Synthetic Biology Journal ›› 2021, Vol. 2 ›› Issue (3): 384-398.DOI: 10.12211/2096-8280.2020-085
• Invited Review • Previous Articles Next Articles
The pivotal biochemical methods in DNA data storage
GAO Yanmin, TANG Mengtong, LIU Qian, QIAO Hongyan, WANG Taoxue, QI Hao
- Key Laboratory of Systems Bioengineering (Ministry of Education),School of Chemical Engineering and Technology,Tianjin University,Tianjin 300350,China
-
Received:2020-11-30Revised:2021-02-08Online:2021-07-13Published:2021-06-30 -
Contact:QI Hao
DNA信息存储中关键生化方法的研究
郜艳敏, 唐梦童, 刘倩, 乔宏艳, 王桃雪, 齐浩
- 天津大学化工学院,系统生物工程教育部重点实验室,天津 300350
-
通讯作者:齐浩 -
作者简介:郜艳敏 (1990—),女,博士研究生。研究方向为DNA信息存储和核酸检测。E-mail:xiaomingao@tju.edu.cn唐梦童 (1995—),女,硕士研究生。研究方向为DNA信息存储。E-mail:tangmengtong126@126.com齐浩 (1978—),男,教授,博士生导师。研究方向为合成生物学和分子生物学。E-mail:haoq@tju.edu.cn -
基金资助:国家重点研发计划“变革性技术关键科学问题”重点专项(2020YFA0712104);国家自然科学基金(21778039)
CLC Number:
Cite this article
GAO Yanmin, TANG Mengtong, LIU Qian, QIAO Hongyan, WANG Taoxue, QI Hao. The pivotal biochemical methods in DNA data storage[J]. Synthetic Biology Journal, 2021, 2(3): 384-398.
郜艳敏, 唐梦童, 刘倩, 乔宏艳, 王桃雪, 齐浩. DNA信息存储中关键生化方法的研究[J]. 合成生物学, 2021, 2(3): 384-398.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2020-085
| 1 | EXTANCE A. How DNA could store all the world's data[J]. Nature, 2016, 537(7618): 22-24. |
| 2 | CARMEAN D, CEZE L, SEELIG G, et al. DNA data storage and hybrid molecular-electronic computing[J]. Proceedings of the IEEE, 2019, 107(1): 63-72. |
| 3 | DE SILVA P Y, GANEGODA G U. New trends of digital data storage in DNA[J]. BioMed Research International, 2016, 2016: 8072463. |
| 4 | AKRAM F, HAQ I U, ALI H, et al. Trends to store digital data in DNA: an overview[J]. Molecular Biology Reports, 2018, 45(5): 1479-1490. |
| 5 | WATERS D L E, SHAPTER F M.The polymerase chain reaction (PCR): general methods[J]. Methods in Molecular Biology, 2014, 1099: 65-75. |
| 6 | TERRANCE W G, FRAISER M S, SCHRAM J L, et al. Strand displacement amplification-an isothermal, in vitro DNA amplification technique[J]. Nucleic Acids Research, 1992, 20(7): 1691-1696. |
| 7 | MAYBORODA O, KATAKIS I, O'SULLIVAN C K. Multiplexed isothermal nucleic acid amplification[J]. Analytical Biochemistry, 2018, 545: 20-30. |
| 8 | KOSURI S, CHURCH G M. Large-scale de novo DNA synthesis: technologies and applications[J]. Nature Methods, 2014, 11(5): 499-507. |
| 9 | GOODWIN S, MCPHERSON J D, Richard MCCOMBIE W. Coming of age: ten years of next-generation sequencing technologies[J]. Nature Reviews Genetics, 2016, 17(6): 333-351. |
| 10 | WIENER N. Interview: machines smarter than men? [J]. US News World Report, 1964, 56: 84-86. |
| 11 | NEIMAN M S. On the molecular memory systems and the directed mutations[J]. Radiotekhnika, 1965, 6: 1-8. |
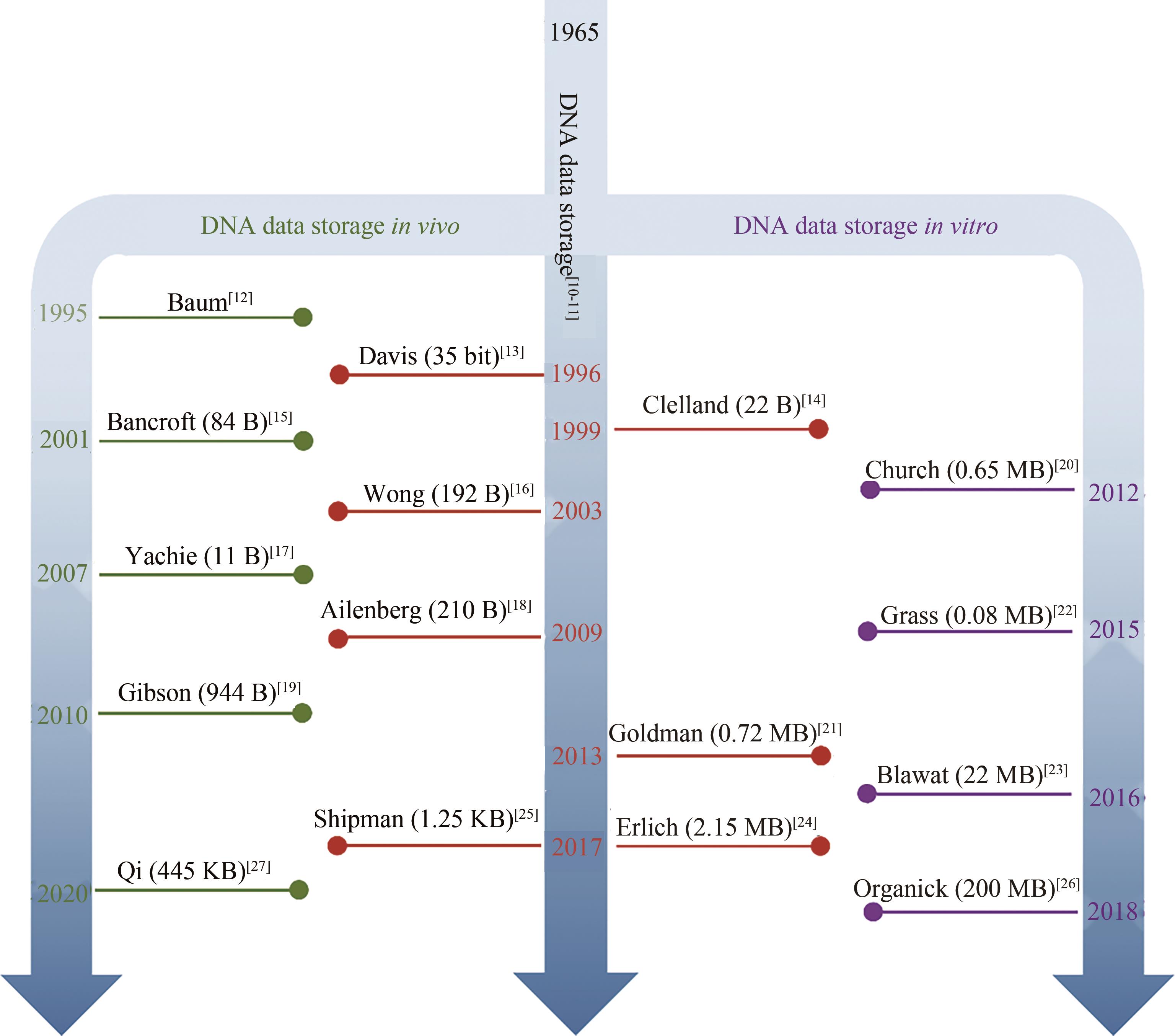
| 12 | BAUM E B. Building an associative memory vastly larger than the brain[J]. Science, 1995, 268(5210): 583-585. |
| 13 | DAVIS J. Microvenus[J]. Art Journal, 1996, 55(1): 70-74. |
| 14 | CLELLAND C T, RISCA V, BANCROFT C. Hiding messages in DNA microdots[J]. Nature, 1999, 399(6736): 533-534. |
| 15 | BANCROFT C, BOWLER T, BLOOM B, et al. Long-term storage of information in DNA[J]. Science, 2001, 293(5536): 1763. |
| 16 | WONG P C, WONG K K, FOOTE H. Organic data memory using the DNA approach[J]. Communications of the ACM, 2003, 46(1): 95-98. |
| 17 | YACHIE N, SEKIYAMA K, SUGAHARA J, et al. Alignment-based approach for durable data storage into living organisms[J]. Biotechnology Progress, 2007, 23(2): 501-505. |
| 18 | AILENBERG M, ROTSTEIN O. An improved Huffman coding method for archiving text, images, and music characters in DNA[J]. Biotechniques, 2009, 47(3): 747-754. |
| 19 | GIBSON D G, GLASS J I, LARTIGUE C, et al. Creation of a bacterial cell controlled by a chemically synthesized genome[J]. Science, 2010, 329(5987): 52-56. |
| 20 | CHURCH G M, GAO Y, KOSURI S. Next-generation digital information storage in DNA[J]. Science, 2012, 337(6102): 1628. |
| 21 | GOLDMAN N, BERTONE P, CHEN S, et al. Towards practical, high-capacity, low-maintenance information storage in synthesized DNA[J]. Nature, 2013, 494(7435): 77-80. |
| 22 | GRASS R N, HECKEL R, PUDDU M, et al. Robust chemical preservation of digital information on DNA in silica with error-correcting codes[J]. Angewandte Chemie International Edition, 2015, 54(8): 2552-2555. |
| 23 | BLAWAT M, GAEDKE K, HVTTER I, et al. Forward error correction for DNA data storage[J]. Procedia Computer Science, 2016, 80: 1011-1022. |
| 24 | ERLICH Y, ZIELINSKI D. DNA fountain enables a robust and efficient storage architecture[J]. Science, 2017, 355(6328): 950-954. |
| 25 | SHIPMAN S L, NIVALA J, MACKLIS J D, et al. CRISPR-Cas encoding of a digital movie into the genomes of a population of living bacteria[J]. Nature, 2017, 547(7663): 345-349. |
| 26 | ORGANICK L, ANG S D, CHEN Y J, et al. Random access in large-scale DNA data storage[J]. Nature Biotechnology, 2018, 36(3): 242-248. |
| 27 | HAO Min, QIAO Hongyan, GAO Yanmin, et al. A mixed culture of bacterial cells enables an economic DNA storage on a large scale[J]. Communications Biology, 2020, 3(1): 416. |
| 28 | BORNHOLT J, LOPEZ R, CARMEAN D M, et al. Toward a DNA-based archival storage system[J]. IEEE Micro, 2017, 37(3): 98-104. |
| 29 | CEZE L, NIVALA J, STRAUSS K. Molecular digital data storage using DNA[J]. Nature Review Genetics, 2019, 20(8): 456-466. |
| 30 | TAKAHASHI C N, NGUYEN B H, STRAUSS K, et al. Demonstration of end-to-end automation of DNA data storage[J]. Scientific Reports, 2019, 9(1): 4998. |
| 31 | CHEN Y J, TAKAHASHI C N, ORGANICK L, et al. Quantifying molecular bias in DNA data storage[J]. Nature Communications, 2020, 11(1): 3264. |
| 32 | HECKEL R, MIKUTIS G, GRASS R N. A Characterization of the DNA data storage channel[J]. Scientific Reports, 2019, 9(1): 9663. |
| 33 | BHAN N J, STRUTZ J, GLASER J, et al. Recording temporal data onto DNA with minutes resolution[J], 2019. doi:10.1101/634790 |
| 34 | HENDLING M, BARISIC I. In-silico design of DNA oligonucleotides: challenges and approaches[J]. Computational and Structural Biotechnology Journal, 2019, 17: 1056-1065. |
| 35 | TIAN Jingdong, MA K, SAAEM I. Advancing high-throughput gene synthesis technology[J]. Molecular Biosystems, 2009, 5(7): 714-722. |
| 36 | 王霞, 赵鹃, 李炳志, 等. DNA合成技术及应用[J]. 生命科学, 2013, 25(10): 993-999. |
| WANG Xia, ZHAO Juan, LI Bingzhi, et al. DNA synthesis methods and applications [J]. Chinese Bulletin of Life Sciences, 2013, 25(10):993-999. | |
| 37 | CLORE A. A new route to synthetic DNA[J]. Nature Biotechnology, 2018, 36(7): 593-595. |
| 38 | LEE H H, KALHOR R, GOELA N, et al. Terminator-free template-independent enzymatic DNA synthesis for digital information storage[J]. Nature Communications, 2019, 10(1): 2383. |
| 39 | MURGHA Y E, ROUILLARD J M, GULARI E.Methods for the preparation of large quantities of complex single-stranded oligonucleotide libraries[J]. PLoS One, 2014, 9(4): e94752. |
| 40 | TIAN Jingdong, GONG Hui, SHENG Nijing, et al. Accurate multiplex gene synthesis from programmable DNA microchips[J]. Nature, 2004, 432(7020): 1050-1054. |
| 41 | KLEIN J C, LAJOIE M J, SCHWARTZ J J, et al. Multiplex pairwise assembly of array-derived DNA oligonucleotides[J]. Nucleic Acids Research, 2016, 44(5): e43. |
| 42 | ANTKOWIAK P L, LIETARD J, DARESTANI M Z, et al.Low cost DNA data storage using photolithographic synthesis and advanced information reconstruction and error correction[J]. Nature Communications, 2020, 11(1): 5345. |
| 43 | TABATABAEI S K, WANG B, ATHREYA N B M, et al. DNA punch cards for storing data on native DNA sequences via enzymatic nicking[J]. Nature Communications, 2020, 11(1): 1742. |
| 44 | ELLINGTON A, POLLARD J D. Synthesis and purification of oligonucleotides[J]. Current Protocols Molecular Biology, 2001, 42(1). DOI:10.1002/0471142727.mb0211s42 . |
| 45 | PINTO A, CHEN S X, ZHANG D Y. Simultaneous and stoichiometric purification of hundreds of oligonucleotides[J]. Nature Communications, 2018, 9(1): 2467. |
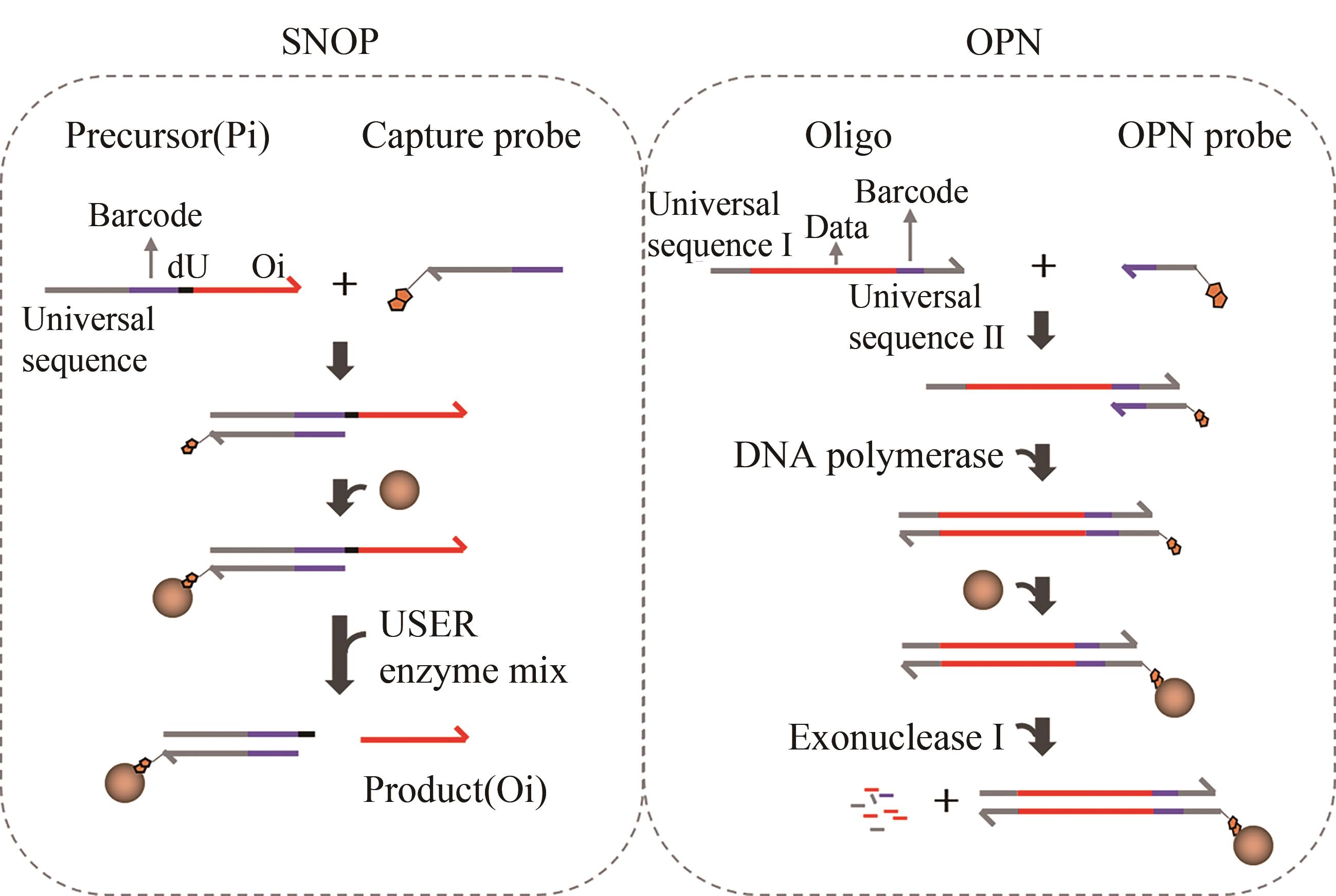
| 46 | GAO Yanmin, CHEN Xin, QIAO Hongyan, et al. Low-bias manipulation of DNA oligo pool for robust data storage[J]. ACS Synthetic Biology, 2020, 9(12): 3344-3352 |
| 47 | WILLERSLEV E, COOPER A. Ancient DNA[J]. Proceedings of the Royal Society B: Biological Sciences, 2005, 272(1558): 3-16. |
| 48 | MITCHELL D, WILLERSLEV E, HANSEN A.Damage and repair of ancient DNA[J]. Mutation Research, 2005, 571(1-2): 265-276. |
| 49 | BONNET J, COLOTTE M, COUDY D, et al. Chain and conformation stability of solid-state DNA: implications for room temperature storage[J]. Nucleic Acids Research, 2010, 38(5): 1531-1546. |
| 50 | MIKUTIS G, SCHMID L, STARK W J, et al. Length-dependent DNA degradation kinetic model: decay compensation in DNA tracer concentration measurements[J]. AIChE Journal, 2019, 65(1): 40-48. |
| 51 | KNEBELSBERGER T, STOGER I.DNA extraction, preservation, and amplification[J]. Methods in Molecular Biology, 2012, 858: 311-338. |
| 52 | WAN E, AKANA M, PONS J, et al. Green technologies for room temperature nucleic acid storage[J]. Current Issues Molecular Biology, 2010, 12(3): 135-142. |
| 53 | BURGOYNE L A. Solid medium and method for DNA storage: US5985327[P].1996-03-05. |
| 54 | IVANOVA N V, KUZMINA M L. Protocols for dry DNA storage and shipment at room temperature[J]. Molecular Ecology Resources, 2013, 13(5): 890-898. |
| 55 | ALKHAMIS K A. Influence of solid-state acidity on the decomposition of sucrose in amorphous systems [J]. International Journal of Pharmaceutics, 2008, 362(1/2): 74-80. |
| 56 | KARNI M, ZIDON D, POLAK P, et al.Thermal degradation of DNA[J]. DNA and Cell Biology, 2013, 32(6): 298-301. |
| 57 | ZHU B, FURUKI T, OKUDA T, et al. Natural DNA mixed with trehalose persists in B-form double-stranding even in the dry state[J]. Journal of Physical Chemistry B, 2007, 111(20): 5542-5544. |
| 58 | ANCHORDOQUY T J, MOLINA M C.Preservation of DNA[J]. Cell Preservation Technology, 2007, 5(4): 180-188. |
| 59 | NEWMAN S, STEPHENSON A P, WILLSEY M, et al. High density DNA data storage library via dehydration with digital microfluidic retrieval[J]. Nature Communications, 2019, 10(1): 1706. |
| 60 | HOWLETT S E, CASTILLO H S, GIOENI L J, et al. Evaluation of DNA stable for DNA storage at ambient temperature[J]. Forensic Science International Genetics, 2014, 8(1): 170-178. |
| 61 | ORGANICK L, NGUYEN B H, MCAMIS R, et al. An empirical comparison of preservation methods for synthetic DNA data storage [J]. Small Methods, 2021. DOI:10.1002/smtd 202001094 . |
| 62 | CLERMONT D, SANTONI S, SAKER S, et al. Assessment of DNA encapsulation, a new room-temperature DNA storage method[J]. Biopreservation and Biobanking, 2014, 12(3): 176-183. |
| 63 | PAUNESCU D, FUHRER R, GRASS R N. Protection and deprotection of DNA-high-temperature stability of nucleic acid barcodes for polymer labeling[J]. Angewandte Chemie International Edition, 2013, 52(15): 4269-4272. |
| 64 | KOCH J, GANTENBEIN S, MASANIA K, et al. A DNA-of-things storage architecture to create materials with embedded memory[J]. Nature Biotechnology, 2020, 38(1): 39-43. |
| 65 | CHEN W D, KOHLL A X, NGUYEN B H, et al. Combining data longevity with high storage capacity—layer-by-layer DNA encapsulated in magnetic nanoparticles[J]. Advanced Functional Materials, 2019. DOI:10.1002/adfm.201901672 . |
| 66 | VALLE L J DEL, BERTRAN O, CHAVES G, et al. DNA adsorbed on hydroxyapatite surfaces[J]. Journal of Materials Chemistry B, 2014, 2(40): 6953-6966. |
| 67 | KOHLL A X, ANTKOWIAK P L, CHEN W D, et al. Stabilizing synthetic DNA for long-term data storage with earth alkaline salts[J]. Chemical Communications, 2020, 56(25): 3613-3616. |
| 68 | SCHONING K, SCHOLZ P, GUNTHA S, et al. Chemical etiology of nucleic acid structure: the alpha-threofuranosyl-(3'->2') oligonucleotide system[J]. Science, 2000, 290(5495): 1347-1351. |
| 69 | YANG K, MCCLOSKEY C M, CHAPUT J C. Reading and writing digital information in TNA[J]. ACS Synthetic Biology, 2020, 9(11): 2936-2942. |
| 70 | DAGHER G G, MACHADO A P, DAVIS E C, et al. Data storage in cellular DNA: contextualizing diverse encoding schemes[J]. Evolutionary Intelligence, 2019. DOI: 10.1007/s12065-019-00202-z . |
| 71 | AKHMETOV A, ELLINGTON A D, MARCOTTE E M. A highly parallel strategy for storage of digital information in living cells[J]. BMC Biotechnology, 2018, 18(1): 64. |
| 72 | NGUYEN H, PARK J, PARK S, et al. Long-term stability and integrity of plasmid-based DNA data storage[J]. Polymers, 2018, 10(1): 28. |
| 73 | SHIPMAN S L, NIVALA J, MACKLIS J D, et al. Molecular recordings by directed CRISPR spacer acquisition[J]. Science, 2016, 353(6298): aaf1175. |
| 74 | DABNEY J, MEYER M. Length and GC-biases during sequencing library amplification: a comparison of various polymerase-buffer systems with ancient and modern DNA sequencing libraries[J]. BioTechniques, 2012, 52(2): 87-94. |
| 75 | ROUX K H. Optimization and troubleshooting in PCR[J]. Cold Spring Harbor Protocols, 2009, 2009(4): pdb, ip66. |
| 76 | AIRD D, ROSS M G, CHEN W S, et al. Analyzing and minimizing PCR amplification bias in Illumina sequencing libraries[J]. Genome Biology, 2011, 12(2): R18. |
| 77 | BENJAMINI Y, Speed T P. Summarizing and correcting the GC content bias in high-throughput sequencing[J]. Nucleic Acids Research, 2012, 40(10): e72. |
| 78 | CLINE J, BRAMAN J C, HOGREFE H H. PCR fidelity of pfu DNA polymerase and other thermostable DNA polymerases[J]. Nucleic Acids Research, 1996, 24(18): 3546-3551. |
| 79 | CZERNY T. High primer concentration improves PCR amplification from random pools[J]. Nucleic Acids Research, 1996, 24(5): 985-986. |
| 80 | WANG Y, NOOR-A-RAHIM M, ZHANG J, et al. Oligo design with single primer binding site for high capacity DNA-based data storage[J]. IEEE/ACM Transactions Computational Biology and Bioinformatics 2020, 17 (6): 2176-2182. |
| 81 | WINSHIP P R. An improved method for directly sequencing PCR-amplified material using dimethyl sulphoxide[J]. Nucleic Acids Research, 1989, 17(3): 1266. |
| 82 | VARADARAJ K, SKINNER D M. Denaturants or cosolvents improve the specificity of PCR amplification of a G+C-rich DNA using genetically engineered DNA polymerases[J]. Gene, 1994, 140(1): 1-5. |
| 83 | HENKE W, HERDEL K, JUNG K, et al. Betaine improves the PCR amplification of GC-rich DNA sequences[J]. Nucleic Acids Research, 1997, 25(19): 3957-3958. |
| 84 | 王同同,张利平,尹康权. 甜菜碱改善水稻高GC含量DNA序列的PCR扩增[J]. 生物技术通报, 2018, 34(3): 80-86. |
| WANG Tongtong, ZHANG Liping, YIN Kangquan. Betaine improves the PCR amplification of rice GC-rich DNA sequence [J]. Biotechnology Bulletin, 2018,34(3):80-86. | |
| 85 | FARELL E M, ALEXANDRE G. Bovine serum albumin further enhances the effects of organic solvents on increased yield of polymerase chain reaction of GC-rich templates[J]. BMC Research Notes, 2012, 5: 257. |
| 86 | MCCONLOGUE L, BROW M A D., INNIS Michael A. Structure-independent DNA amplification by PCR using 7-deaza-2'-deoxyguanosine[J]. Nucleic Acids Research, 1988, 16(20): 9869. |
| 87 | PAN Wenjing, BYRNE-STEELE M, WANG Chunlin, et al. DNA polymerase preference determines PCR priming efficiency[J]. BMC Biotechnology, 2014, 14: 10. |
| 88 | DON R H, COX P T, WAINWRIGHT B J, et al. 'Touchdown' PCR to circumvent spurious priming during gene amplification[J]. Nucleic Acids Research, 1991, 19(14): 4008. |
| 89 | HECKER K H, ROUX K H. High and low annealing temperatures increase both specificity and yield in touchdown and stepdown PCR[J]. Biotechniques, 1996, 20(3): 478-485. |
| 90 | FREY U H, BACHMANN H S, PETERS J, et al. PCR-amplification of GC-rich regions: 'slowdown PCR'[J]. Natural Protocols, 2008, 3(8): 1312-1317. |
| 91 | KANAGAL-SHAMANNA R. EMULSION PCR: Techniques and applications[J]. Methods in Molecular Biology, 2016, 1392: 33-42. |
| 92 | VERMA V, GUPTA A, CHAUDHARY V K. Emulsion PCR made easy[J]. BioTechniques, 2020, 69(1): 421-426. |
| 93 | SHAO K, DING Weifeng, WANG Feng, et al. Emulsion PCR: a high efficient way of PCR amplification of random DNA libraries in aptamer selection[J]. PLoS One, 2011, 6(9): e24910. |
| 94 | POLZ M F, CAVANAUGH C M. Bias in template-to-product ratios in multitemplate PCR[J]. Applied and Environmental Microbiology, 1998, 64(10): 3724-3730. |
| 95 | QIAN Cheng, WANG Rui, WU Hui, et al. Nicking enzyme-assisted amplification (NEAA) technology and its applications: a review[J]. Analytica Chimica Acta, 2019, 1050: 1-15. |
| 96 | ZHAO Yongxi, CHEN Feng, LI Qian, et al. Isothermal amplification of nucleic acids[J]. Chemical Review, 2015, 115(22): 12491-12545. |
| 97 | CHOI Y, BAE H J, LEE A C, et al. DNA micro-disks for the management of DNA-based data storage with index and write-once-read-many (WORM) memory features[J]. Advanced Materials, 2020, 32(37): e2001249. |
| 98 | WANG Ting, LIU Yong, SUN Huanhuan, et al. An RNA-guided Cas9 nickase-based method for universal isothermal DNA amplification[J]. Angewandte ChemieInternational Edition, 2019, 58(16): 5382-5386. |
| 99 | ZHOU Wenhua, HU Li, YING Liming, et al. A CRISPR-Cas9-triggered strand displacement amplification method for ultrasensitive DNA detection[J]. Nature Communications, 2018, 9(1): 5012. |
| 100 | WILLSEY M, STRAUSS K, CEZE L, et al. Puddle: a dynamic, error-correcting, full-stack microfluidics platform[C]∥ASPLOS' 19 . New York: ACM, 2019: 183-197. |
| 101 | PRESS W H, HAWKINS J A, JONES S K, et al. HEDGES error-correcting code for DNA storage corrects indels and allows sequence constraints[J]. Proceedings of the National Academy of Sciences of the United States of America, 2020, 117(31): 18489-18496. |
| 102 | SONG Wentu, CAI Kui, ZHANG Mu, et al. Codes with run-length and GC-content constraints for DNA-based data storage[J]. IEEE Communications Letters, 2018, 22(10): 2004-2007. |
| [1] | JIAO Hongtao, QI Meng, SHAO Bin, JIANG Jinsong. Legal issues for the storage of DNA data [J]. Synthetic Biology Journal, 2025, 6(1): 177-189. |
| [2] | HUANG Xiaoluo, DAI Junbiao. DNA synthesis technology: foundation of DNA data storage [J]. Synthetic Biology Journal, 2021, 2(3): 335-353. |
| [3] | CHEN Weigang, GE Qi, WANG Panpan, HAN Mingzhe, GUO Jian. Multiple interleaved RS codes for data storage using up to Mb-scale synthetic DNA in living cells [J]. Synthetic Biology Journal, 2021, 2(3): 428-443. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||