Synthetic Biology Journal ›› 2022, Vol. 3 ›› Issue (6): 1061-1080.DOI: 10.12211/2096-8280.2021-021
• Invited Review • Previous Articles Next Articles
Design, optimization and application of whole-cell microbial biosensors with engineered genetic circuits
YANG Lu1, WU Nan1, BAI Rongrong1, DONG Weiliang1,2, ZHOU Jie1,2, JIANG Min1,2
- 1.College of Biotechnology and Pharmaceutical Engineering,Nanjing Tech University,Nanjing 211816,Jiangsu,China
2.State Key Laboratory of Materials-Oriented Chemical Engineering,Nanjing Tech University,Nanjing 211816,Jiangsu,China
-
Received:2021-02-07Revised:2021-04-23Online:2023-01-17Published:2022-12-31 -
Contact:ZHOU Jie, JIANG Min
基因回路型全细胞微生物传感器的设计、优化与应用
杨璐1, 吴楠1, 白茸茸1, 董维亮1,2, 周杰1,2, 姜岷1,2
- 1.南京工业大学生物与制药工程学院,江苏 南京 211816
2.南京工业大学材料化学工程国家重点实验室,江苏 南京 211816
-
通讯作者:周杰,姜岷 -
作者简介:杨璐 (1997—),女,硕士研究生。研究方向为合成生物传感器的设计与应用。E-mail:luyang97@njtech.edu.cn周杰 (1991—),男,博士,副教授。研究方向为合成生物传感器的设计与应用。E-mail:jayzhou@njtech.edu.cn姜岷 (1972—),男,博士,教授。研究方向为人工多细胞体系设计与构建。E-mail:bioengine@njtech.edu.cn -
基金资助:国家重点研发计划“合成生物学”重点专项(2018YFA0902200);国家自然科学基金(21727818)
CLC Number:
Cite this article
YANG Lu, WU Nan, BAI Rongrong, DONG Weiliang, ZHOU Jie, JIANG Min. Design, optimization and application of whole-cell microbial biosensors with engineered genetic circuits[J]. Synthetic Biology Journal, 2022, 3(6): 1061-1080.
杨璐, 吴楠, 白茸茸, 董维亮, 周杰, 姜岷. 基因回路型全细胞微生物传感器的设计、优化与应用[J]. 合成生物学, 2022, 3(6): 1061-1080.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2021-021
| 评价指标 | 描述 |
|---|---|
| 特异性 | 在复杂环境中专一性识别待测底物的能力 |
| 灵敏度 | 对待测物浓度变化的敏感程度 |
| 动态范围 | 最大输出信号和最小输出信号的比例 |
| 操作范围 | 输入与输出呈单一正相关或负相关的输入信号浓度范围 |
| 稳定性 | 样品检测过程中保持性能恒定的能力 |
| 响应时间 | 信号输入到信号输出的总时间 |
| 安全性 | 是否存在基因泄露转移、吸收外源DNA或潜在有害突变等风险 |
Tab. 1 Evaluation of whole cell microbial sensor [44]
| 评价指标 | 描述 |
|---|---|
| 特异性 | 在复杂环境中专一性识别待测底物的能力 |
| 灵敏度 | 对待测物浓度变化的敏感程度 |
| 动态范围 | 最大输出信号和最小输出信号的比例 |
| 操作范围 | 输入与输出呈单一正相关或负相关的输入信号浓度范围 |
| 稳定性 | 样品检测过程中保持性能恒定的能力 |
| 响应时间 | 信号输入到信号输出的总时间 |
| 安全性 | 是否存在基因泄露转移、吸收外源DNA或潜在有害突变等风险 |
| 领域 | 检测物 | 传感元件 | 操作范围 | 应用 | 参考文献 |
|---|---|---|---|---|---|
| 生物制造过程监控 | 木糖 | XylR | 0~20 g/L | 木糖转运蛋白筛选 | [ |
| 甘氨酸 | 甘氨酸核糖开关 | 20~200 μmol/L | 动态调控5-氨基乙酰丙酸合成 | [ | |
| 丙酮酸 | PdhR | 10~35 nmol/g | 动态调控葡萄糖二酸合成 | [ | |
| 衣康酸 | YpLysR | 16~160 μmol/L | 衣康酸高产菌株筛选 | [ | |
| 环境监测食品安全 | 4-甲基苯二酚 | ChpR | 25~500 nmol/L | 农药毒死蜱检测 | [ |
| As3+ /As5+ | ArsR | 0.74~60 μg/L | 水样重金属离子检测 | [ | |
四环素 土霉素 | TetR | >20 μg/kg >50 μg/kg | 鱼肉四环素、土霉素含量检测 | [ | |
| 医疗诊断与监护 | 硫代硫酸盐 | ThsR | 0.1~1 mmol/L | 小鼠结肠炎诊断 | [ |
| 茶碱 | 茶碱核糖开关 | 0~5 mmol/L | 支气管扩张剂药物摄取监测 | [ | |
| 尿酸 | HucR | 0~5 mmol/L | 血液中尿酸稳态维持 | [ | |
| 葡萄糖 | GBP | 1~10 mmol/L | 糖尿病患者血糖监测 | [ |
Tab. 2 Application examples of whole-cell microbial biosensor
| 领域 | 检测物 | 传感元件 | 操作范围 | 应用 | 参考文献 |
|---|---|---|---|---|---|
| 生物制造过程监控 | 木糖 | XylR | 0~20 g/L | 木糖转运蛋白筛选 | [ |
| 甘氨酸 | 甘氨酸核糖开关 | 20~200 μmol/L | 动态调控5-氨基乙酰丙酸合成 | [ | |
| 丙酮酸 | PdhR | 10~35 nmol/g | 动态调控葡萄糖二酸合成 | [ | |
| 衣康酸 | YpLysR | 16~160 μmol/L | 衣康酸高产菌株筛选 | [ | |
| 环境监测食品安全 | 4-甲基苯二酚 | ChpR | 25~500 nmol/L | 农药毒死蜱检测 | [ |
| As3+ /As5+ | ArsR | 0.74~60 μg/L | 水样重金属离子检测 | [ | |
四环素 土霉素 | TetR | >20 μg/kg >50 μg/kg | 鱼肉四环素、土霉素含量检测 | [ | |
| 医疗诊断与监护 | 硫代硫酸盐 | ThsR | 0.1~1 mmol/L | 小鼠结肠炎诊断 | [ |
| 茶碱 | 茶碱核糖开关 | 0~5 mmol/L | 支气管扩张剂药物摄取监测 | [ | |
| 尿酸 | HucR | 0~5 mmol/L | 血液中尿酸稳态维持 | [ | |
| 葡萄糖 | GBP | 1~10 mmol/L | 糖尿病患者血糖监测 | [ |
| 1 | 周益康, 吴亦楠, 王天民, 等. 代谢物生物传感器:微生物细胞工厂构建中的合成生物学工具[J]. 生物技术通报, 2017, 33(1): 1-11. |
| ZHOU Y K, WU Y N, WANG T M, et al. Metabolite biosensor: a useful synthetic biology tool to assist the construction of microbial cell factory[J]. Biotechnology Bulletin, 2017, 33(1): 1-11. | |
| 2 | 秦伟彤, 田健, 伍宁丰. 全细胞生物传感器的设计及其在环境监测中的应用[J]. 生物技术进展, 2018, 8(5): 369-375. |
| QIN W T, TIAN J, WU N F. Design of the whole-cell biosenor and its application in environmental monitoring[J]. Current Biotechnology, 2018, 8(5): 369-375. | |
| 3 | 张静, 吕雪飞,邓玉林. 基因工程微生物传感器及其应用研究进展[J]. 生命科学仪器, 2019, 17(1): 11-16. |
| ZHANG J, LV X F, DENG Y L. Application research of genetically engineered microbial biosensors[J]. Life Science Instruments, 2019, 17(1): 11-16. | |
| 4 | 施冬艳, 何珣, 陈怡露. 合成生物学在微生物传感器中的应用[J]. 东南大学学报, 2012, 31(3): 363-369. |
| SHI D Y, HE X, CHEN Y L. Application of synthetic biology in microbial sensors[J]. Journal of Southeast University, 2012, 31(3): 363-369. | |
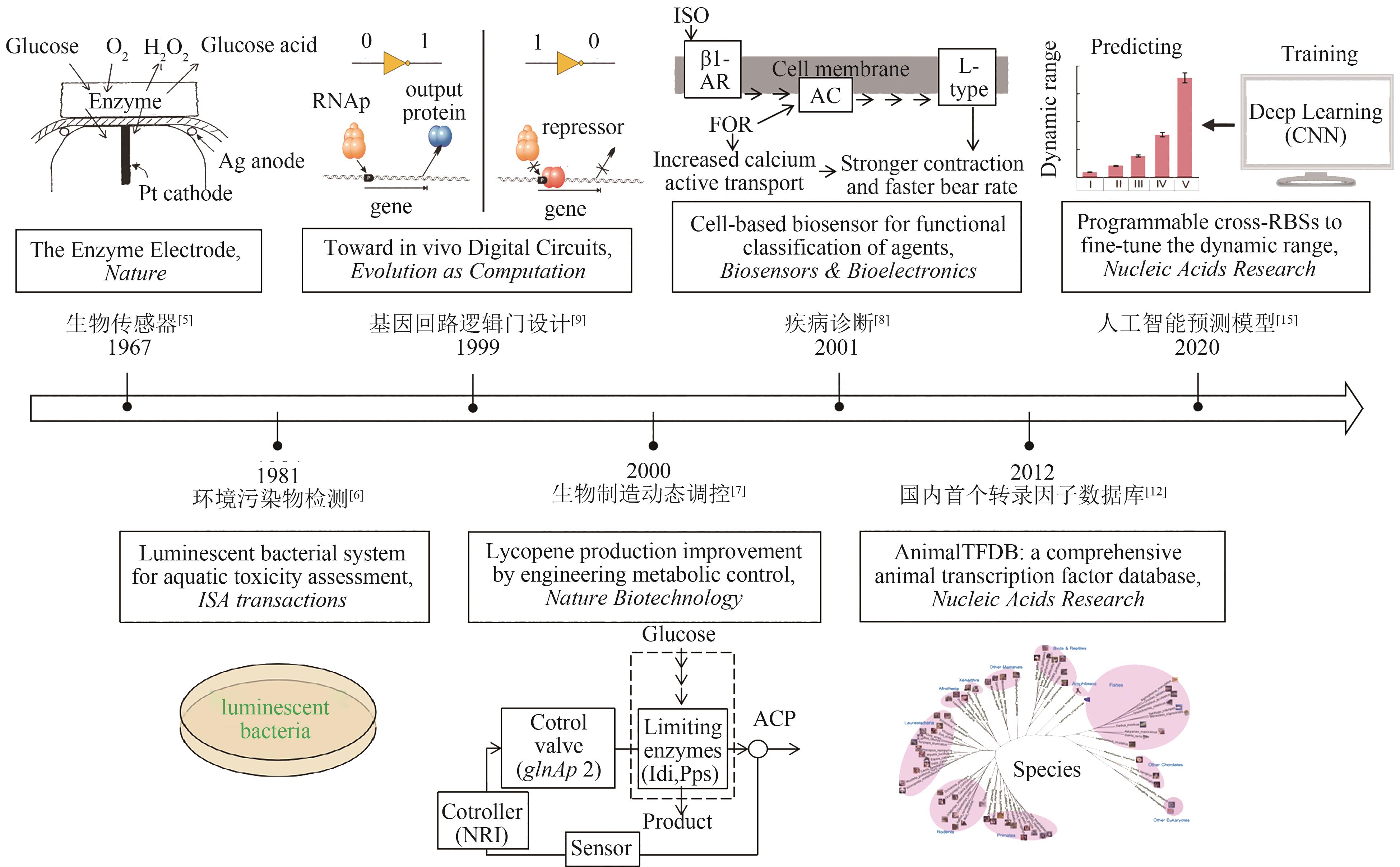
| 5 | UPDIKE S J, HICKS G P. The enzyme electrode[J]. Nature, 1967, 214(5092): 986-988. |
| 6 | BULICH A A, ISENBERG D L. Use of the luminescent bacterial system for the rapid assessment of aquatic toxicity[J]. ISA transactions, 1981, 20(1): 29-33. |
| 7 | FARMER W R, LIAO J C. Improving lycopene production in Escherichia coli by engineering metabolic control[J]. Nature Biotechnology, 2000, 18(5): 533-537. |
| 8 | ARAVANIS A M, DE BUSSCHERE B D, CHRUSCINSKI A J, et al. A genetically engineered cell-based biosensor for functional classification of agents[J]. Biosensors and Bioelectronics, 2001, 16(7/8): 571-577. |
| 9 | WEISS R, HOMSY G E, KNIGHT T F. Toward in vivo digital circuits[C]//LANDWEBER L F, WINFREE E. Evolution as Computation. Berlin: Springer, 2002:275-295. |
| 10 | SALTEPE B, KEHRIBAR E S, YIRMIBESOGLU S S S, et al. Cellular biosensors with engineered genetic circuits[J]. ACS Sensors, 2018, 3(1): 13-26. |
| 11 | AUSLANDER S, AUSLANDER D, MULLER M, et al. Programmable single-cell mammalian biocomputers[J]. Nature, 2012, 487(7405): 123-127. |
| 12 | ZHANG H M, CHEN H, LIU W, et al. AnimalTFDB: a comprehensive animal transcription factor database[J]. Nucleic Acids Research, 2011, 40(D1): D144-D149. |
| 13 | ANG J, HARRIS E, HUSSEY B J, et al. Tuning response curves for synthetic biology[J]. ACS Synthetic Biology, 2013, 2(10): 547-567. |
| 14 | RICCI F, VALLEE-BELISLE A, SIMON A J, et al. Using nature's "tricks" to rationally tune the binding properties of biomolecular receptors[J]. Accounts of Chemical Research, 2016, 49(9): 1884-1892. |
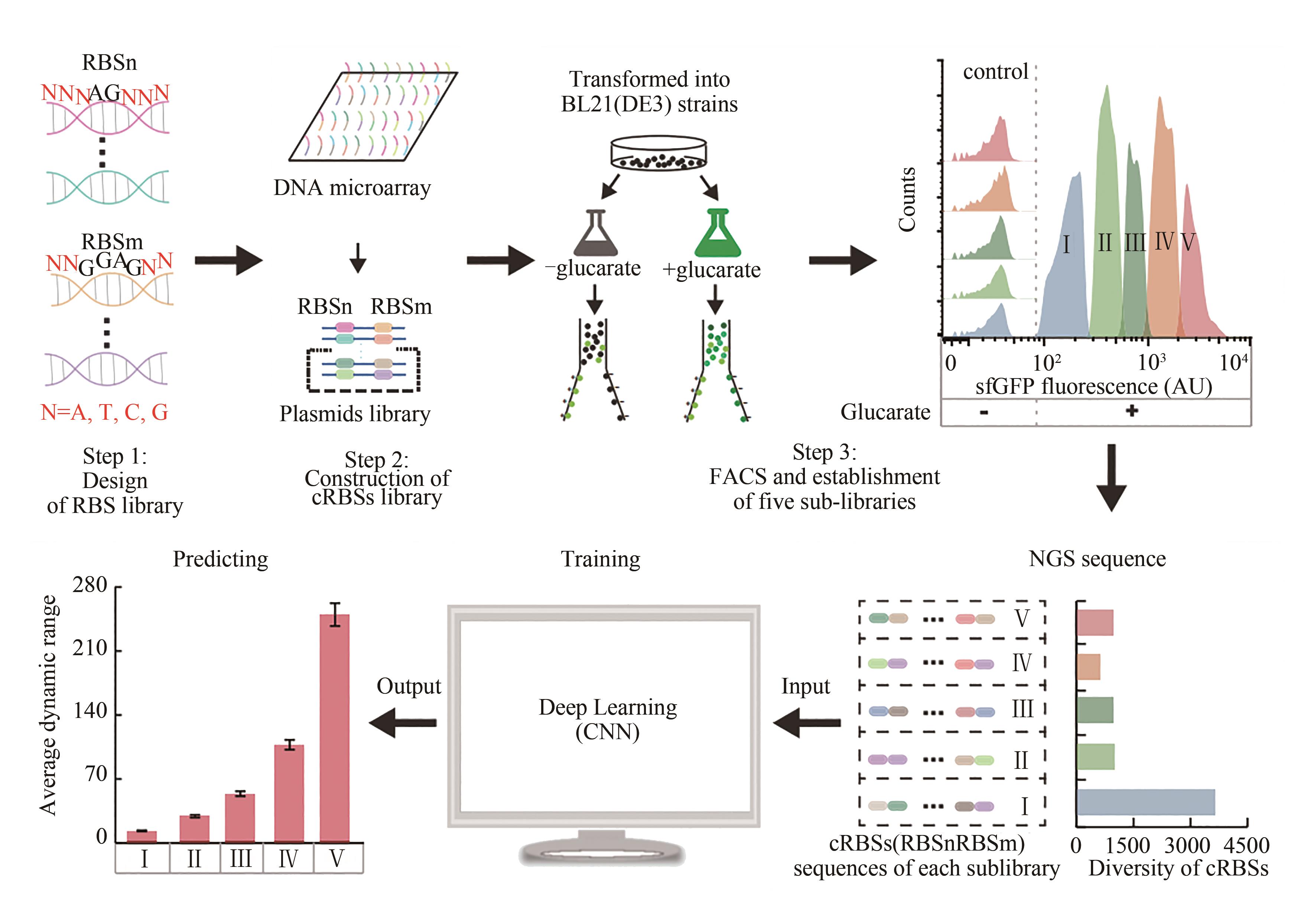
| 15 | DING N N, YUAN Z Q, ZHANG X J, et al. Programmable cross-ribosome-binding sites to fine-tune the dynamic range of transcription factor-based biosensor[J]. Nucleic Acids Research, 2020, 48(18): 10602-10613. |
| 16 | DE PAEPE B, PETERS G, COUSSEMENT P, et al. Tailor-made transcriptional biosensors for optimizing microbial cell factories[J]. Journal of Industrial Microbiology and Biotechnology, 2017, 44(4/5): 623-645. |
| 17 | XU X H, LI X L, LIU Y F, et al. Pyruvate-responsive genetic circuits for dynamic control of central metabolism[J]. Nature Chemical Biology, 2020, 16(11): 1261-1268. |
| 18 | TANG R Q, WAGNER J M, ALPER H S, et al. Design, evolution, and characterization of a xylose biosensor in Escherichia coli using the XylR/xylO system with an expanded operating range[J]. ACS Synthetic Biology, 2020, 9(10): 2714-2722. |
| 19 | TANG S Y, FAZELINIA H, CIRINO P C. AraC regulatory protein mutants with altered effector specificity[J]. Journal of the American Chemical Society, 2008, 130(15): 5267-5271. |
| 20 | XU P, LI L Y, ZHANG F M, et al. Improving fatty acids production by engineering dynamic pathway regulation and metabolic control[J]. Proceedings of the National Academy of Sciences of the United States of America, 2014, 111(31): 11299-11304. |
| 21 | YANG Y P, LIN Y H, WANG J, et al. Sensor-regulator and RNAi based bifunctional dynamic control network for engineered microbial synthesis[J]. Nature Communications, 2018, 9(1): 3043-3052. |
| 22 | WU Y K, CHEN T C, LIU Y F, et al. Design of a programmable biosensor-CRISPRi genetic circuits for dynamic and autonomous dual-control of metabolic flux in Bacillus subtilis [J]. Nucleic Acids Research, 2020, 48(2): 996-1009. |
| 23 | TUNGTUR S, EGAN S M, SWINT-KRUSE L. Functional consequences of exchanging domains between LacI and PurR are mediated by the intervening linker sequence[J]. Proteins: Structure, Function, and Bioinformatics, 2007, 68(1): 375-388. |
| 24 | WINKLER W, NAHVI A, BREAKER R R. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression[J]. Nature, 2002, 419(6910): 952-956. |
| 25 | ZHOU L B, REN J, LI Z D, et al. Characterization and engineering of a clostridium glycine riboswitch and its use to control a novel metabolic pathway for 5-aminolevulinic acid production in Escherichia coli [J]. ACS Synthetic Biology, 2019, 8(10): 2327-2335. |
| 26 | ZHOU L B, ZENG A P. Exploring lysine riboswitch for metabolic flux control and improvement of L-lysine synthesis in Corynebacterium glutamicum [J]. ACS Synthetic Biology, 2015, 4(6): 729-734. |
| 27 | ONTIVEROS-PALACIOS N, SMITH A M, GRUNDY F J, et al. Molecular basis of gene regulation by the THI-box riboswitch [J]. Molecular Microbiology, 2008, 67(4): 793-803. |
| 28 | WINKLER W C, COHEN-CHALAMISH S, BREAKER R R. An mRNA structure that controls gene expression by binding FMN [J]. Proceedings of the National Academy of Sciences of the United States of America, 2002, 99(25): 15908-15913. |
| 29 | CROMIE M J, SHI Y X, LATIFI T, et al. An RNA sensor for intracellular Mg2+ [J]. Cell, 2006, 125(1): 71-84. |
| 30 | SHI Y X, ZHAO G, KONG W. Genetic analysis of riboswitch-mediated transcriptional regulation responding to Mn2+ in Salmonella [J]. Journal of Biological Chemistry, 2014, 289(16): 11353-11366. |
| 31 | SUESS B, HANSON S, BERENS C, et al. Conditional gene expression by controlling translation with tetracycline-binding aptamers[J]. Nucleic Acids Research, 2003, 31(7): 1853-1858. |
| 32 | TRAUSCH J J, CERES P, REYES F E, et al. The structure of a tetrahydrofolate-sensing riboswitch reveals two ligand binding sites in a single aptamer[J]. Structure, 2011, 19(10): 1413-1423. |
| 33 | DESAI S K, GALLIVAN J P. Genetic screens and selections for small molecules based on a synthetic riboswitch that activates protein translation[J]. Journal of the American Chemical Society, 2004, 126(41): 13247-13254. |
| 34 | BEREZA-MALCOLM L T, MANN G, FRANKS A E. Environmental sensing of heavy metals through whole cell microbial biosensors: a synthetic biology approach[J]. ACS Synthetic Biology, 2015, 4(5): 535-546. |
| 35 | TAO H C, PENG Z W, LI P S, et al. Optimizing cadmium and mercury specificity of CadR-based E. coli biosensors by redesign of CadR[J]. Biotechnology Letters, 2013, 35(8): 1253-1258. |
| 36 | HANKO E K R, MINTON N P, MALYS N. A Transcription factor-based biosensor for detection of itaconic acid[J]. ACS Synthetic Biology, 2018, 7(5): 1436-1446. |
| 37 | YAGI K. Applications of whole-cell bacterial sensors in biotechnology and environmental science[J]. Applied Microbiology & Biotechnology, 2007, 73(6): 1251-1258. |
| 38 | FUJIMOTO H, WAKABAYASHI M, YAMASHIRO H, et al. Whole-cell arsenite biosensor using photosynthetic bacterium Rhodovulum sulfidophilum [J]. Applied Microbiology and Biotechnology, 2006, 73(2): 332-338. |
| 39 | ALEKSIC J, BIZZARI F, CAI Y, et al. Development of a novel biosensor for the detection of arsenic in drinking water[J]. IET Synthetic Biology, 2007, 1(1/2): 87-90. |
| 40 | FEENEY K A, PUTKER M, BRANCACCIO M, et al. In-depth characterization of firefly luciferase as a reporter of circadian gene expression in mammalian cells[J]. Journal of Biological Rhythms, 2016, 31(6): 540-550. |
| 41 | TINIKUL R, CHUNTHABOON P, PHONBUPPHA J, et al. Bacterial luciferase: molecular mechanisms and applications[J]. The Enzymes, 2020, 47: 427-455. |
| 42 | LOPRESIDE A, CALABRETTA M M, MONTALI L, et al. Prêt-à-porter nanoYESα and nanoYESβ bioluminescent cell biosensors for ultrarapid and sensitive screening of endocrine-disrupting chemicals[J]. Analytical and Bioanalytical Chemistry, 2019, 411(19): 4937-4949. |
| 43 | LOPRESIDE A, WAN X Y, MICHELINI E, et al. Comprehensive profiling of diverse genetic reporters with application to whole-cell and cell-free biosensors[J]. Analytical Chemistry, 2019, 91(23): 15284-15292. |
| 44 | DIETRICH J A, MCKEE A E, KEASLING J D. High-throughput metabolic engineering: advances in small-molecule screening and selection[J]. Annual Review of Biochemistry, 2010, 79:563-590. |
| 45 | CHONG H Q, CHING C B. Development of colorimetric-based whole-cell biosensor for organophosphorus compounds by engineering transcription regulator DmpR [J]. ACS Synthetic Biology, 2016, 5(11): 1290-1298. |
| 46 | SHIN H J. Development of highly-sensitive microbial biosensors by mutation of the nahR regulatory gene[J]. Journal of Biotechnology, 2010, 150(2): 246-250. |
| 47 | TEO W S, HEE K S, CHANG M W. Bacterial FadR and synthetic promoters function as modular fatty acid sensor- regulators in Saccharomyces cerevisiae [J]. Engineering in Life Sciences, 2013, 13(5): 456-463. |
| 48 | TEO W S, CHANG M W. Bacterial XylRs and synthetic promoters function as genetically encoded xylose biosensors in Saccharomyces cerevisiae [J]. Biotechnology Journal, 2015, 10(2): 315-322. |
| 49 | LIANG C N, ZHANG X X, WU J Y, et al. Dynamic control of toxic natural product biosynthesis by an artificial regulatory circuit[J]. Metabolic Engineering, 2020, 57: 239-246. |
| 50 | TANG S Y, CIRINO P C. Design and application of a mevalonate-responsive regulatory protein[J]. Angewandte Chemie-International Edition, 2011, 50(5): 1084-1086. |
| 51 | TANG S Y, QIAN S, AKINTERINWA O, et al. Screening for enhanced triacetic acid lactone production by recombinant Escherichia coli expressing a designed triacetic acid lactone reporter[J]. Journal of the American Chemical Society, 2013, 135(27): 10099-10103. |
| 52 | FREI C S, WANG Z Q, QIAN S, et al. Analysis of amino acid substitutions in AraC variants that respond to triacetic acid lactone[J]. Protein Science, 2016, 25(4): 804-814. |
| 53 | CHEN W, ZHANG S, JIANG P X, et al. Design of an ectoine-responsive AraC mutant and its application in metabolic engineering of ectoine biosynthesis[J]. Metabolic Engineering, 2015, 30: 149-155. |
| 54 | XIONG D D, LU S K, WU J Y, et al. Improving key enzyme activity in phenylpropanoid pathway with a designed biosensor[J]. Metabolic Engineering, 2017, 40: 115-123. |
| 55 | YEOM S J, KIM M, KWON K K, et al. A synthetic microbial biosensor for high-throughput screening of lactam biocatalysts[J]. Nature Communications, 2018, 9: 5053. |
| 56 | MEINHARDT S, MANLEY M W, BECKER N A, et al. Novel insights from hybrid LacI/GalR proteins: family-wide functional attributes and biologically significant variation in transcription repression[J]. Nucleic Acids Research, 2012, 40(21): 11139-11154. |
| 57 | CHOU H H, KEASLING J D. Programming adaptive control to evolve increased metabolite production[J]. Nature Communications, 2013, 4: 2595. |
| 58 | ZHANG F Z, CAROTHERS J M, KEASLING J D. Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids[J]. Nature Biotechnology, 2012, 30(4): 354-359. |
| 59 | DABIRIAN Y, LI X W, CHEN Y, et al. Expanding the dynamic range of a transcription factor-based biosensor in Saccharomyces cerevisiae [J]. ACS Synthetic Biology, 2019, 8(9): 1968-1975. |
| 60 | DAVID F, NIELSEN J, SIEWERS V. Flux control at the malonyl-CoA node through hierarchical dynamic pathway regulation in Saccharomyces cerevisiae [J]. ACS Synthetic Biology, 2016, 5(3): 224-233. |
| 61 | CHEN Y, HO J M L, SHIS D L, et al. Tuning the dynamic range of bacterial promoters regulated by ligand-inducible transcription factors[J]. Nature Communications, 2018, 9: 64. |
| 62 | LI S J, SI T, WANG M, et al. Development of a synthetic malonyl-CoA sensor in Saccharomyces cerevisiae for intracellular metabolite monitoring and genetic screening [J]. ACS Synthetic Biology, 2015, 4(12): 1308-1315. |
| 63 | PANG Q X, HAN H, LIU X Q, et al. In vivo evolutionary engineering of riboswitch with high-threshold for N-acetylneuraminic acid production[J]. Metabolic Engineering, 2020, 59: 36-43. |
| 64 | JANG S H, JANG S Y, XIU Y, et al. Development of artificial riboswitches for monitoring of naringenin in vivo [J]. ACS Synthetic Biology, 2017, 6(11): 2077-2085. |
| 65 | JANG S Y, JANG S H, IM D K, et al. Artificial caprolactam-specific riboswitch as an intracellular metabolite sensor[J]. ACS Synthetic Biology, 2019, 8(6): 1276-1283. |
| 66 | DWIDAR M, SEIKE Y, KOBORI S, et al. Programmable artificial cells using histamine-responsive synthetic riboswitch[J]. Journal of the American Chemical Society, 2019, 141(28): 11103-11114. |
| 67 | YANG J, SEO S W, JANG S H, et al. Synthetic RNA devices to expedite the evolution of metabolite-producing microbes[J]. Nature Communications, 2013, 4: 1413. |
| 68 | WEIGAND J E, SANCHEZ M, GUNNESCH EB, et al. Screening for engineered neomycin riboswitches that control translation initiation[J]. RNA, 2008, 14(1): 89-97. |
| 69 | GROHER F, BOFILL-BOSCH C, SCHNEIDER C, et al. Riboswitching with ciprofloxacin-development and characterization of a novel RNA regulator[J]. Nucleic Acids Research, 2018, 46(4): 2121-2132. |
| 70 | BOUSSEBAYLE A, TORKA D, OLLIVAUD S, et al. Next-level riboswitch development-implementation of Capture-SELEX facilitates identification of a new synthetic riboswitch[J]. Nucleic Acids Research, 2019, 47(9): 4883-4895. |
| 71 | WERSTUCK G, GREEN M R. Controlling gene expression in living cells through small molecule-RNA interactions[J]. Science, 1998, 282(5387): 296-298. |
| 72 | GRATE D, WILSON C. Inducible regulation of the S. cerevisiae cell cycle mediated by an RNA aptamer-ligand complex[J]. Bioorganic & Medicinal Chemistry, 2001, 9(10): 2565-2570. |
| 73 | HARVEY I, GARNEAU P, PELLETIER J. Inhibition of translation by RNA-small molecule interactions[J]. RNA, 2002, 8(4): 452-463. |
| 74 | SUESS B, FINK B, BERENS C, et al. A theophylline responsive riboswitch based on helix slipping controls gene expression in vivo [J]. Nucleic Acids Research, 2004, 32(4): 1610-1614. |
| 75 | WIELAND M, BENZ A, KLAUSER B, et al. Artificial ribozyme switches containing natural riboswitch aptamer domains[J]. Angewandte Chemie-International Edition, 2009, 48: 2715-2718. |
| 76 | WACHSMUTH M, FINDEISS S, WEISSHEIMER N, et al. De novo design of a synthetic riboswitch that regulates transcription termination[J]. Nucleic Acids Research, 2013, 41(4): 2541-2551. |
| 77 | TEO W S, CHANG M W. Development and characterization of AND-gate dynamic controllers with a modular synthetic GAL1 core promoter in Saccharomyces cerevisiae [J]. Biotechnology and Bioengineering, 2014, 111(1): 144-151. |
| 78 | ANDERSON J C, VOIGT C A, ARKIN A P. Environmental signal integration by a modular AND gate[J]. Molecular Systems Biology, 2007, 3: 133. |
| 79 | WANG B J, KITNEY R I, JOLY N, et al. Engineering modular and orthogonal genetic logic gates for robust digital-like synthetic biology[J]. Nature Communications, 2011, 2: 508. |
| 80 | TAMSIR A, TABOR J J, VOIGT C A. Robust multicellular computing using genetically encoded NOR gates and chemical 'wires' [J]. Nature, 2011, 469(7329): 212-215. |
| 81 | WHANGSUK W, THIENGMAG S, DUBBS J, et al. Specific detection of the pesticide chlorpyrifos by a sensitive genetic-based whole cell biosensor[J]. Analytical Biochemistry, 2016, 493: 11-13. |
| 82 | HU Q, LI L, WANG Y J, et al. Construction of WCB-11: a novel phiYFP arsenic-resistant whole-cell biosensor[J]. Journal of Environmental Sciences, 2010, 9(22): 1469-1474. |
| 83 | PELLINEN T, BYLUND G, VIRTA M, et al. Detection of traces of tetracyclines from fish with a bioluminescent sensor strain incorporating bacterial luciferase reporter genes[J]. Journal of Agricultural and Food Chemistry, 2002, 50(17): 4812-4815. |
| 84 | DAEFFLER K N M, GALLEY J D, SHETH R U, et al. Engineering bacterial thiosulfate and tetrathionate sensors for detecting gut inflammation[J]. Molecular Systems Biology, 2017, 13(4): 923. |
| 85 | JO J J, SHIN J S. Construction of intragenic synthetic riboswitches for detection of a small molecule[J]. Biotechnology Letters, 2009, 31(10): 1577-1581. |
| 86 | KEMMER C, GITZINGER M, DAOUD-EL BABA M, et al. Self-sufficient control of urate homeostasis in mice by a synthetic circuit[J]. Nature Biotechnology, 2010, 28(4): 355-360. |
| 87 | PICKUP J C, KHAN F, ZHI Z L. Fluorescence intensity- and lifetime-based glucose sensing using glucose/galactose-binding protein[J]. Journal of Diabetes Science and Technology, 2013, 7(1): 62-71. |
| 88 | 于政, 申晓林, 孙新晓, 等. 动态调控策略在代谢工程中的应用研究进[J]. 合成生物学, 2020, 1(4): 440-453. |
| YU Z, SHEN X L, SUN X X, et al. Application of dynamic regulation strategies in metabolic engineering[J]. Synthetic Biology Journal, 2020, 1(4): 440-453. | |
| 89 | LIU D, XIAO Y, EVANS B S, et al. Negative feedback regulation of fatty acid production based on a malonyl-CoA sensor-actuator [J]. ACS Synthetic Biology, 2015, 4(2): 132-140. |
| 90 | DE LAS HERAS A, MARTINEZ-GARCIA E, DOMINGO-SANANES M R, et al. Rationally rewiring the connectivity of the XylR/Pu regulatory node of the m-xylene degradation pathway in Pseudomonas putida [J]. Integrative Biology, 2016, 8(4): 571-576. |
| 91 | WERLEN C, JASPERS M C M, VAN DER MEER J R. Measurement of biologically available naphthalene in gas and aqueous phases by use of a Pseudomonas putida biosensor [J]. Applied and Environmental Microbiology, 2004, 70(1): 43-51. |
| 92 | MADHUSHANI A, DEL PESO-SANTOS T, MORENO R, et al. Transcriptional and translational control through the 5'-leader region of the dmpR master regulatory gene of phenol metabolism[J]. Environmental Microbiology, 2015, 17(1): 119-133. |
| 93 | JOE M H, LEE K H, LIM S Y, et al. Pigment-based whole-cell biosensor system for cadmium detection using genetically engineered Deinococcus radiodurans [J]. Bioprocess and Biosystems Engineering, 2012, 35(1/2): 265-272. |
| 94 | RAVIKUMAR S, GANESH I, YOO I K, et al. Construction of a bacterial biosensor for zinc and copper and its application to the development of multifunctional heavy metal adsorption bacteria[J]. Process Biochemistry, 2012, 47(5): 758-765. |
| 95 | KISKER C, HINRICHS W, TOVAR K, et al. The complex formed between Tet repressor and tetracycline-Mg2+ reveals mechanism of antibiotic resistance[J]. Journal of Molecular Biology, 1995, 247(2): 260-280. |
| 96 | CHANG Y M, CHEN C K M, KO T P, et al. Structural analysis of the antibiotic-recognition mechanism of MarR proteins[J]. Acta Crystallographica Section D-Structural Biology, 2013, 69(6): 1138-1149. |
| 97 | WANG B J, BARAHONA M, BUCK M. A modular cell-based biosensor using engineered genetic logic circuits to detect and integrate multiple environmental signals[J]. Biosensors and Bioelectronics, 2013, 40(1): 368-376. |
| 98 | FERNANDEZ-LOPEZ R, RUIZ R, DE LA CRUZ F, et al. Transcription factor-based biosensors enlightened by the analyte[J]. Frontiers in Microbiology, 2015, 6: 648. |
| 99 | CHEN X, LV Y, WU R Q. Current technologies of synthetic biosensors for disease detection: design, classification and future perspectives[J]. Chinese Medical Sciences Journal, 2018, 33(4): 240-251. |
| 100 | AUSLANDER D, EGGERSCHWILER B, KEMMER C, et al. A designer cell-based histamine-specific human allergy profiler[J]. Nature Communications, 2014, 5: 4408. |
| 101 | BAI P, YE H F, XIE M Q, et al. A synthetic biology-based device prevents liver injury in mice[J]. Journal of Hepatology, 2016, 65(1): 84-94. |
| 102 | YE H F, XIE M Q, XUE S, et al. Self-adjusting synthetic gene circuit for correcting insulin resistance[J]. Nature Biomedical Engineering, 2016, 1: 0005. |
| 103 | SAXENA P, CHARPIN-EL HAMRI G, FOLCHER M, et al. Synthetic gene network restoring endogenous pituitary-thyroid feedback control in experimental Graves' disease[J]. Proceedings of the National Academy of Sciences of the United States of America, 2016, 113(5): 1244-1249. |
| 104 | SMOLE A, LAINSCEK D, BEZELJAK U, et al. A synthetic mammalian therapeutic gene circuit for sensing and suppressing inflammation[J]. Molecular Therapy, 2017, 25(1): 102-119. |
| 105 | ZAGER V, CEMAZAR M, HRELJAC I, et al. Development of human cell biosensor system for genotoxicity detection based on DNA damage-induced gene expression[J]. Radiology and Oncology, 2010, 44(1): 42-51. |
| 106 | 刘泽辉, 张亚同, 胡欣. 我院茶碱血药浓度监测及其影响因素综合性评[J]. 中国新药杂志, 2016, 25(21): 2514-2520. |
| LIU Z H, ZHANG Y T, HU X. Analysis of influence factors on blood concentration monitoring of theophylline in our hospital [J]. Chinese Journal of New Drug, 2016, 25(21): 2514-2520. | |
| 107 | TIAN J Z, YANG G H, GU Y, et al. Developing an endogenous quorum-sensing based CRISPRi circuit for autonomous and tunable dynamic regulation of multiple targets in Streptomyces[J]. Nucleic Acids Research, 2020, 48(14): 8188-8202. |
| 108 | CUNNINGHAM-BRYANT D, SUN J W, FERNANDEZ B, et al. CRISPR-Cas-mediated chemical control of transcriptional dynamics in yeast[J]. ChemBioChem, 2019, 20(12): 1519-1523. |
| 109 | WAN X Y, VOLPETTI F, PETROVA E, et al. Cascaded amplifying circuits enable ultrasensitive cellular sensors for toxic metals[J]. Nature Chemical Biology, 2019, 15(5): 540-548. |
| [1] | GAO Ge, BIAN Qi, WANG Baojun. Synthetic genetic circuit engineering: principles, advances and prospects [J]. Synthetic Biology Journal, 2025, 6(1): 45-64. |
| [2] | LI Geng, SHEN Xiaolin, SUN Xinxiao, WANG Jia, YUAN Qipeng. Research progress in recombinant expression and application of peroxidases [J]. Synthetic Biology Journal, 2024, 5(6): 1498-1517. |
| [3] | DU Yao, GAO Hongdan, LIU Jiakun, LIU Xiaorong, XING Zhihao, ZHANG Tao, MA Dongli. Research progress of the CRISPR-Cas system in the detecting pathogen nucleic acids [J]. Synthetic Biology Journal, 2024, 5(1): 202-216. |
| [4] | MA Cui, YANG Fan, ZHANG Juntai, HE Kai. Multifunction microplate reader for automated foundry platform [J]. Synthetic Biology Journal, 2023, 4(5): 1036-1049. |
| [5] | QIN Weitong, YANG Guangyu. Research and application progress of microdroplets high throughput screening methods [J]. Synthetic Biology Journal, 2023, 4(5): 966-979. |
| [6] | SUN Zhi, YANG Ning, LOU Chunbo, TANG Chao, YANG Xiaojing. Rational design for functional topology and its applications in synthetic biology [J]. Synthetic Biology Journal, 2023, 4(3): 444-463. |
| [7] | Jiaqi HOU, Nan JIANG, Lianju MA, Yuan LU. Cell-free protein synthesis: from basic research to engineering applications [J]. Synthetic Biology Journal, 2022, 3(3): 465-486. |
| [8] | XU Xinxin, KUANG Hua. Advances in the biological detection of food contaminants based on synthetic receptors [J]. Synthetic Biology Journal, 2022, 3(2): 399-414. |
| [9] | Pan CHU, Jingwen ZHU, Wenqi HUANG, Chenli LIU, Xiongfei FU. Host-circuit coupling: toward a new framework for genetic circuit design [J]. Synthetic Biology Journal, 2021, 2(1): 91-105. |
| [10] | Xiaomeng LI, Wei JIANG, Quanfeng LIANG, Qingsheng QI. Application of bacterial quorum sensing system in intercellular communication and its progress in synthetic biology [J]. Synthetic Biology Journal, 2020, 1(5): 540-555. |
| [11] | Bo ZHANG, Yongshuo MA, Yi SHANG, Sanwen HUANG. Recent advances in plant synthetic biology [J]. Synthetic Biology Journal, 2020, 1(2): 121-140. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||