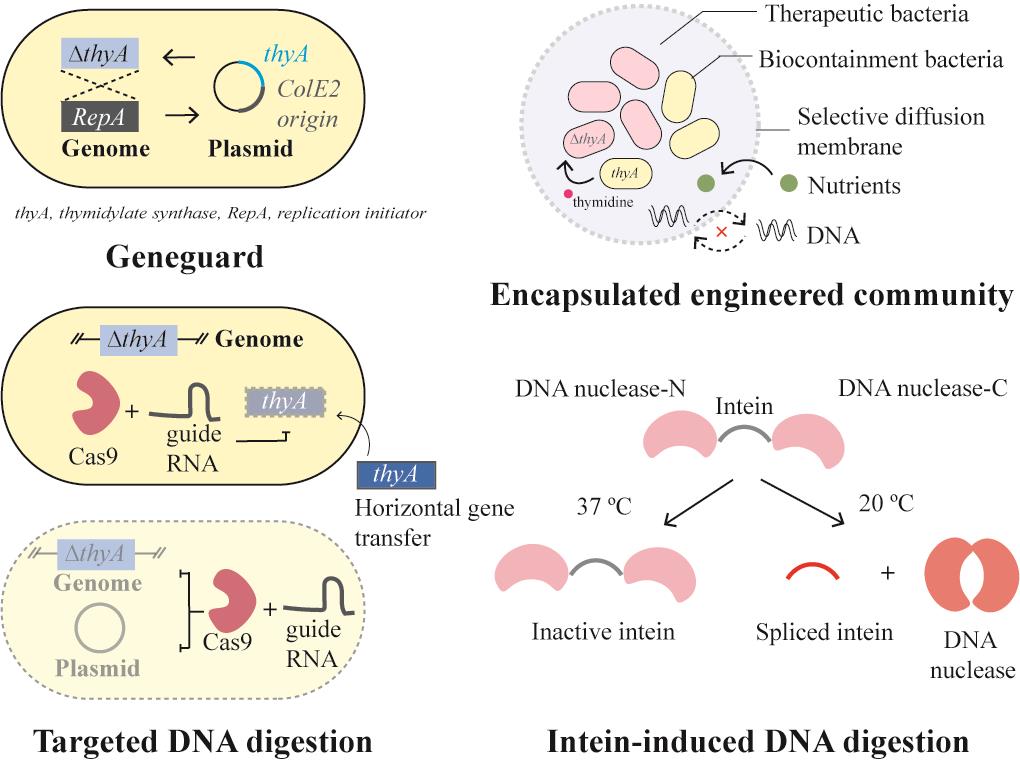
| 基因元件拆分 | Geneguard质粒宿主依赖系统[ |
| DNA降解 | 靶向外源基因的Cas9系统[ |
| 降低突变 | 基因组插入序列元件IS敲除[ |
| 物理隔离 | GelMA包裹的工程菌[ |
Table 1 Examples of biocontainment strategies
| 生物安全防控策略 | 实例 |
|---|---|
| 营养缺陷 | dapA基因敲除的营养缺陷[ |
| 自杀基因调控开关 | ATc调控的Deadman自杀基因开关;LacI-GalR融合转录因子的Passcode自杀基因开关[ |
| 毒素-抗毒素 | 温度响应的CcdB-CcdA毒素-抗毒素分子对[ |
| 基因元件拆分 | Geneguard质粒宿主依赖系统[ |
| DNA降解 | 靶向外源基因的Cas9系统[ |
| 降低突变 | 基因组插入序列元件IS敲除[ |
| 物理隔离 | GelMA包裹的工程菌[ |
| 1 | GURBATRI C R, ARPAIA N, DANINO T. Engineering bacteria as interactive cancer therapies[J]. Science, 2022, 378(6622): 858-864. |
| 2 | KIM K, KANG M, CHO B K. Systems and synthetic biology-driven engineering of live bacterial therapeutics[J]. Frontiers in Bioengineering and Biotechnology, 2023, 11: 1267378. |
| 3 | RAMAN V, DESHPANDE C P, KHANDUJA S, et al. Build-a-bug workshop: using microbial-host interactions and synthetic biology tools to create cancer therapies[J]. Cell Host & Microbe, 2023, 31(10): 1574-1592. |
| 4 | CIOCAN D, ELINAV E. Engineering bacteria to modulate host metabolism[J]. Acta Physiologica, 2023, 238(3): e14001. |
| 5 | ZHAI L, FU L Y, WEI W, et al. Advances of bacterial biomaterials for disease therapy[J]. ACS Synthetic Biology, 2024, 13(5): 1400-1411. |
| 6 | YAN S Z, GAN Y, XU H Z, et al. Bacterial carrier-mediated drug delivery systems: a promising strategy in cancer therapy[J]. Frontiers in Bioengineering and Biotechnology, 2025, 12: 1526612. |
| 7 | DEY S, SANKARAN S. Engineered bacterial therapeutics with material solutions[J]. Trends in Biotechnology, 2024, 42(12): 1663-1676. |
| 8 | LEE J W, CHAN C T Y, SLOMOVIC S, et al. Next-generation biocontainment systems for engineered organisms[J]. Nature Chemical Biology, 2018, 14(6): 530-537. |
| 9 | ASIN-GARCIA E, KALLERGI A, LANDEWEERD L, et al. Genetic safeguards for safety-by-design: so close yet so far[J]. Trends in Biotechnology, 2020, 38(12): 1308-1312. |
| 10 | PARKER M T, KUNJAPUR A M. Deployment of engineered microbes: contributions to the bioeconomy and considerations for biosecurity[J]. Health Security, 2020, 18(4): 278-296. |
| 11 | ARNOLDS K L, DAHLIN L R, DING L, et al. Biotechnology for secure biocontainment designs in an emerging bioeconomy[J]. Current Opinion in Biotechnology, 2021, 71: 25-31. |
| 12 | OU Y K, GUO S J. Safety risks and ethical governance of biomedical applications of synthetic biology[J]. Frontiers in Bioengineering and Biotechnology, 2023, 11: 1292029. |
| 13 | HALAWA E M, FADEL M, AL-RABIA M W, et al. Antibiotic action and resistance: updated review of mechanisms, spread, influencing factors, and alternative approaches for combating resistance[J]. Frontiers in Pharmacology, 2024, 14: 1305294. |
| 14 | PANTOJA ANGLES A, VALLE-PÉREZ A U, HAUSER C, et al. Microbial biocontainment systems for clinical, agricultural, and industrial applications[J]. Frontiers in Bioengineering and Biotechnology, 2022, 10: 830200. |
| 15 | HUANG Y, LIN X J, YU S Y, et al. Intestinal engineered probiotics as living therapeutics: chassis selection, colonization enhancement, gene circuit design, and biocontainment[J]. ACS Synthetic Biology, 2022, 11(10): 3134-3153. |
| 16 | ARBOLEDA-GARCÍA A, ALARCON-RUIZ I, BOADA-ACOSTA L, et al. Advancements in synthetic biology-based bacterial cancer therapy: a modular design approach[J]. Critical Reviews in Oncology/Hematology, 2023, 190: 104088. |
| 17 | MOE-BEHRENS G H G, DAVIS R, HAYNES K A. Preparing synthetic biology for the world[J]. Frontiers in Microbiology, 2013, 4: 5. |
| 18 | WILSON D J. NIH guidelines for research involving recombinant DNA molecules[J]. Accountability in Research, 1993, 3(2-3): 177-185. |
| 19 | FENG G Q, HUANG H N, CHEN Y G. Effects of emerging pollutants on the occurrence and transfer of antibiotic resistance genes: a review[J]. Journal of Hazardous Materials, 2021, 420: 126602. |
| 20 | WANG S M, LI W, XI B D, et al. Mechanisms and influencing factors of horizontal gene transfer in composting system: a review[J]. Science of The Total Environment, 2024, 955: 177017. |
| 21 | 中华人民共和国科学技术部. 基因工程安全管理办法[EB/OL]. [2025-02-12]. . |
| 22 | NELSON M T, CHARBONNEAU M R, COIA H G, et al. Characterization of an engineered live bacterial therapeutic for the treatment of phenylketonuria in a human gut-on-a-chip[J]. Nature Communications, 2021, 12: 2805. |
| 23 | LEVENTHAL D S, SOKOLOVSKA A, LI N, et al. Immunotherapy with engineered bacteria by targeting the STING pathway for anti-tumor immunity[J]. Nature Communications, 2020, 11: 2739. |
| 24 | CHAN C T Y, LEE J W, CAMERON D E, et al. 'Deadman' and 'Passcode' microbial kill switches for bacterial containment[J]. Nature Chemical Biology, 2016, 12(2): 82-86. |
| 25 | FERRY Q R V, LYUTOVA R, FULGA T A. Rational design of inducible CRISPR guide RNAs for de novo assembly of transcriptional programs[J]. Nature Communications, 2017, 8: 14633. |
| 26 | STIRLING F, NAYDICH A, BRAMANTE J, et al. Synthetic cassettes for pH-mediated sensing, counting, and containment[J]. Cell Reports, 2020, 30(9): 3139-3148.e4. |
| 27 | WRIGHT O, DELMANS M, STAN G B, et al. GeneGuard: a modular plasmid system designed for biosafety[J]. ACS Synthetic Biology, 2015, 4(3): 307-316. |
| 28 | HUANG S Q, LEE A J, TSOI R, et al. Coupling spatial segregation with synthetic circuits to control bacterial survival[J]. Molecular Systems Biology, 2016, 12(2): 859. |
| 29 | HAYASHI N, LAI Y, FUERTE-STONE J, et al. Cas9-assisted biological containment of a genetically engineered human commensal bacterium and genetic elements[J]. Nature Communications, 2024, 15: 2096. |
| 30 | FOO G W, LEICHTHAMMER C D, SAITA I M, et al. Intein-based thermoregulated meganucleases for containment of genetic material[J]. Nucleic Acids Research, 2024, 52(4): 2066-2077. |
| 31 | CAI Y Z, AGMON N, CHOI W J, et al. Intrinsic biocontainment: multiplex genome safeguards combine transcriptional and recombinational control of essential yeast genes[J]. Proceedings of the National Academy of Sciences of the United States of America, 2015, 112(6): 1803-1808. |
| 32 | CALLAWAY E. 'Minimal' cell raises stakes in race to harness synthetic life[J]. Nature, 2016, 531(7596): 557-558. |
| 33 | WU F, LIN S S, LUO H L, et al. Noncontact microbiota transplantation by core-shell microgel-enabled nonleakage envelopment[J]. Science Advances, 2025, 11(6): eadr7373. |
| 34 | ZOU Z P, CAI Z H, ZHANG X P, et al. Delivery of encapsulated intelligent engineered probiotic for inflammatory bowel disease therapy[J]. Advanced Healthcare Materials, 2025, 14(3): 2403704. |
| 35 | INDA-WEBB M E, JIMENEZ M, LIU Q, et al. Sub-1.4 Cm3 capsule for detecting labile inflammatory biomarkers in situ [J]. Nature, 2023, 620(7973): 386-392. |
| 36 | HIROTA R, ABE K, KATSUURA Z I, et al. A novel biocontainment strategy makes bacterial growth and survival dependent on phosphite[J]. Scientific Reports, 2017, 7: 44748. |
| 37 | AGMON N, TANG Z J, YANG K, et al. Low escape-rate genome safeguards with minimal molecular perturbation of Saccharomyces cerevisiae [J]. Proceedings of the National Academy of Sciences of the United States of America, 2017, 114(8): E1470-E1479. |
| 38 | ASIN-GARCIA E, BATIANIS C, LI Y S, et al. Phosphite synthetic auxotrophy as an effective biocontainment strategy for the industrial chassis Pseudomonas putida [J]. Microbial Cell Factories, 2022, 21(1): 156. |
| 39 | SEBESTA J, XIONG W, GUARNIERI M T, et al. Biocontainment of genetically engineered algae[J]. Frontiers in Plant Science, 2022, 13: 839446. |
| 40 | BONGAERTS N, EDOO Z, ABUKAR A A, et al. Low-cost anti-mycobacterial drug discovery using engineered E. coli [J]. Nature Communications, 2022, 13: 3905. |
| 41 | HEDIN K A, KRUSE V, VAZQUEZ-URIBE R, et al. Biocontainment strategies for in vivo applications of Saccharomyces boulardii [J]. Frontiers in Bioengineering and Biotechnology, 2023, 11: 1136095. |
| 42 | RUEDA-MEJIA M P, BÜHLMANN A, ORTIZ-MERINO R A, et al. Pantothenate auxotrophy in a naturally occurring biocontrol yeast[J]. Applied and Environmental Microbiology, 2023, 89(7): e00884-23. |
| 43 | LINDNER S N, RAMIREZ L C, KRÜSEMANN J L, et al. NADPH-auxotrophic E. coli: a sensor strain for testing in vivo regeneration of NADPH[J]. ACS Synthetic Biology, 2018, 7(12): 2742-2749. |
| 44 | ISABELLA V M, HA B N, CASTILLO M J, et al. Development of a synthetic live bacterial therapeutic for the human metabolic disease phenylketonuria[J]. Nature Biotechnology, 2018, 36(9): 857-864. |
| 45 | KURTZ C B, MILLET Y A, PUURUNEN M K, et al. An engineered E. coli Nissle improves hyperammonemia and survival in mice and shows dose-dependent exposure in healthy humans[J]. Science Translational Medicine, 2019, 11(475): eaau7975. |
| 46 | LUKE J J, PIHA-PAUL S A, MEDINA T, et al. Phase I study of SYNB1891, an engineered E. coli nissle strain expressing STING agonist, with and without atezolizumab in advanced malignancies[J]. Clinical Cancer Research, 2023, 29(13): 2435-2444. |
| 47 | ATA Ö, MATTANOVICH D. Into the metabolic wild: Unveiling hidden pathways of microbial metabolism[J]. Microbial Biotechnology, 2024, 17(8): e14548. |
| 48 | JONES B S, LAMB L S, GOLDMAN F, et al. Improving the safety of cell therapy products by suicide gene transfer[J]. Frontiers in Pharmacology, 2014, 5: 254. |
| 49 | ČELEŠNIK H, TANŠEK A, TAHIROVIĆ A, et al. Biosafety of biotechnologically important microalgae: intrinsic suicide switch implementation in Cyanobacterium synechocystis sp. PCC 6803[J]. Biology Open, 2016, 5(4): 519-528. |
| 50 | ZHOU Y Q, SUN T, CHEN Z X, et al. Development of a new biocontainment strategy in model Cyanobacterium Synechococcus strains[J]. ACS Synthetic Biology, 2019, 8(11): 2576-2584. |
| 51 | HOFFMANN S A, CAI Y Z. Engineering stringent genetic biocontainment of yeast with a protein stability switch[J]. Nature Communications, 2024, 15: 1060. |
| 52 | VARMA S, GULATI K A, SRIRAMAKRISHNAN J, et al. Environment signal dependent biocontainment systems for engineered organisms: Leveraging triggered responses and combinatorial systems[J]. Synthetic and Systems Biotechnology, 2025, 10(2): 356-364. |
| 53 | MU Z P, ZOU Z N, YANG Y, et al. A genetically engineered Escherichia coli that senses and degrades tetracycline antibiotic residue[J]. Synthetic and Systems Biotechnology, 2018, 3(3): 196-203. |
| 54 | HALVORSEN T M, RICCI D P, PARK D M, et al. Comparison of kill switch toxins in plant-beneficial Pseudomonas fluorescens reveals drivers of lethality, stability, and escape[J]. ACS Synthetic Biology, 2022, 11(11): 3785-3796. |
| 55 | JURĖNAS D, FRAIKIN N, GOORMAGHTIGH F, et al. Biology and evolution of bacterial toxin–antitoxin systems[J]. Nature Reviews Microbiology, 2022, 20(6): 335-350. |
| 56 | SOUCY S M, HUANG J L, GOGARTEN J P. Horizontal gene transfer: building the web of life[J]. Nature Reviews Genetics, 2015, 16(8): 472-482. |
| 57 | WOLFF J H, MIKKELSEN J G. Delivering genes with human immunodeficiency virus-derived vehicles: still state-of-the-art after 25 years[J]. Journal of Biomedical Science, 2022, 29(1): 79. |
| 58 | ENG A, BORENSTEIN E. Microbial community design: methods, applications, and opportunities[J]. Current Opinion in Biotechnology, 2019, 58: 117-128. |
| 59 | IBRAHIM M, RAAJARAAM L, RAMAN K. Modelling microbial communities: Harnessing consortia for biotechnological applications[J]. Computational and Structural Biotechnology Journal, 2021, 19: 3892-3907. |
| 60 | MIMEE M, CITORIK R J, LU T K. Microbiome therapeutics- advances and challenges[J]. Advanced Drug Delivery Reviews, 2016, 105(Pt A): 44-54. |
| 61 | KE J, WANG B, YOSHIKUNI Y. Microbiome engineering: synthetic biology of plant-associated microbiomes in sustainable agriculture[J]. Trends in Biotechnology, 2021, 39(3): 244-261. |
| 62 | RAPP K M, JENKINS J P, BETENBAUGH M J. Partners for life: building microbial consortia for the future[J]. Current Opinion in Biotechnology, 2020, 66: 292-300. |
| 63 | OZDEMIR T, FEDOREC A J H, DANINO T, et al. Synthetic biology and engineered live biotherapeutics: toward increasing system complexity[J]. Cell Systems, 2018, 7(1): 5-16. |
| 64 | DOU J, BENNETT M R. Synthetic biology and the gut microbiome[J]. Biotechnology Journal, 2018, 13(5): 1700159. |
| 65 | NEVOT G, SANTOS-MORENO J, CAMPAMÀ-SANZ N, et al. Synthetically programmed antioxidant delivery by a domesticated skin commensal[J]. Cell Systems, 2025, 16(2): 101169. |
| 66 | VANARSDALE E, NAVID A, CHU M J, et al. Electrogenetic signaling and information propagation for controlling microbial consortia via programmed lysis[J]. Biotechnology and Bioengineering, 2023, 120(5): 1366-1381. |
| 67 | CALIANDO B J, VOIGT C A. Targeted DNA degradation using a CRISPR device stably carried in the host genome[J]. Nature Communications, 2015, 6: 6989. |
| 68 | ROTTINGHAUS A G, FERREIRO A, FISHBEIN S R S, et al. Genetically stable CRISPR-based kill switches for engineered microbes[J]. Nature Communications, 2022, 13: 672. |
| 69 | ASIN-GARCIA E, MARTIN-PASCUAL M, GARCIA-MORALES L, et al. ReScribe: an unrestrained tool combining multiplex recombineering and minimal-PAM ScCas9 for genome recoding Pseudomonas putida [J]. ACS Synthetic Biology, 2021, 10(10): 2672-2688. |
| 70 | HARTIG A M, DAI W T, ZHANG K, et al. Influence of environmental conditions on the escape rates of biocontained genetically engineered microbes[J]. Environmental Science & Technology, 2024, 58(51): 22657-22667. |
| 71 | ASIN-GARCIA E, MARTIN-PASCUAL M, DE BUCK C, et al. GenoMine: a CRISPR-Cas9-based kill switch for biocontainment of Pseudomonas putida [J]. Frontiers in Bioengineering and Biotechnology, 2024, 12: 1426107. |
| 72 | PANTOJA ANGLES A, ALI Z, MAHFOUZ M. CS-cells: a CRISPR-Cas12 DNA device to generate chromosome-shredded cells for efficient and safe molecular biomanufacturing[J]. ACS Synthetic Biology, 2022, 11(1): 430-440. |
| 73 | YANG B, WU C, TENG Y X, et al. Tailoring microbial fitness through computational steering and CRISPRi-driven robustness regulation[J]. Cell Systems, 2024, 15(12): 1133-1147.e4. |
| 74 | YANG Z K, LUO H, ZHANG Y M, et al. Pan-genomic analysis provides novel insights into the association of E.coli with human host and its minimal genome[J]. Bioinformatics, 2019, 35(12): 1987-1991. |
| 75 | TANG T C, THAM E, LIU X Y, et al. Hydrogel-based biocontainment of bacteria for continuous sensing and computation[J]. Nature Chemical Biology, 2021, 17(6): 724-731. |
| 76 | DATTA D, WEISS E L, WANGPRASEURT D, et al. Phenotypically complex living materials containing engineered cyanobacteria[J]. Nature Communications, 2023, 14: 4742. |
| 77 | CHE H C, WANG Z Y, LI Y, et al. A stable and sensitive engineering bacterial sensor via physical biocontainment and two-stage signal amplification[J]. Analytical Chemistry, 2024, 96(21): 8807-8813. |
| 78 | ROMESBERG F E. Creation, optimization, and use of semi-synthetic organisms that store and retrieve increased genetic information[J]. Journal of Molecular Biology, 2022, 434(8): 167331. |
| 79 | KIMOTO M, HIRAO I. Genetic alphabet expansion technology by creating unnatural base pairs[J]. Chemical Society Reviews, 2020, 49(21): 7602-7626. |
| 80 | KIMOTO M, HIRAO I. Genetic code engineering by natural and unnatural base pair systems for the site-specific incorporation of non-standard amino acids into proteins[J]. Frontiers in Molecular Biosciences, 2022, 9: 851646. |
| 81 | GERECHT K, FREUND N, LIU W, et al. The expanded central dogma: genome resynthesis, orthogonal biosystems, synthetic genetics[J]. Annual Review of Biophysics, 2023, 52: 413-432. |
| 82 | DÖRRENHAUS R, WAGNER P K, KATH-SCHORR S. Two are not enough: synthetic strategies and applications of unnatural base pairs[J]. Biological Chemistry, 2023, 404(10): 883-896. |
| 83 | AWAWDEH A, RADECKI A A, VARGAS-RODRIGUEZ O. Suppressor tRNAs at the interface of genetic code expansion and medicine[J]. Frontiers in Genetics, 2024, 15: 1420331. |
| 84 | COSTELLO A, PETERSON A A, CHEN P H, et al. Genetic code expansion history and modern innovations[J]. Chemical Reviews, 2024, 124(21): 11962-12005. |
| 85 | YI H B, LEE S, SEO K, et al. Cellular and biophysical applications of genetic code expansion[J]. Chemical Reviews, 2024, 124(11): 7465-7530. |
| 86 | KIM Y J, CHO S H, KIM J C, et al. tRNA engineering strategies for genetic code expansion[J]. Frontiers in Genetics, 2024, 15: 1373250. |
| 87 | GÓMEZ-TATAY L, HERNÁNDEZ-ANDREU J M. Xenobiology for the biocontainment of synthetic organisms: opportunities and challenges[J]. Life, 2024, 14(8): 996. |
| 88 | MARLIÈRE P, PATROUIX J, DÖRING V, et al. Chemical evolution of a bacterium’s genome[J]. Angewandte Chemie International Edition, 2011, 50(31): 7109-7114. |
| 89 | MALYSHEV D A, DHAMI K, LAVERGNE T, et al. A semi-synthetic organism with an expanded genetic alphabet[J]. Nature, 2014, 509(7500): 385-388. |
| 90 | ZHANG Y, PTACIN J L, FISCHER E C, et al. A semi-synthetic organism that stores and retrieves increased genetic information[J]. Nature, 2017, 551(7682): 644-647. |
| 91 | MUKAI T, LAJOIE M J, ENGLERT M, et al. Rewriting the genetic code[J]. Annual Review of Microbiology, 2017, 71: 557-577. |
| 92 | WANG L, BROCK A, HERBERICH B, et al. Expanding the genetic code of Escherichia coli [J]. Science, 2001, 292(5516): 498-500. |
| 93 | KATO Y. Translational control using an expanded genetic code[J]. International Journal of Molecular Sciences, 2019, 20(4): 887. |
| 94 | CHANG T T, DING W C, YAN S R, et al. A robust yeast biocontainment system with two-layered regulation switch dependent on unnatural amino acid[J]. Nature Communications, 2023, 14: 6487. |
| 95 | KATO Y. An engineered bacterium auxotrophic for an unnatural amino acid: a novel biological containment system[J]. PeerJ, 2015, 3: e1247. |
| 96 | KATO Y. Extremely low leakage expression systems using dual transcriptional-translational control for toxic protein production[J]. International Journal of Molecular Sciences, 2020, 21(3): 705. |
| 97 | KURU E, MÄÄTTÄLÄ R M, NOGUERA K, et al. Release factor inhibiting antimicrobial peptides improve nonstandard amino acid incorporation in wild-type bacterial cells[J]. ACS Chemical Biology, 2020, 15(7): 1852-1861. |
| 98 | GAO X W, SUN Y J, YANG Y H, et al. Directed evolution of hydroxylase XcP4H for enhanced 5-HTP production in engineered probiotics to treat depression[J]. International Journal of Biological Macromolecules, 2025, 307: 142250. |
| 99 | ROVNER A J, HAIMOVICH A D, KATZ S R, et al. Recoded organisms engineered to depend on synthetic amino acids[J]. Nature, 2015, 518(7537): 89-93. |
| 100 | XUAN W M, SCHULTZ P G. A strategy for creating organisms dependent on noncanonical amino acids[J]. Angewandte Chemie International Edition, 2017, 56(31): 9170-9173. |
| 101 | TACK D S, ELLEFSON J W, THYER R, et al. Addicting diverse bacteria to a noncanonical amino acid[J]. Nature Chemical Biology, 2016, 12(3): 138-140. |
| 102 | MANDELL D J, LAJOIE M J, MEE M T, et al. Biocontainment of genetically modified organisms by synthetic protein design[J]. Nature, 2015, 518(7537): 55-60. |
| 103 | KOH M, NASERTORABI F, HAN G W, et al. Generation of an orthogonal protein-protein interface with a noncanonical amino acid[J]. Journal of the American Chemical Society, 2017, 139(16): 5728-5731. |
| 104 | GAN F, LIU R H, WANG F, et al. Functional replacement of histidine in proteins to generate noncanonical amino acid dependent organisms[J]. Journal of the American Chemical Society, 2018, 140(11): 3829-3832. |
| 105 | LAJOIE M J, ROVNER A J, GOODMAN D B, et al. Genomically recoded organisms expand biological functions[J]. Science, 2013, 342(6156): 357-360. |
| 106 | FREDENS J, WANG K H, DE LA TORRE D, et al. Total synthesis of Escherichia coli with a recoded genome[J]. Nature, 2019, 569(7757): 514-518. |
| 107 | NYERGES A, VINKE S, FLYNN R, et al. A swapped genetic code prevents viral infections and gene transfer[J]. Nature, 2023, 615(7953): 720-727. |
| 108 | CHOI Y N, KIM D, LEE S, et al. Quadruplet Codon decoding-based versatile genetic biocontainment system[J]. Nucleic Acids Research, 2025, 53(1): gkae1292. |
| 109 | LAWSON C E, MARTÍ J M, RADIVOJEVIC T, et al. Machine learning for metabolic engineering: a review[J]. Metabolic Engineering, 2021, 63: 34-60. |
| 110 | DIANAWATI D, MISHRA V, SHAH N P. Survival of microencapsulated probiotic bacteria after processing and during storage: a review[J]. Critical Reviews in Food Science and Nutrition, 2016, 56(10): 1685-1716. |
| 111 | NGUYEN T T, NGUYEN P T, PHAM M N, et al. Synbiotics: a new route of self-production and applications to human and animal health[J]. Probiotics and Antimicrobial Proteins, 2022, 14(5): 980-993. |
| 112 | HASSANISAADI M, VATANKHAH M, KENNEDY J F, et al. Advancements in xanthan gum: a macromolecule for encapsulating plant probiotic bacteria with enhanced properties[J]. Carbohydrate Polymers, 2025, 348: 122801. |
| 113 | 中华人民共和国生态环境部. 中华人民共和国生物安全法[EB/OL]. [2025-02-12]. . |
| [1] | CHEN Yingying, LIU Yang, SHI Junjie, MA Junying, JU Jianhua. CRISPR/Cas systems and their applications in gene editing with filamentous fungi [J]. Synthetic Biology Journal, 2024, 5(3): 672-693. |
| [2] | XU Zhimeng, XIE Zhen. Research progress and biotechnological applications of the prime editing [J]. Synthetic Biology Journal, 2024, 5(1): 1-15. |
| [3] | CHEN Yaru, CAO Yingxiu, SONG Hao. Advances and applications of gene editing and transcriptional regulation in electroactive microorganisms [J]. Synthetic Biology Journal, 2023, 4(6): 1281-1299. |
| [4] | LIN Jicong, ZOU Gen, LIU Hongmin, WEI Yongjun. Application of CRISPR/Cas genome editing technology in the synthesis of secondary metabolites of filamentous fungi [J]. Synthetic Biology Journal, 2023, 4(4): 738-755. |
| [5] | LIU Ke, LIN Guihong, LIU Kun, ZHOU Wei, WANG Fengqing, WEI Dongzhi. Mining, engineering and functional expansion of CRISPR/Cas systems [J]. Synthetic Biology Journal, 2023, 4(1): 47-66. |
| [6] | LIANG Liya, LIU Rongming. Protein engineering of DNA targeting type Ⅱ CRISPR/Cas systems [J]. Synthetic Biology Journal, 2023, 4(1): 86-101. |
| [7] | LIU Jiaxin, CHENG Chi, LI Xinqi, WANG Chaojun, ZHANG Ying, XUE Chuang. Recent progress in the molecular genetic modification tools of Clostridium [J]. Synthetic Biology Journal, 2022, 3(6): 1201-1217. |
| [8] | BI Jiacheng, TIAN Zhigang. Synthetic immunology and future NK cell immunotherapy [J]. Synthetic Biology Journal, 2022, 3(1): 22-34. |
| [9] | XIAO Han, LIU Yixin. Progress and challenge of the CRISPR-Cas system in gene editing for filamentous fungi [J]. Synthetic Biology Journal, 2021, 2(2): 274-286. |
| [10] | LI Yang, SHEN Xiaolin, SUN Xinxiao, YUAN Qipeng, YAN Yajun, WANG Jia. Advances of CRISPR gene editing in microbial synthetic biology [J]. Synthetic Biology Journal, 2021, 2(1): 106-120. |
| [11] | YANG Yongfu, GENG Binan, SONG Haoyue, HE Qiaoning, HE Mingxiong, BAO Jie, BAI Fengwu, YANG Shihui. Progress and perspectives on developing Zymomonas mobilis as a chassis cell [J]. Synthetic Biology Journal, 2021, 2(1): 59-90. |
| [12] | YUAN Feiyan, YU Yang, LI Chun. Artificial enzyme designs and its application based on non-natural structural elements [J]. Synthetic Biology Journal, 2020, 1(6): 685-696. |
| [13] | CAO Zhongzheng, ZHANG Xinyi, XU Yiyuan, ZHOU Zhuo, WEI Wensheng. Genome editing technology and its applications in synthetic biology [J]. Synthetic Biology Journal, 2020, 1(4): 413-426. |
| [14] | LIN Lu, LV Xueqin, LIU Yanfeng, DU Guocheng, CHEN Jian, LIU Long. Advances in design, construction and applications of Bacillus subtilis chassis cells [J]. Synthetic Biology Journal, 2020, 1(2): 247-265. |
| [15] | ZHANG Bo, MA Yongshuo, SHANG Yi, HUANG Sanwen. Recent advances in plant synthetic biology [J]. Synthetic Biology Journal, 2020, 1(2): 121-140. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||