合成生物学 ›› 2024, Vol. 5 ›› Issue (2): 338-352.DOI: 10.12211/2096-8280.2023-054
合成生物学在多糖结合疫苗研发中的应用
叶精勤, 黄文华, 潘超, 朱力, 王恒樑
- 军事科学院军事医学研究院,病原微生物生物安全全国重点实验室,北京 100071
-
收稿日期:2023-08-09修回日期:2023-11-02出版日期:2024-04-30发布日期:2024-04-28 -
通讯作者:王恒樑 -
作者简介:叶精勤 (1996—),女, 博士研究生。研究方向为传染病疫苗研究。E-mail:yejq0922@163.com王恒樑 (1971—),男,博士,研究员,博士生导师。军事科学院军事医学研究院生物工程研究所副所长,病原微生物生物安全国家重点实验室副主任。主要从事病原细菌致病机理及基因工程疫苗研究。E-mail:wanghl@bmi.ac.cn -
基金资助:国家重点研发计划(2021YFC2102100);国家自然科学基金(32271507);北京市科技新星计划(2022045)
Applications of synthetic biology in developing polysaccharide conjugate vaccines
YE Jingqin, HUANG Wenhua, PAN Chao, ZHU Li, WANG Hengliang
- State Key Laboratory of Pathogen and Biosecurity,Academy of Military Medical Sciences,Academy of Military Sciences,Beijing 100071,China
-
Received:2023-08-09Revised:2023-11-02Online:2024-04-30Published:2024-04-28 -
Contact:WANG Hengliang
摘要:
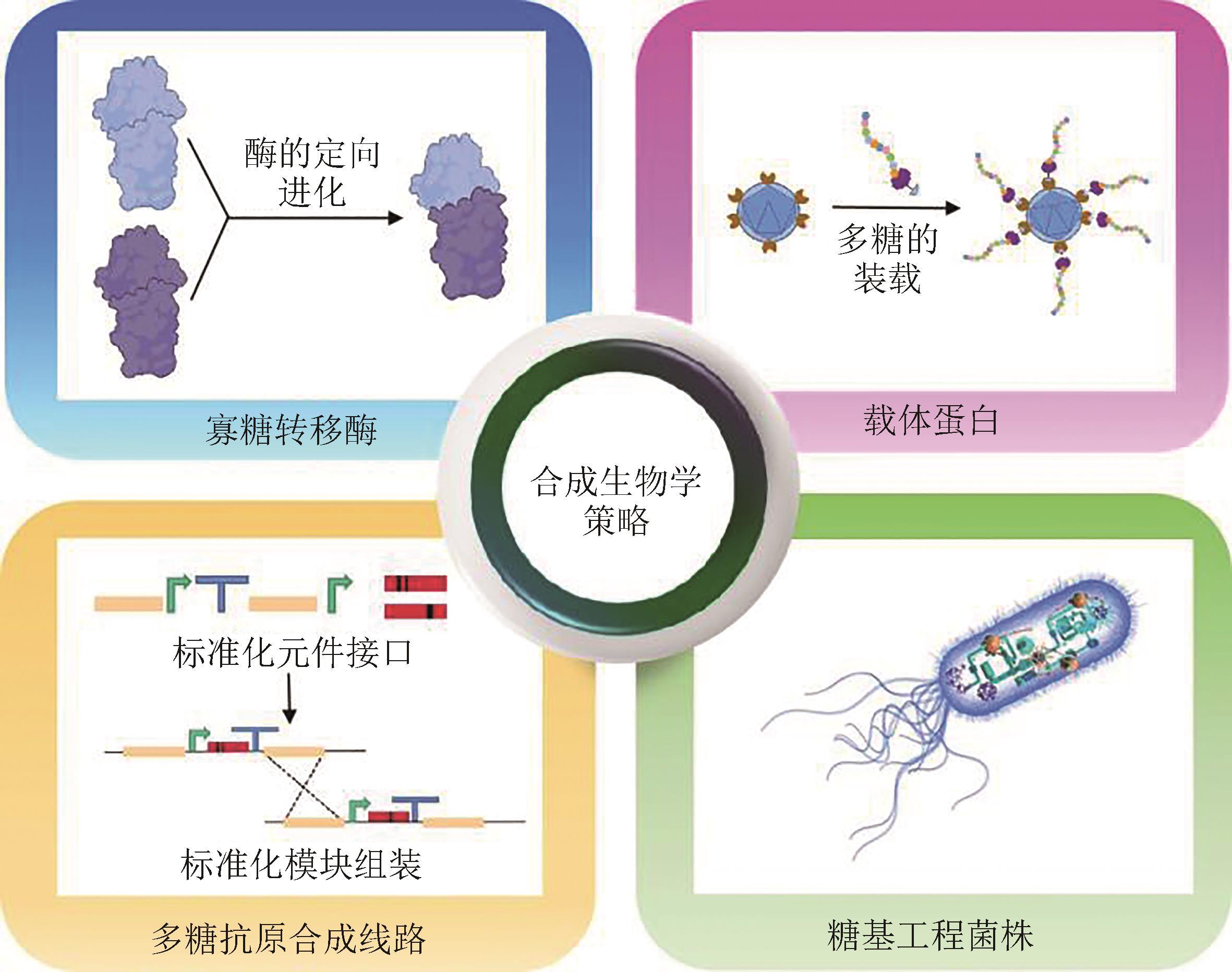
由于合成生物学的快速发展,目前已经实现了人工设计DNA和蛋白质的合成,对具有重要生物学功能、结构也更为复杂的糖类物质进行精准设计合成,也将是合成生物学未来发展的重要方向。近年来,一种基于细菌寡糖转移酶体系的蛋白多糖偶联技术发展迅速,已被广泛应用于病原细菌多糖结合疫苗的生物合成制备。本文综述了该技术体系中的寡糖转移酶元件、载体蛋白元件、异源多糖抗原合成线路以及工程菌株改造等核心关键模块的最新研究进展。使用生物活体系统,发酵生产多糖结合疫苗,具有产物均一性好、步骤简便、绿色环保等优势,是一种亟待发展的新兴技术,同时也存在一些技术细节需要完善。未来,寡糖转移酶的定向进化、纳米颗粒型蛋白载体的应用、多糖合成基因的组合重排、工程菌株的代谢途径优化,将有望进一步促进多糖结合疫苗的生物合成研究。
中图分类号:
引用本文
叶精勤, 黄文华, 潘超, 朱力, 王恒樑. 合成生物学在多糖结合疫苗研发中的应用[J]. 合成生物学, 2024, 5(2): 338-352.
YE Jingqin, HUANG Wenhua, PAN Chao, ZHU Li, WANG Hengliang. Applications of synthetic biology in developing polysaccharide conjugate vaccines[J]. Synthetic Biology Journal, 2024, 5(2): 338-352.
| 菌株 | 抗原 | 载体蛋白 | 偶联技术 | 研究阶段 |
|---|---|---|---|---|
| B型流感嗜血杆菌 | 天然多糖 | TT | 化学偶联 | 已授权 |
| 寡糖 | CRM197 | 化学偶联 | 已授权 | |
| C型脑膜炎奈瑟氏菌 | 天然多糖 | TT | 化学偶联 | 已授权 |
| 寡糖 | CRM197 | 化学偶联 | 已授权 | |
| ACWY型脑膜炎奈瑟氏菌 | 寡糖 | CRM197 | 化学偶联 | 已授权 |
| 肺炎链球菌 | 天然多糖 | PD, DT, TT | 化学偶联 | 已授权 |
| 肺炎链球菌 | 天然多糖 | 链霉亲和素融合蛋白 | 生物偶联 | 临床Ⅱ期 |
| 肠外致病大肠杆菌 | 寡糖 | EPA | 生物偶联 | 临床Ⅲ期 |
| 志贺氏菌2a | 寡糖 | EPA | 生物偶联 | 临床Ⅰ期 |
| 寡糖 | TT | 化学反应 | 临床Ⅰ期 | |
| 肺炎克雷伯氏菌 | O-抗原寡糖 | EPA | 生物偶联 | 临床Ⅰ期 |
表1 多糖结合疫苗中所采用的载体蛋白情况汇总
Table 1 Summary of carrier proteins used in polysaccharide conjugate vaccines
| 菌株 | 抗原 | 载体蛋白 | 偶联技术 | 研究阶段 |
|---|---|---|---|---|
| B型流感嗜血杆菌 | 天然多糖 | TT | 化学偶联 | 已授权 |
| 寡糖 | CRM197 | 化学偶联 | 已授权 | |
| C型脑膜炎奈瑟氏菌 | 天然多糖 | TT | 化学偶联 | 已授权 |
| 寡糖 | CRM197 | 化学偶联 | 已授权 | |
| ACWY型脑膜炎奈瑟氏菌 | 寡糖 | CRM197 | 化学偶联 | 已授权 |
| 肺炎链球菌 | 天然多糖 | PD, DT, TT | 化学偶联 | 已授权 |
| 肺炎链球菌 | 天然多糖 | 链霉亲和素融合蛋白 | 生物偶联 | 临床Ⅱ期 |
| 肠外致病大肠杆菌 | 寡糖 | EPA | 生物偶联 | 临床Ⅲ期 |
| 志贺氏菌2a | 寡糖 | EPA | 生物偶联 | 临床Ⅰ期 |
| 寡糖 | TT | 化学反应 | 临床Ⅰ期 | |
| 肺炎克雷伯氏菌 | O-抗原寡糖 | EPA | 生物偶联 | 临床Ⅰ期 |
| 1 | ANDRIANANTOANDRO E, BASU S, KARIG D K, et al. Synthetic biology: new engineering rules for an emerging discipline[J]. Molecular Systems Biology, 2006, 2: 2006.0028. |
| 2 | BRENNER K, YOU L C, ARNOLD F H. Engineering microbial consortia: a new frontier in synthetic biology[J]. Trends in Biotechnology, 2008, 26(9): 483-489. |
| 3 | JAROENTOMEECHAI T, TAW M N, LI M J, et al. Cell-free synthetic glycobiology: designing and engineering glycomolecules outside of living cells[J]. Frontiers in Chemistry, 2020, 8: 645. |
| 4 | KHOURY G A, BALIBAN R C, FLOUDAS C A. Proteome-wide post-translational modification statistics: frequency analysis and curation of the swiss-prot database[J]. Scientific Reports, 2011, 1: 90. |
| 5 | HELENIUS A, AEBI A M. Intracellular functions of N-linked glycans[J]. Science, 2001, 291(5512): 2364-2369. |
| 6 | RUDD P M, WORMALD M R, STANFIELD R L, et al. Roles for glycosylation of cell surface receptors involved in cellular immune recognition[J]. Journal of Molecular Biology, 1999, 293(2): 351-366. |
| 7 | TYTGAT H L P, LEBEER S. The sweet tooth of bacteria: common themes in bacterial glycoconjugates[J]. Microbiology and Molecular Biology Reviews: MMBR, 2014, 78(3): 372-417. |
| 8 | STRENG-OUWEHAND I, HO N I, LITJENS M, et al. Glycan modification of antigen alters its intracellular routing in dendritic cells, promoting priming of T cells[J]. eLife, 2016, 5: e11765. |
| 9 | PHANSE Y, CARRILLO-CONDE B R, RAMER-TAIT A E, et al. A systems approach to designing next generation vaccines: combining α-galactose modified antigens with nanoparticle platforms[J]. Scientific Reports, 2014, 4: 3775. |
| 10 | ELLIOTT S, LORENZINI T, ASHER S, et al. Enhancement of therapeutic protein in vivo activities through glycoengineering[J]. Nature Biotechnology, 2003, 21(4): 414-421. |
| 11 | OHTSUBO K, MARTH J D. Glycosylation in cellular mechanisms of health and disease[J]. Cell, 2006, 126(5): 855-867. |
| 12 | ZHANG Q, JOHNSTON E V, SHIEH J H, et al. Synthesis of granulocyte-macrophage colony-stimulating factor as homogeneous glycoforms and early comparisons with yeast cell-derived material[J]. Proceedings of the National Academy of Sciences of the United States of America, 2014, 111(8): 2885-2890. |
| 13 | CHUNG C H, MIRAKHUR B, CHAN E, et al. Cetuximab-induced anaphylaxis and IgE specific for galactose-α-1,3-galactose[J]. New England Journal of Medicine, 2008, 358(11): 1109-1117. |
| 14 | LI H J, SETHURAMAN N, STADHEIM T A, et al. Optimization of humanized IgGs in glycoengineered Pichia pastoris [J]. Nature Biotechnology, 2006, 24(2): 210-215. |
| 15 | TIAN W H, YE Z L, WANG S J, et al. The glycosylation design space for recombinant lysosomal replacement enzymes produced in CHO cells[J]. Nature Communications, 2019, 10: 1785. |
| 16 | ABREU A G, BARBOSA A S. How Escherichia coli circumvent complement-mediated killing[J]. Frontiers in Immunology, 2017, 8: 452. |
| 17 | BUNDLE D. Antibacterials: a sweet vaccine[J]. Nature Chemistry, 2016, 8(3): 201-202. |
| 18 | SZYMANSKI C M, YAO R J, EWING C P, et al. Evidence for a system of general protein glycosylation in Campylobacter jejuni [J]. Molecular Microbiology, 1999, 32(5): 1022-1030. |
| 19 | PARKHILL J, WREN B W, MUNGALL K, et al. The genome sequence of the food-borne pathogen Campylobacter jejuni reveals hypervariable sequences[J]. Nature, 2000, 403(6770): 665-668. |
| 20 | FELDMAN M F, WACKER M, HERNANDEZ M, et al. Engineering N-linked protein glycosylation with diverse O antigen lipopolysaccharide structures in Escherichia coli [J]. Proceedings of the National Academy of Sciences of the United States of America, 2005, 102(8): 3016-3021. |
| 21 | BERTANI B, RUIZ N. Function and biogenesis of lipopolysaccharides[J]. EcoSal Plus, 2018, 8(1): ESP-0001-2018. |
| 22 | WHITFIELD C, WEAR S S, SANDE C. Assembly of bacterial capsular polysaccharides and exopolysaccharides[J]. Annual Review of Microbiology, 2020, 74: 521-543. |
| 23 | KOWARIK M, YOUNG N M, NUMAO S, et al. Definition of the bacterial N-glycosylation site consensus sequence[J]. The EMBO Journal, 2006, 25(9): 1957-1966. |
| 24 | IHSSEN J, KOWARIK M, DILETTOSO S, et al. Production of glycoprotein vaccines in Escherichia coli [J]. Microbial Cell Factories, 2010, 9: 61. |
| 25 | HATZ C F R, BALLY B, ROHRER S, et al. Safety and immunogenicity of a candidate bioconjugate vaccine against Shigella dysenteriae type 1 administered to healthy adults: a single blind, partially randomized Phase Ⅰ study[J]. Vaccine, 2015, 33(36): 4594-4601. |
| 26 | RAVENSCROFT N, BRAUN M, SCHNEIDER J, et al. Characterization and immunogenicity of a Shigella flexneri 2a O-antigen bioconjugate vaccine candidate[J]. Glycobiology, 2019, 29(9): 669-680. |
| 27 | VAN DEN DOBBELSTEEN G P J M, FAÉ K C, SERROYEN J, et al. Immunogenicity and safety of a tetravalent E. coli O-antigen bioconjugate vaccine in animal models[J]. Vaccine, 2016, 34(35): 4152-4160. |
| 28 | MARSHALL L E, NELSON M, DAVIES C H, et al. An O-antigen glycoconjugate vaccine produced using protein glycan coupling technology is protective in an inhalational rat model of tularemia[J]. Journal of Immunology Research, 2018, 2018: 8087916. |
| 29 | GARCIA-QUINTANILLA F, IWASHKIW J A, PRICE N L, et al. Production of a recombinant vaccine candidate against Burkholderia pseudomallei exploiting the bacterial N-glycosylation machinery[J]. Frontiers in Microbiology, 2014, 5: 381. |
| 30 | IWASHKIW J A, FENTABIL M A, FARIDMOAYER A, et al. Exploiting the Campylobacter jejuni protein glycosylation system for glycoengineering vaccines and diagnostic tools directed against brucellosis[J]. Microbial Cell Factories, 2012, 11: 13. |
| 31 | WACKER M, WANG L H, KOWARIK M, et al. Prevention of Staphylococcus aureus infections by glycoprotein vaccines synthesized in Escherichia coli [J]. The Journal of Infectious Diseases, 2014, 209(10): 1551-1561. |
| 32 | HERBERT J A, KAY E J, FAUSTINI S E, et al. Production and efficacy of a low-cost recombinant pneumococcal protein polysaccharide conjugate vaccine[J]. Vaccine, 2018, 36(26): 3809-3819. |
| 33 | LIZAK C, GERBER S, NUMAO S, et al. X-ray structure of a bacterial oligosaccharyltransferase[J]. Nature, 2011, 474(7351): 350-355. |
| 34 | NAPIÓRKOWSKA M, BOILEVIN J, SOVDAT T, et al. Molecular basis of lipid-linked oligosaccharide recognition and processing by bacterial oligosaccharyltransferase[J]. Nature Structural & Molecular Biology, 2017, 24(12): 1100-1106. |
| 35 | IHSSEN J, HAAS J, KOWARIK M, et al. Increased efficiency of Campylobacter jejuni N-oligosaccharyltransferase PglB by structure-guided engineering[J]. Open Biology, 2015, 5(4): 140227. |
| 36 | STIMSON E, VIRJI M, BARKER S, et al. Discovery of a novel protein modification: α-glycerophosphate is a substituent of meningococcal pilin[J]. Biochemical Journal, 1996, 316(1): 29-33. |
| 37 | FARIDMOAYER A, FENTABIL M A, MILLS D C, et al. Functional characterization of bacterial oligosaccharyltransferases involved in O-linked protein glycosylation[J]. Journal of Bacteriology, 2007, 189(22): 8088-8098. |
| 38 | FARIDMOAYER A, FENTABIL M A, HAURAT M F, et al. Extreme substrate promiscuity of the Neisseria oligosaccharyl transferase involved in protein O-glycosylation[J]. Journal of Biological Chemistry, 2008, 283(50): 34596-34604. |
| 39 | PAN C, SUN P, LIU B, et al. Biosynthesis of conjugate vaccines using an O-linked glycosylation system[J]. mBio, 2016, 7(2): e00443-16. |
| 40 | SUN P, PAN C, ZENG M, et al. Design and production of conjugate vaccines against S. Paratyphi A using an O-linked glycosylation system in vivo [J]. NPJ Vaccines, 2018, 3: 4. |
| 41 | LI S L, HUANG J, WANG K F, et al. A bioconjugate vaccine against Brucella abortus produced by engineered Escherichia coli [J]. Frontiers in Bioengineering and Biotechnology, 2023, 11: 1121074. |
| 42 | LIU Y, PAN C, WANG K F, et al. Preparation of a Klebsiella pneumoniae conjugate nanovaccine using glycol-engineered Escherichia coli [J]. Microbial Cell Factories, 2023, 22(1): 95. |
| 43 | SCHULZ B L, JEN F E C, POWER P M, et al. Identification of bacterial protein O-oligosaccharyltransferases and their glycoprotein substrates[J]. PLoS One, 2013, 8(5): e62768. |
| 44 | GEBHART C, IELMINI M V, REIZ B, et al. Characterization of exogenous bacterial oligosaccharyltransferases in Escherichia coli reveals the potential for O-linked protein glycosylation in Vibrio cholerae and Burkholderia thailandensis [J]. Glycobiology, 2012, 22(7): 962-974. |
| 45 | ELHENAWY W, SCOTT N E, TONDO M L, et al. Protein O-linked glycosylation in the plant pathogen Ralstonia solanacearum [J]. Glycobiology, 2016, 26(3): 301-311. |
| 46 | HADJINEOPHYTOU C, ANONSEN J H, SVINGERUD T, et al. Sculpting the bacterial O-glycoproteome: functional analyses of orthologous oligosaccharyltransferases with diverse targeting specificities[J]. mBio, 2022, 13(3): e0379721. |
| 47 | HARDING C M, NASR M A, KINSELLA R L, et al. Acinetobacter strains carry two functional oligosaccharyltransferases, one devoted exclusively to type Ⅳ pilin, and the other one dedicated to O-glycosylation of multiple proteins[J]. Molecular Microbiology, 2015, 96(5): 1023-1041. |
| 48 | HARDING C M, NASR M A, SCOTT N E, et al. A platform for glycoengineering a polyvalent pneumococcal bioconjugate vaccine using E. coli as a host[J]. Nature Communications, 2019, 10: 891. |
| 49 | KNOOT C J, ROBINSON L S, HARDING C M. A minimal sequon sufficient for O-linked glycosylation by the versatile oligosaccharyltransferase PglS[J]. Glycobiology, 2021, 31(9): 1192-1203. |
| 50 | FELDMAN M F, MAYER BRIDWELL A E, SCOTT N E, et al. A promising bioconjugate vaccine against hypervirulent Klebsiella pneumoniae [J]. Proceedings of the National Academy of Sciences of the United States of America, 2019, 116(37): 18655-18663. |
| 51 | DUKE J A, PASCHALL A V, ROBINSON L S, et al. Development and immunogenicity of a prototype multivalent group B Streptococcus bioconjugate vaccine[J]. ACS Infectious Diseases, 2021, 7(11): 3111-3123. |
| 52 | KNOOT C J, WANTUCH P L, ROBINSON L S, et al. Discovery and characterization of a new class of O-linking oligosaccharyltransferases from the Moraxellaceae family[J]. Glycobiology, 2023, 33(1): 57-74. |
| 53 | HARVEY H, KUS J V, TESSIER L, et al. Pseudomonas aeruginosa D-arabinofuranose biosynthetic pathway and its role in type Ⅳ pilus assembly[J]. Journal of Biological Chemistry, 2011, 286(32): 28128-28137. |
| 54 | HARDING C M, FELDMAN M F. Glycoengineering bioconjugate vaccines, therapeutics, and diagnostics in E. coli [J]. Glycobiology, 2019, 29(7): 519-529. |
| 55 | ZARSCHLER K, JANESCH B, PABST M, et al. Protein tyrosine O-glycosylation—a rather unexplored prokaryotic glycosylation system[J]. Glycobiology, 2010, 20(6): 787-798. |
| 56 | MAES E, KRZEWINSKI F, GARENAUX E, et al. Glycosylation of BclA glycoprotein from Bacillus cereus and Bacillus anthracis exosporium is domain-specific[J]. Journal of Biological Chemistry, 2016, 291(18): 9666-9677. |
| 57 | DEL BINO L, ØSTERLID K E, WU D Y, et al. Synthetic glycans to improve current glycoconjugate vaccines and fight antimicrobial resistance[J]. Chemical Reviews, 2022, 122(20): 15672-15716. |
| 58 | MICOLI F, COSTANTINO P, ADAMO R. Potential targets for next generation antimicrobial glycoconjugate vaccines[J]. FEMS Microbiology Reviews, 2018, 42(3): 388-423. |
| 59 | HUANG Y L, WU C Y. Carbohydrate-based vaccines: challenges and opportunities[J]. Expert Review of Vaccines, 2010, 9(11): 1257-1274. |
| 60 | WILDER-SMITH A. Meningococcal disease: risk for international travellers and vaccine strategies[J]. Travel Medicine and Infectious Disease, 2008, 6(4): 182-186. |
| 61 | Prevention and control of meningococcal disease: recommendations of the Advisory Committee on Immunization Practices (ACIP) [J/OL]. MMWR Recommendations and Reports, 2000, 49(RR07): 1-10[2023-09-01]. . |
| 62 | SCHUERMAN L, PRYMULA R, HENCKAERTS I, et al. ELISA IgG concentrations and opsonophagocytic activity following pneumococcal protein D conjugate vaccination and relationship to efficacy against acute otitis media[J]. Vaccine, 2007, 25(11): 1962-1968. |
| 63 | PRYMULA R, PEETERS P, CHROBOK V, et al. Pneumococcal capsular polysaccharides conjugated to protein D for prevention of acute otitis media caused by both Streptococcus pneumoniae and non-typable Haemophilus influenzae: a randomised double-blind efficacy study[J]. The Lancet, 2006, 367(9512): 740-748. |
| 64 | ZHU H, ROLLIER C S, POLLARD A J. Recent advances in lipopolysaccharide-based glycoconjugate vaccines[J]. Expert Review of Vaccines, 2021, 20(12): 1515-1538. |
| 65 | COHEN D, ATSMON J, ARTAUD C, et al. Safety and immunogenicity of a synthetic carbohydrate conjugate vaccine against Shigella flexneri 2a in healthy adult volunteers: a phase 1, dose-escalating, single-blind, randomised, placebo-controlled study[J]. The Lancet Infectious Diseases, 2021, 21(4): 546-558. |
| 66 | FRENCK R W JR, ERVIN J, CHU L, et al. Safety and immunogenicity of a vaccine for extra-intestinal pathogenic Escherichia coli (ESTELLA): a phase 2 randomised controlled trial[J]. The Lancet Infectious Diseases, 2019, 19(6): 631-640. |
| 67 | DAVID S, REUTER S, HARRIS S R, et al. Epidemic of carbapenem-resistant Klebsiella pneumoniae in Europe is driven by nosocomial spread[J]. Nature Microbiology, 2019, 4(11): 1919-1929. |
| 68 | SHEN X, LAGERGÅRD T, YANG Y, et al. Group B Streptococcus capsular polysaccharide-cholera toxin B subunit conjugate vaccines prepared by different methods for intranasal immunization[J]. Infection and Immunity, 2001, 69(1): 297-306. |
| 69 | DOW J M, MAURI M, SCOTT T A, et al. Improving protein glycan coupling technology (PGCT) for glycoconjugate vaccine production[J]. Expert Review of Vaccines, 2020, 19(6): 507-527. |
| 70 | ROMANO M R, LEUZZI R, CAPPELLETTI E, et al. Recombinant Clostridium difficile toxin fragments as carrier protein for PSⅡ surface polysaccharide preserve their neutralizing activity[J]. Toxins, 2014, 6(4): 1385-1396. |
| 71 | NILO A, PASSALACQUA I, FABBRINI M, et al. Exploring the effect of conjugation site and chemistry on the immunogenicity of an anti-group B streptococcus glycoconjugate vaccine based on GBS67 pilus protein and type Ⅴ polysaccharide[J]. Bioconjugate Chemistry, 2015, 26(8): 1839-1849. |
| 72 | YUE H, MA G H. Polymeric micro/nanoparticles: particle design and potential vaccine delivery applications[J]. Vaccine, 2015, 33(44): 5927-5936. |
| 73 | NEEK M, KIM T I, WANG S W. Protein-based nanoparticles in cancer vaccine development[J]. Nanomedicine: Nanotechnology, Biology, and Medicine, 2019, 15(1): 164-174. |
| 74 | CHEN J Y, WANG P, YUAN L Z, et al. A live attenuated virus-based intranasal COVID-19 vaccine provides rapid, prolonged, and broad protection against SARS-CoV-2[J]. Science Bulletin, 2022, 67(13): 1372-1387. |
| 75 | CHARLTON HUME H K, VIDIGAL J, CARRONDO M J T, et al. Synthetic biology for bioengineering virus-like particle vaccines[J]. Biotechnology and Bioengineering, 2019, 116(4): 919-935. |
| 76 | LI X, PAN C, SUN P, et al. Orthogonal modular biosynthesis of nanoscale conjugate vaccines for vaccination against infection[J]. Nano Research, 2022, 15(2): 1645-1653. |
| 77 | PAN C, WU J, QING S, et al. Biosynthesis of self-assembled proteinaceous nanoparticles for vaccination[J]. Advanced Materials, 2020, 32(42): e2002940. |
| 78 | HUANG B, XU Y, HU X H, et al. A backbone-centred energy function of neural networks for protein design[J]. Nature, 2022, 602(7897): 523-528. |
| 79 | PENG Z H, WU J, WANG K F, et al. Production of a promising biosynthetic self-assembled nanoconjugate vaccine against Klebsiella pneumoniae serotype O2 in a general Escherichia coli host[J]. Advanced Science, 2021, 8(14): e2100549. |
| 80 | SHI Y X, PAN C, WANG K F, et al. Construction of orthogonal modular proteinaceous nanovaccine delivery vectors based on mSA-biotin binding[J]. Nanomaterials, 2022, 12(5): 734. |
| 81 | TEMME K, ZHAO D H, VOIGT C A. Refactoring the nitrogen fixation gene cluster from Klebsiella oxytoca [J]. Proceedings of the National Academy of Sciences of the United States of America, 2012, 109(18): 7085-7090. |
| 82 | SMANSKI M J, BHATIA S, ZHAO D H, et al. Functional optimization of gene clusters by combinatorial design and assembly[J]. Nature Biotechnology, 2014, 32(12): 1241-1249. |
| 83 | ZELCBUCH L, ANTONOVSKY N, BAR-EVEN A, et al. Spanning high-dimensional expression space using ribosome-binding site combinatorics[J]. Nucleic Acids Research, 2013, 41(9): e98. |
| 84 | TAYLOR G M, MORDAKA P M, HEAP J T. Start-Stop Assembly: a functionally scarless DNA assembly system optimized for metabolic engineering[J]. Nucleic Acids Research, 2019, 47(3): e17. |
| 85 | KAY E J, MAURI M, WILLCOCKS S J, et al. Engineering a suite of E. coli strains for enhanced expression of bacterial polysaccharides and glycoconjugate vaccines[J]. Microbial Cell Factories, 2022, 21(1): 66. |
| 86 | GASPERINI G, RASO M M, SCHIAVO F, et al. Rapid generation of Shigella flexneri GMMA displaying natural or new and cross-reactive O-Antigens[J]. NPJ Vaccines, 2022, 7: 69. |
| 87 | JAROENTOMEECHAI T, KWON Y H, LIU Y W, et al. A universal glycoenzyme biosynthesis pipeline that enables efficient cell-free remodeling of glycans[J]. Nature Communications, 2022, 13: 6325. |
| 88 | GLASSCOCK C J, YATES L E, JAROENTOMEECHAI T, et al. A flow cytometric approach to engineering Escherichia coli for improved eukaryotic protein glycosylation[J]. Metabolic Engineering, 2018, 47: 488-495. |
| 89 | CERONI F, ALGAR R, STAN G B, et al. Quantifying cellular capacity identifies gene expression designs with reduced burden[J]. Nature Methods, 2015, 12(5): 415-418. |
| 90 | LINTON D, DORRELL N, HITCHEN P G, et al. Functional analysis of the Campylobacter jejuni N-linked protein glycosylation pathway[J]. Molecular Microbiology, 2005, 55(6): 1695-1703. |
| 91 | NEUHARD J, THOMASSEN E. Altered deoxyribonucleotide pools in P2 eductants of Escherichia coli K-12 due to deletion of the dcd gene[J]. Journal of Bacteriology, 1976, 126(2): 999-1001. |
| 92 | ALAIMO C, CATREIN I, MORF L, et al. Two distinct but interchangeable mechanisms for flipping of lipid-linked oligosaccharides[J]. The EMBO Journal, 2006, 25(5): 967-976. |
| 93 | PÉREZ J M, MCGARRY M A, MAROLDA C L, et al. Functional analysis of the large periplasmic loop of the Escherichia coli K-12 WaaL O-antigen ligase[J]. Molecular Microbiology, 2008, 70(6): 1424-1440. |
| 94 | MUSUMECI M A, FARIDMOAYER A, WATANABE Y, et al. Evaluating the role of conserved amino acids in bacterial O-oligosaccharyltransferases by in vivo, in vitro and limited proteolysis assays[J]. Glycobiology, 2014, 24(1): 39-50. |
| 95 | STRUTTON B, JAFFÉ S R P, PANDHAL J, et al. Producing a glycosylating Escherichia coli cell factory: the placement of the bacterial oligosaccharyl transferase pglB onto the genome[J]. Biochemical and Biophysical Research Communications, 2018, 495(1): 686-692. |
| 96 | YATES L E, NATARAJAN A, LI M J, et al. Glyco-recoded Escherichia coli: recombineering-based genome editing of native polysaccharide biosynthesis gene clusters[J]. Metabolic Engineering, 2019, 53: 59-68. |
| 97 | NATARAJAN A, JAROENTOMEECHAI T, LI M J, et al. Metabolic engineering of glycoprotein biosynthesis in bacteria[J]. Emerging Topics in Life Sciences, 2018, 2(3): 419-432. |
| 98 | YU H, CHEN X. One-pot multienzyme (OPME) systems for chemoenzymatic synthesis of carbohydrates[J]. Organic & Biomolecular Chemistry, 2016, 14(10): 2809-2818. |
| 99 | GUARINO C, DELISA M P. A prokaryote-based cell-free translation system that efficiently synthesizes glycoproteins[J]. Glycobiology, 2012, 22(5): 596-601. |
| 100 | SHIMIZU Y, INOUE A, TOMARI Y, et al. Cell-free translation reconstituted with purified components[J]. Nature Biotechnology, 2001, 19(8): 751-755. |
| 101 | STARK J C, JAROENTOMEECHAI T, MOELLER T D, et al. On-demand, cell-free biomanufacturing of conjugate vaccines at the point-of-care[EB/OL]. bioRxiv, 2019: 681841[2023-09-01]. . |
| 102 | NATARAJAN A, JAROENTOMEECHAI T, CABRERA-SÁNCHEZ M, et al. Engineering orthogonal human O-linked glycoprotein biosynthesis in bacteria[J]. Nature Chemical Biology, 2020, 16(10): 1062-1070. |
| [1] | 孙扬, 陈立超, 石艳云, 王珂, 吕丹丹, 徐秀美, 张立新. 作物光合作用合成生物学的策略与展望[J]. 合成生物学, 2025, (): 1-16. |
| [2] | 钟奶才, 陈缘, 潘文锋, 苏小凤, 廖景文, 钟近艺. 等离子体微生物育种技术在生物制造中的应用进展[J]. 合成生物学, 2025, (): 1-17. |
| [3] | 狄泽燕, 周铄, 杨铭方, 刘昕, 陈瑶. 功能框架材料在一碳生物转化中的应用[J]. 合成生物学, 2025, (): 1-18. |
| [4] | 张成辛. 基于文本数据挖掘的蛋白功能预测的机遇与挑战[J]. 合成生物学, 2025, (): 1-14. |
| [5] | 姜源旭, 范盈盈, 魏平. 生物振荡的设计原理与人工合成[J]. 合成生物学, 2025, (): 1-15. |
| [6] | 黄姝涵, 马赫, 罗云孜. 生物合成红景天苷的研究进展[J]. 合成生物学, 2025, (): 1-16. |
| [7] | 李倩, James E. Ferrell, 陈于平. 细胞质浓度:细胞生物学的老问题、新参数[J]. 合成生物学, 2025, (): 1-18. |
| [8] | 姜百翼, 钱珑. 活细胞记录器在细胞谱系追踪中的应用和前景[J]. 合成生物学, 2025, (): 1-17. |
| [9] | 徐怀胜, 石晓龙, 刘晓光, 徐苗苗. DNA存储的关键技术:编码、纠错、随机访问与安全性[J]. 合成生物学, 2025, 6(1): 157-176. |
| [10] | 温艳华, 刘合栋, 曹春来, 巫瑞波. 蛋白质工程在医药产业中的应用[J]. 合成生物学, 2025, 6(1): 65-86. |
| [11] | 仲泉周, 单依怡, 裴清云, 金艳芸, 王艺涵, 孟璐远, 王歆韵, 张雨鑫, 刘坤媛, 王慧中, 冯尚国. 生物合成法生产α-熊果苷的研究进展[J]. 合成生物学, 2025, 6(1): 118-135. |
| [12] | 张以恒, 陈雪梅, 石婷. 生物制造的市本率(PC值):定义与应用[J]. 合成生物学, 2025, 6(1): 8-17. |
| [13] | 刘晓悦, 王盼娣, 吴刚, 刘芳. 基因工程辅助萝卜硫苷在十字花科作物中的高效生物合成[J]. 合成生物学, 2025, 6(1): 136-156. |
| [14] | 任家卫, 张金鹏, 徐国强, 张晓梅, 许正宏, 张晓娟. 大肠杆菌中终止子对下游转录单元基因表达的影响[J]. 合成生物学, 2025, 6(1): 213-227. |
| [15] | 唐兴华, 陆钱能, 胡翌霖. 人类世中对合成生物学的哲学反思[J]. 合成生物学, 2025, 6(1): 203-212. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||