合成生物学 ›› 2024, Vol. 5 ›› Issue (3): 548-560.DOI: 10.12211/2096-8280.2023-090
细菌聚酮合酶间的杂合方式及聚酮化合物生物合成逻辑
张瑞, 金文铮, 陈依军
- 中国药科大学生命科学与技术学院化学生物学教研室,江苏 南京 211198
-
收稿日期:2023-11-28修回日期:2024-03-04出版日期:2024-06-30发布日期:2024-07-12 -
通讯作者:陈依军 -
作者简介:张瑞 (1999—),女,硕士研究生。研究方向为聚酮化合物的生物合成。E-mail:zhangrui19990303@163.com陈依军 (1962—),男,教授。研究方向为药物合成生物学。E-mail:yjchen@cpu.edu.cn
Bacterial inter-PKS hybrids and the biosynthetic algorithm of polyketides
ZHANG Rui, JIN Wenzheng, CHEN Yijun
- Laboratory of Chemical Biology,School of Life Science and Technology,China Pharmaceutical University,Nanjing 211198,Jiangsu,China
-
Received:2023-11-28Revised:2024-03-04Online:2024-06-30Published:2024-07-12 -
Contact:CHEN Yijun
摘要:
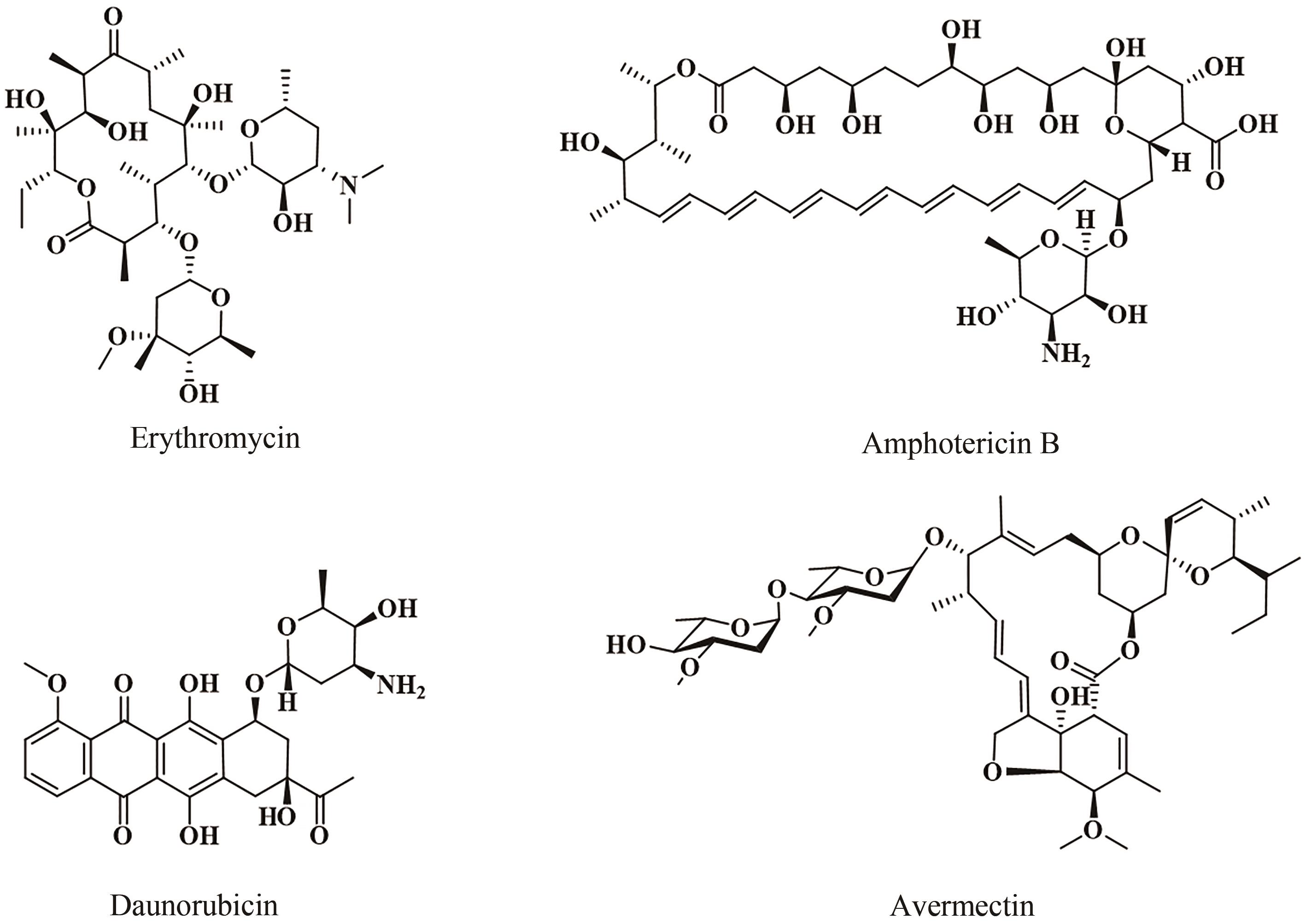
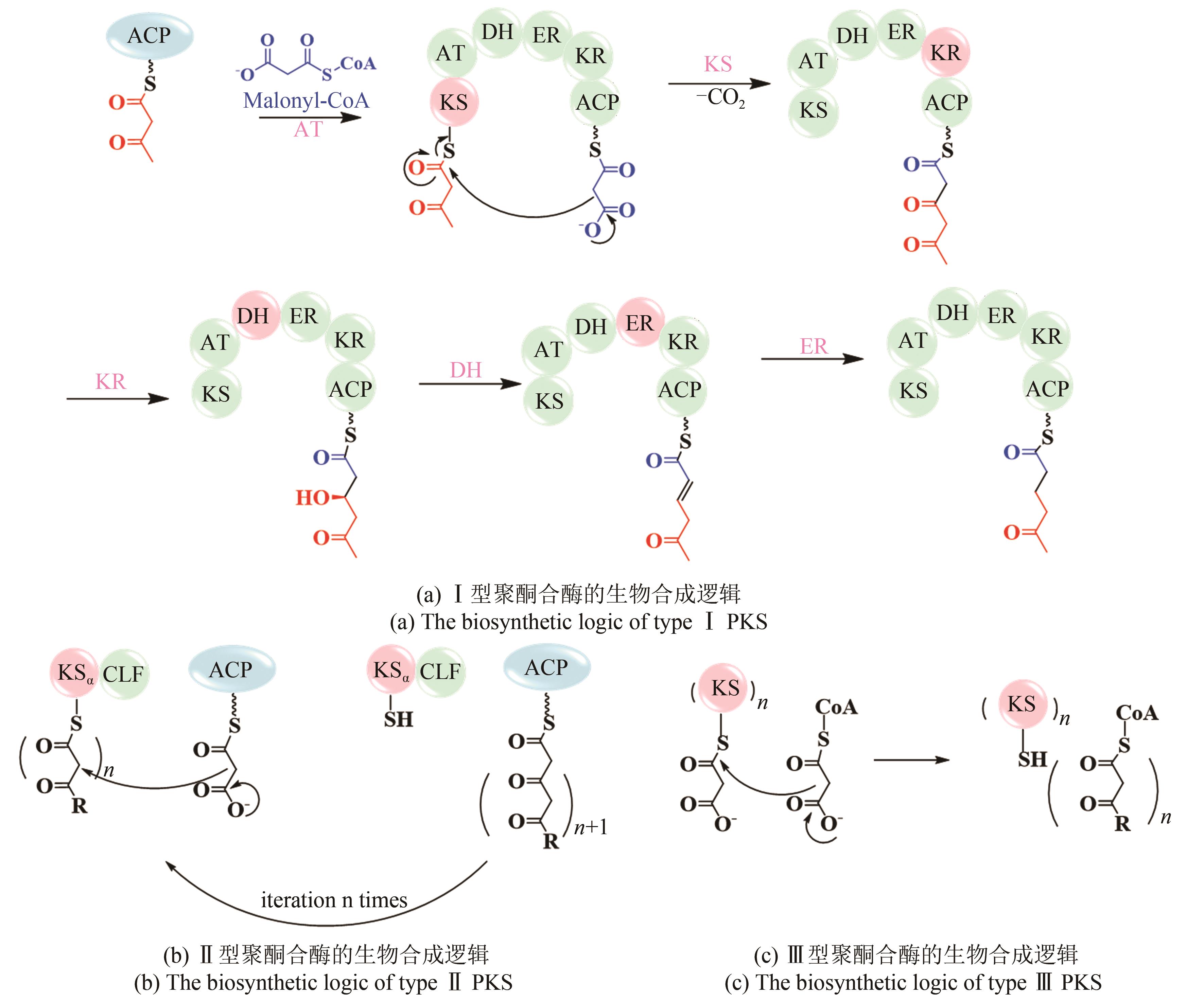
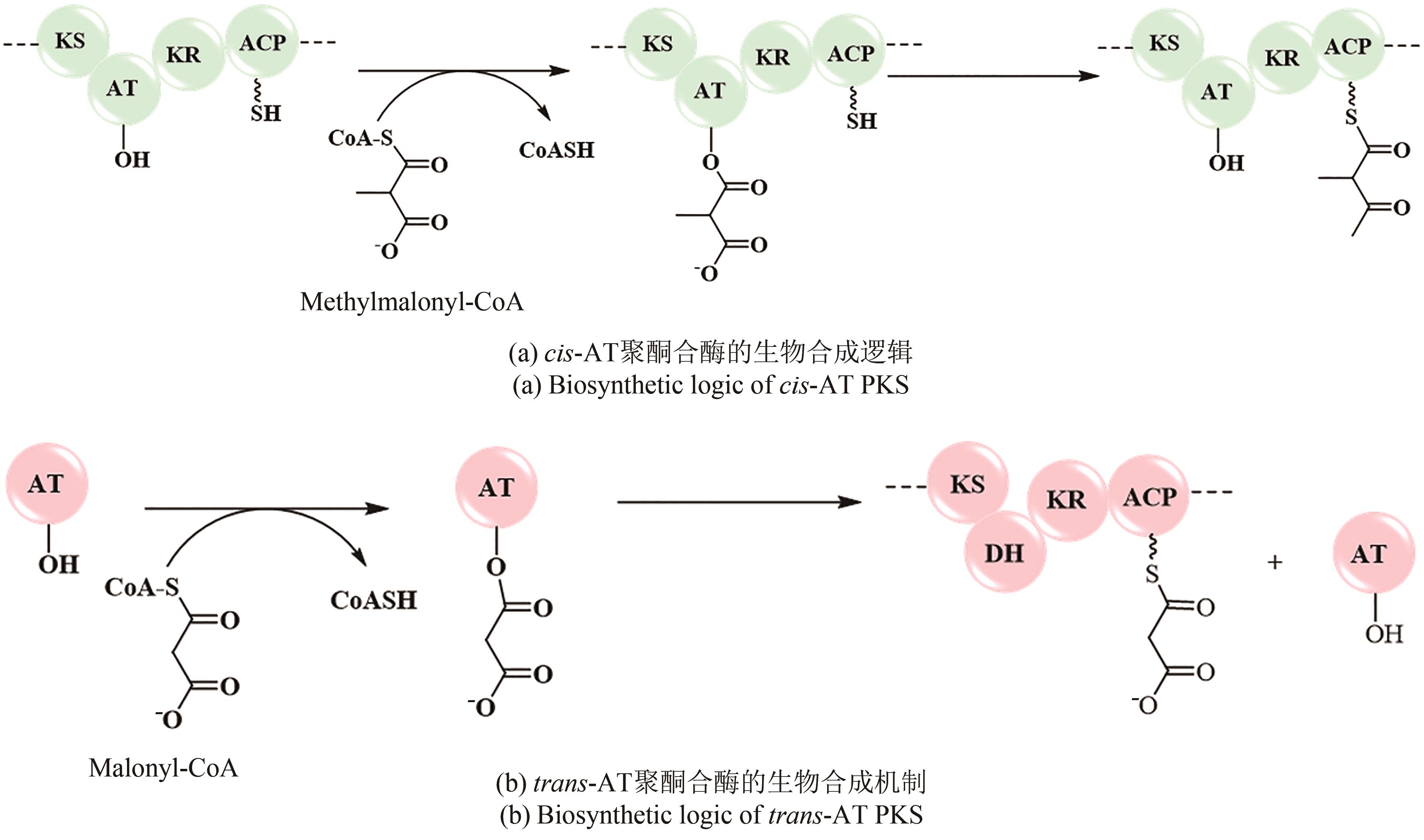
聚酮化合物(polyketide)是一类来源广泛、结构多样的活性天然产物,聚酮合酶(polyketide synthase, PKS)负责聚酮骨架的生物合成。细菌次级代谢中PKS广泛存在,不同类型的PKS在组成和生物合成机制上各不相同,从而产生截然不同的聚酮骨架。根据细菌PKS功能和生物合成途径的不同,可以将其分为Ⅰ型、Ⅱ型和Ⅲ型。PKS通常能与其他生物合成酶系杂合以产生结构更为复杂的天然产物。同时,不同类型PKS之间也可以形成多种内部杂合,产生更多样的聚酮骨架。本文总结和比较PKS间的内部杂合,包括Ⅰ型PKS内部杂合、Ⅰ型/Ⅱ型PKS杂合以及Ⅰ型/Ⅲ型PKS杂合,归纳各种杂合基因簇的形成方式及其杂合特征。通过比较杂合聚酮化合物的生物合成机制并讨论杂合聚酮工程化改造的进展,展望了多种潜在的聚酮杂合模式,合理假设存在合成过程相反的Ⅰ型/Ⅱ型PKS杂合模式,或随着化合物的挖掘发现迄今未报道的Ⅱ型/Ⅲ型PKS杂合模式等,指出可以充分和全面地利用细菌基因组信息,通过酶和基因的生物勘探,发现更多更特殊的PKS杂合化合物等一系列针对新颖聚酮化合物进行基因组挖掘的方向,同时也提出了工程化改造trans-AT PKS在cis-AT模块中实现不同寻常的骨架修饰等多种PKS的工程化改造设想,为后续PKS内部杂合基因簇挖掘和表征提供一些新思路。
中图分类号:
引用本文
张瑞, 金文铮, 陈依军. 细菌聚酮合酶间的杂合方式及聚酮化合物生物合成逻辑[J]. 合成生物学, 2024, 5(3): 548-560.
ZHANG Rui, JIN Wenzheng, CHEN Yijun. Bacterial inter-PKS hybrids and the biosynthetic algorithm of polyketides[J]. Synthetic Biology Journal, 2024, 5(3): 548-560.
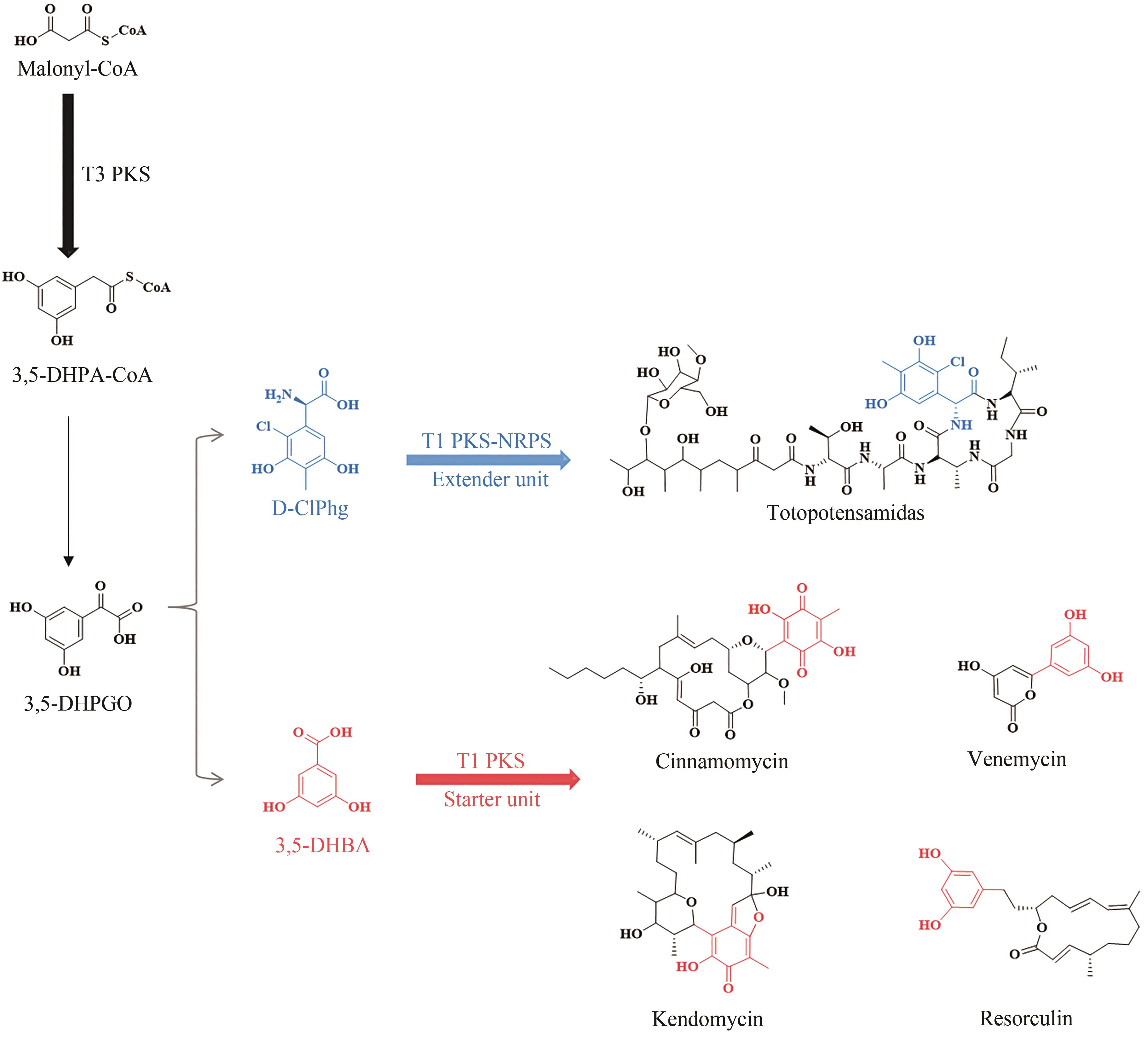
图5 Ⅱ型PKS和Ⅰ/Ⅱ型杂合PKS对应产物的化学结构(由聚酮合酶的起始单元引入的基团以红色加粗显示)
Fig. 5 Structures of products synthesized under the catalysis of type Ⅱ PKS and type Ⅰ/Ⅱ PKS hybrids(The moieties from starter units in polyketidess are highlighted in red.)
| 1 | TOOPAANG W, BUNNAK W, SRISUKSAM C, et al. Microbial polyketides and their roles in insect virulence: from genomics to biological functions[J]. Natural Product Reports, 2022, 39(11): 2008-2029. |
| 2 | HUR G H, VICKERY C R, BURKART M D. Explorations of catalytic domains in non-ribosomal peptide synthetase enzymology[J]. Natural Product Reports, 2012, 29(10): 1074-1098. |
| 3 | AVALOS M, GARBEVA P, VADER L, et al. Biosynthesis, evolution and ecology of microbial terpenoids[J]. Natural Product Reports, 2022, 39(2): 249-272. |
| 4 | MONTALBÁN-LÓPEZ M, SCOTT T A, RAMESH S, et al. New developments in RiPP discovery, enzymology and engineering[J]. Natural Product Reports, 2021, 38(1): 130-239. |
| 5 | LI S S, YANG B W, TAN G Y, et al. Polyketide pesticides from actinomycetes[J]. Current Opinion in Biotechnology, 2021, 69: 299-307. |
| 6 | GRAHAM E M. Erythromycin[J]. Obstetrics and Gynecology Clinics of North America, 1992, 19(3): 539-549. |
| 7 | HAMILL R J. Amphotericin B formulations: a comparative review of efficacy and toxicity[J]. Drugs, 2013, 73(9): 919-934. |
| 8 | POURMADADI M, GHAEMI A, SHAMSABADIPOUR A, et al. Nanoparticles loaded with Daunorubicin as an advanced tool for cancer therapy[J]. European Journal of Medicinal Chemistry, 2023, 258: 115547. |
| 9 | IKEDA H, OMURA S. Avermectin biosynthesis[J]. Chemical Reviews, 1997, 97(7): 2591-2610. |
| 10 | HOPWOOD D A. Genetic contributions to understanding polyketide synthases[J]. Chemical Reviews, 1997, 97(7): 2465-2498. |
| 11 | SHEN B. Polyketide biosynthesis beyond the type Ⅰ, Ⅱ and Ⅲ polyketide synthase paradigms[J]. Current Opinion in Chemical Biology, 2003, 7(2): 285-295. |
| 12 | KEATINGE-CLAY A T. The structures of type Ⅰ polyketide synthases[J]. Natural Product Reports, 2012, 29(10): 1050-1073. |
| 13 | HERTWECK C. The biosynthetic logic of polyketide diversity[J]. Angewandte Chemie International Edition, 2009, 48(26): 4688-4716. |
| 14 | KEATINGE-CLAY A T. Stereocontrol within polyketide assembly lines[J]. Natural Product Reports, 2016, 33(2): 141-149. |
| 15 | CHEN H T, DU L C. Iterative polyketide biosynthesis by modular polyketide synthases in bacteria[J]. Applied Microbiology and Biotechnology, 2016, 100(2): 541-557. |
| 16 | SONG L J, JENNER M, MASSCHELEIN J, et al. Discovery and biosynthesis of gladiolin: a Burkholderia gladioli antibiotic with promising activity against Mycobacterium tuberculosis [J]. Journal of the American Chemical Society, 2017, 139(23): 7974-7981. |
| 17 | DAS A, KHOSLA C. Biosynthesis of aromatic polyketides in bacteria[J]. Accounts of Chemical Research, 2009, 42(5): 631-639. |
| 18 | DREIER J, KHOSLA C. Mechanistic analysis of a type Ⅱ polyketide synthase. Role of conserved residues in the beta-ketoacyl synthase-chain length factor heterodimer[J]. Biochemistry, 2000, 39(8): 2088-2095. |
| 19 | HERTWECK C, LUZHETSKYY A, REBETS Y, et al. Type Ⅱ polyketide synthases: gaining a deeper insight into enzymatic teamwork[J]. Natural Product Reports, 2007, 24(1): 162-190. |
| 20 | XIE S L, ZHANG L H. Type Ⅱ polyketide synthases: a bioinformatics-driven approach[J]. ChemBioChem, 2023, 24(9): e202200775. |
| 21 | YU D Y, XU F C, ZENG J, et al. Type Ⅲ polyketide synthases in natural product biosynthesis[J]. IUBMB Life, 2012, 64(4): 285-295. |
| 22 | LI Y Y, MÜLLER R. Non-modular polyketide synthases in myxobacteria[J]. Phytochemistry, 2009, 70(15-16): 1850-1857. |
| 23 | MIYANAGA A, KUDO F, EGUCHI T. Protein-protein interactions in polyketide synthase-nonribosomal peptide synthetase hybrid assembly lines[J]. Natural Product Reports, 2018, 35(11): 1185-1209. |
| 24 | JENKE-KODAMA H, DITTMANN E. Bioinformatic perspectives on NRPS/PKS megasynthases: advances and challenges[J]. Natural Product Reports, 2009, 26(7): 874-883. |
| 25 | SCHWECKE T, APARICIO J F, MOLNÁR I, et al. The biosynthetic gene cluster for the polyketide immunosuppressant rapamycin[J]. Proceedings of the National Academy of Sciences of the United States of America, 1995, 92(17): 7839-7843. |
| 26 | CHEN H, O’CONNOR S, CANE D E, et al. Epothilone biosynthesis: assembly of the methylthiazolylcarboxy starter unit on the EpoB subunit[J]. Chemistry & Biology, 2001, 8(9): 899-912. |
| 27 | DU L, SÁNCHEZ C, CHEN M, et al. The biosynthetic gene cluster for the antitumor drug bleomycin from Streptomyces verticillus ATCC15003 supporting functional interactions between nonribosomal peptide synthetases and a polyketide synthase[J]. Chemistry & Biology, 2000, 7(8): 623-642. |
| 28 | KOZAKAI R, ONO T, HOSHINO S, et al. Acyltransferase that catalyses the condensation of polyketide and peptide moieties of goadvionin hybrid lipopeptides[J]. Nature Chemistry, 2020, 12(9): 869-877. |
| 29 | HUANG Y, HOEFGEN S, VALIANTE V. Biosynthesis of fungal drimane-type sesquiterpene esters[J]. Angewandte Chemie International Edition, 2021, 60(44): 23763-23770. |
| 30 | DONADIO S, STAVER M J, MCALPINE J B, et al. Modular organization of genes required for complex polyketide biosynthesis[J]. Science, 1991, 252(5006): 675-679. |
| 31 | OMURA S, IKEDA H, ISHIKAWA J, et al. Genome sequence of an industrial microorganism Streptomyces avermitilis: deducing the ability of producing secondary metabolites[J]. Proceedings of the National Academy of Sciences of the United States of America, 2001, 98(21): 12215-12220. |
| 32 | STAUNTON J, WEISSMAN K J. Polyketide biosynthesis: a millennium review[J]. Natural Product Reports, 2001, 18(4): 380-416. |
| 33 | HELFRICH E J N, REITER S, PIEL J. Recent advances in genome-based polyketide discovery[J]. Current Opinion in Biotechnology, 2014, 29: 107-115. |
| 34 | WINTER J M, BEHNKEN S, HERTWECK C. Genomics-inspired discovery of natural products[J]. Current Opinion in Chemical Biology, 2011, 15(1): 22-31. |
| 35 | NIEHS S P, KUMPFMÜLLER J, DOSE B, et al. Insect-associated bacteria assemble the antifungal butenolide gladiofungin by non-canonical polyketide chain termination[J]. Angewandte Chemie International Edition, 2020, 59(51): 23122-23126. |
| 36 | ZAZOPOULOS E, HUANG K X, STAFFA A, et al. A genomics-guided approach for discovering and expressing cryptic metabolic pathways[J]. Nature Biotechnology, 2003, 21(2): 187-190. |
| 37 | PENG H Y, ISHIDA K, HERTWECK C. Loss of single-domain function in a modular assembly line alters the size and shape of a complex polyketide[J]. Angewandte Chemie International Edition, 2019, 58(50): 18252-18256. |
| 38 | ZHANG J J, TANG X Y, HUAN T, et al. Pass-back chain extension expands multimodular assembly line biosynthesis[J]. Nature Chemical Biology, 2020, 16(1): 42-49. |
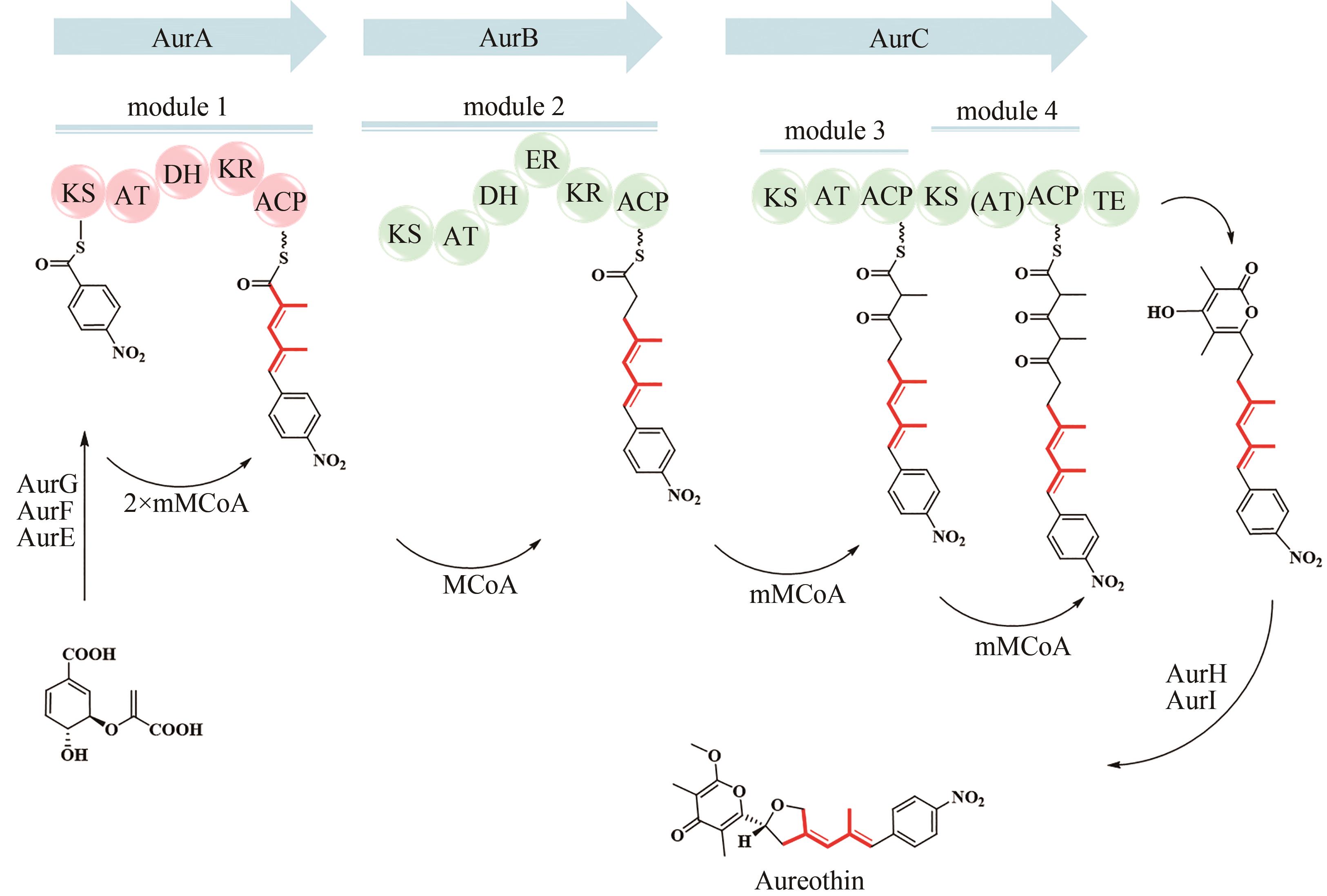
| 39 | HIRATA Y, NAKATA H, YAMADA K, et al. The structure of aureothin, a nitro compound obtained from Streptomyces thioluteus [J]. Tetrahedron, 1961, 14(3-4): 252-274. |
| 40 | HE J, HERTWECK C. Iteration as programmed event during polyketide assembly; molecular analysis of the aureothin biosynthesis gene cluster[J]. Chemistry & Biology, 2003, 10(12): 1225-1232. |
| 41 | HE J, HERTWECK C. Functional analysis of the aureothin iterative type Ⅰ polyketide synthase[J]. ChemBioChem, 2005, 6(5): 908-912. |
| 42 | BUSCH B, UEBERSCHAAR N, SUGIMOTO Y, et al. Interchenar retrotransfer of aureothin intermediates in an iterative polyketide synthase module[J]. Journal of the American Chemical Society, 2012, 134(30): 12382-12385. |
| 43 | ARAI M, HAMANO K. Isolation of three main components. F3, F4 and F5, from azalomycin F-complex[J]. The Journal of Antibiotics, 1970, 23(3): 107-112. |
| 44 | YUAN G J, LIN H P, WANG C, et al. 1H and 13C assignments of two new macrocyclic lactones isolated from Streptomyces sp. 211726 and revised assignments of azalomycins F3a, F4a and F5a [J]. Magnetic Resonance in Chemistry, 2011, 49(1): 30-37. |
| 45 | ZHAI G F, WANG W Y, XU W, et al. Cross-module enoylreduction in the Azalomycin F polyketide synthase[J]. Angewandte Chemie International Edition, 2020, 59(50): 22738-22742. |
| 46 | XU W, ZHAI G F, LIU Y Z, et al. An iterative module in the Azalomycin F polyketide synthase contains a switchable enoylreductase domain[J]. Angewandte Chemie International Edition, 2017, 56(20): 5503-5506. |
| 47 | CHEN A Y, SCHNARR N A, KIM C Y, et al. Extender unit and acyl carrier protein specificity of ketosynthase domains of the 6-deoxyerythronolide B synthase[J]. Journal of the American Chemical Society, 2006, 128(9): 3067-3074. |
| 48 | KAPUR S, CHEN A Y, CANE D E, et al. Molecular recognition between ketosynthase and acyl carrier protein domains of the 6-deoxyerythronolide B synthase[J]. Proceedings of the National Academy of Sciences of the United States of America, 2010, 107(51): 22066-22071. |
| 49 | TAN F H, PUTOCZKI T L, STYLLI S S, et al. Ponatinib: a novel multi-tyrosine kinase inhibitor against human malignancies[J]. OncoTargets and Therapy, 2019, 12: 635-645. |
| 50 | SUGIMOTO Y, ISHIDA K, TRAITCHEVA N, et al. Freedom and constraint in engineered noncolinear polyketide assembly lines[J]. Chemistry & Biology, 2015, 22(2): 229-240. |
| 51 | FISCH K M. Biosynthesis of natural products by microbial iterative hybrid PKS-NRPS[J]. RSC Advances, 2013, 3(40): 18228-18247. |
| 52 | CHENG Y Q, TANG G L, SHEN B. Type Ⅰ polyketide synthase requiring a discrete acyltransferase for polyketide biosynthesis[J]. Proceedings of the National Academy of Sciences of the United States of America, 2003, 100(6): 3149-3154. |
| 53 | DODGE G J, MALONEY F P, SMITH J L. Protein-protein interactions in “cis-AT” polyketide synthases[J]. Natural Product Reports, 2018, 35(10): 1082-1096. |
| 54 | KOSOL S, JENNER M, LEWANDOWSKI J R, et al. Protein-protein interactions in trans-AT polyketide synthases[J]. Natural Product Reports, 2018, 35(10): 1097-1109. |
| 55 | WOLF H, CHINALI G, PARMEGGIANI A. Kirromycin, an inhibitor of protein biosynthesis that acts on elongation factor Tu[J]. Proceedings of the National Academy of Sciences of the United States of America, 1974, 71(12): 4910-4914. |
| 56 | WEBER T, LAIPLE K J, PROSS E K, et al. Molecular analysis of the kirromycin biosynthetic gene cluster revealed beta-alanine as precursor of the pyridone moiety[J]. Chemistry & Biology, 2008, 15(2): 175-188. |
| 57 | ROBERTSEN H L, MUSIOL-KROLL E M, DING L, et al. Filling the gaps in the kirromycin biosynthesis: deciphering the role of genes involved in ethylmalonyl-CoA supply and tailoring reactions[J]. Scientific Reports, 2018, 8(1): 3230. |
| 58 | MUSIOL E M, HÄRTNER T, KULIK A, et al. Supramolecular templating in kirromycin biosynthesis: the acyltransferase KirCⅡ loads ethylmalonyl-CoA extender onto a specific ACP of the trans-AT PKS[J]. Chemistry & Biology, 2011, 18(4): 438-444. |
| 59 | MUSIOL E M, GREULE A, HÄRTNER T, et al. The AT2 domain of KirCⅠ loads malonyl extender units to the ACPs of the kirromycin PKS[J]. ChemBioChem, 2013, 14(11): 1343-1352. |
| 60 | JENKE-KODAMA H, BÖRNER T, DITTMANN E. Natural biocombinatorics in the polyketide synthase genes of the actinobacterium Streptomyces avermitilis [J]. PLoS Computational Biology, 2006, 2(10): 1210-1218.e132. |
| 61 | NGUYEN T, ISHIDA K, JENKE-KODAMA H, et al. Exploiting the mosaic structure of trans-acyltransferase polyketide synthases for natural product discovery and pathway dissection[J]. Nature Biotechnology, 2008, 26(2): 225-233. |
| 62 | PIEL J. Biosynthesis of polyketides by trans-AT polyketide synthases[J]. Natural Product Reports, 2010, 27(7): 996-1047. |
| 63 | YEO W L, HENG E, TAN L, et al. Biosynthetic engineering of the antifungal, anti-MRSA auroramycin[J]. Microbial Cell Factories, 2020, 19(1): 3. |
| 64 | DUNN B J, WATTS K R, ROBBINS T, et al. Comparative analysis of the substrate specificity of trans-versus cis-acyltransferases of assembly line polyketide synthases[J]. Biochemistry, 2014, 53(23): 3796-3806. |
| 65 | SIRIRUNGRUANG S, AD O, PRIVALSKY T M, et al. Engineering site-selective incorporation of fluorine into polyketides[J]. Nature Chemical Biology, 2022, 18(8): 886-893. |
| 66 | SHEN B, NAKAYAMA H, HUTCHINSON C R. Isolation and structural elucidation of tetracenomycin F2 and tetracenomycin F1: early intermediates in the biosynthesis of tetracenomycin C in Streptomyces glaucescens [J]. Journal of Natural Products, 1993, 56(8): 1288-1293. |
| 67 | KOTOWSKA M, PAWLIK K. Roles of type Ⅱ thioesterases and their application for secondary metabolite yield improvement[J]. Applied Microbiology and Biotechnology, 2014, 98(18): 7735-7746. |
| 68 | CHEN A, RE R N, BURKART M D. Type Ⅱ fatty acid and polyketide synthases: deciphering protein-protein and protein-substrate interactions[J]. Natural Product Reports, 2018, 35(10): 1029-1045. |
| 69 | DAS A, KHOSLA C. In vivo and in vitro analysis of the hedamycin polyketide synthase[J]. Chemistry & Biology, 2009, 16(11): 1197-1207. |
| 70 | HAN Y, WANG Y, YANG Y H, et al. Shellmycin A-D, novel bioactive tetrahydroanthra-γ-pyrone antibiotics from marine Streptomyces sp. Shell-016[J]. Marine Drugs, 2020, 18(1): 58. |
| 71 | BILILIGN T, HYUN C G, WILLIAMS J S, et al. The hedamycin locus implicates a novel aromatic PKS priming mechanism[J]. Chemistry & Biology, 2004, 11(7): 959-969. |
| 72 | ABE I. Biosynthesis of medicinally important plant metabolites by unusual type Ⅲ polyketide synthases[J]. Journal of Natural Medicines, 2020, 74(4): 639-646. |
| 73 | MURRAY L A M, MCKINNIE S M K, MOORE B S, et al. Meroterpenoid natural products from Streptomyces bacteria- the evolution of chemoenzymatic syntheses[J]. Natural Product Reports, 2020, 37(10): 1334-1366. |
| 74 | AUSTIN M B, NOEL J P. The chalcone synthase superfamily of type Ⅲ polyketide synthases[J]. Natural Product Reports, 2003, 20(1): 79-110. |
| 75 | CHEN H, TSENG C C, HUBBARD B K, et al. Glycopeptide antibiotic biosynthesis: enzymatic assembly of the dedicated amino acid monomer (S)-3,5-dihydroxyphenylglycine[J]. Proceedings of the National Academy of Sciences of the United States of America, 2001, 98(26): 14901-14906. |
| 76 | WENZEL S C, BODE H B, KOCHEMS I, et al. A type Ⅰ/type Ⅲ polyketide synthase hybrid biosynthetic pathway for the structurally unique ansa compound kendomycin[J]. ChemBioChem, 2008, 9(16): 2711-2721. |
| 77 | SONG R T, SHI H X, ZHU J, et al. A single-component flavoenzyme catalyzed regioselective halogenation of pyrone in the biosynthesis of venemycins[J]. ACS Chemical Biology, 2019, 14(12): 2533-2537. |
| 78 | LACEY H J, CHEN R, VUONG D, et al. Resorculins: hybrid polyketide macrolides from Streptomyces sp. MST-91080[J]. Organic & Biomolecular Chemistry, 2023, 21(12): 2531-2538. |
| 79 | ZHANG B, JIN W Z, ZHANG Y Y, et al. A type Ⅰ/type Ⅲ PKS hybrid generates cinnamomycin A-D[J]. Organic Letters, 2023, 25(15): 2560-2564. |
| 80 | CORTES J, HAYDOCK S F, ROBERTS G A, et al. An unusually large multifunctional polypeptide in the erythromycin-producing polyketide synthase of Saccharopolyspora erythraea [J]. Nature, 1990, 348(6297): 176-178. |
| 81 | GOMES E S, SCHUCH V, DE MACEDO LEMOS E G. Biotechnology of polyketides: new breath of life for the novel antibiotic genetic pathways discovery through metagenomics[J]. Brazilian Journal of Microbiology, 2013, 44(4): 1007-1034. |
| 82 | ZHANG J Y, SUN Y, WANG Y J, et al. Genome mining of novel rubiginones from Streptomyces sp. CB02414 and characterization of the post-PKS modification steps in rubiginone biosynthesis[J]. Microbial Cell Factories, 2021, 20(1): 192. |
| 83 | BAGDE S R, MATHEWS I I, FROMME J C, et al. Modular polyketide synthase contains two reaction chambers that operate asynchronously[J]. Science, 2021, 374(6568): 723-729. |
| 84 | RISDIAN C, MOZEF T, WINK J. Biosynthesis of polyketides in Streptomyces [J]. Microorganisms, 2019, 7(5): 124. |
| 85 | PRUSOV E V. Total synthesis of antibiotics: recent achievements, limitations, and perspectives[J]. Applied Microbiology and Biotechnology, 2013, 97(7): 2773-2795. |
| 86 | ZHANG S W, XIE Q, SUN C L, et al. Cytotoxic kendomycins containing the carbacylic Ansa scaffold from the marine-derived Verrucosispora sp. SCSIO 07399[J]. Journal of Natural Products, 2019, 82(12): 3366-3371. |
| [1] | 程中玉, 李付琸. 基于P450选择性氧化的天然产物化学-酶法合成进展[J]. 合成生物学, 2024, 5(5): 960-980. |
| [2] | 张守祺, 王涛, 孔尧, 邹家胜, 刘元宁, 徐正仁. 天然产物的化学-酶法合成:方法与策略的演进[J]. 合成生物学, 2024, 5(5): 913-940. |
| [3] | 谢向前, 郭雯, 王欢, 李进. 含氨基乙烯半胱氨酸核糖体肽的生物合成与化学合成[J]. 合成生物学, 2024, 5(5): 981-996. |
| [4] | 张俊, 金诗雪, 云倩, 瞿旭东. 聚酮化合物非天然延伸单元的生物合成与结构改造应用[J]. 合成生物学, 2024, 5(3): 561-570. |
| [5] | 虞旭昶, 吴辉, 李雷. 文库构建与基因簇靶向筛选驱动的微生物天然产物高效发现[J]. 合成生物学, 2024, 5(3): 492-506. |
| [6] | 冯金, 潘海学, 唐功利. 近十年天然产物药物的生物合成研究进展[J]. 合成生物学, 2024, 5(3): 408-446. |
| [7] | 奚萌宇, 胡逸灵, 顾玉诚, 戈惠明. 基因组挖掘指导天然药物分子的发现[J]. 合成生物学, 2024, 5(3): 447-473. |
| [8] | 雷茹, 陶慧, 刘天罡. 基因组深度挖掘驱动微生物萜类化合物高效发现[J]. 合成生物学, 2024, 5(3): 507-526. |
| [9] | 施鑫杰, 杜艺岭. 双嵌入家族抗肿瘤非核糖体肽的生物合成研究进展[J]. 合成生物学, 2024, 5(3): 593-611. |
| [10] | 宋永相, 张秀凤, 李艳芹, 肖华, 闫岩. 自抗性基因导向的活性天然产物挖掘[J]. 合成生物学, 2024, 5(3): 474-491. |
| [11] | 惠真, 唐啸宇. CRISPR/Cas9编辑系统在微生物天然产物研究中的应用[J]. 合成生物学, 2024, 5(3): 658-671. |
| [12] | 胡哲辉, 徐娟, 卞光凯. 自动化高通量技术在天然产物生物合成中的应用[J]. 合成生物学, 2023, 4(5): 932-946. |
| [13] | 张凡忠, 相长君, 张骊駻. 进化与大数据导向生物信息学在天然产物研究中的发展及应用[J]. 合成生物学, 2023, 4(4): 629-650. |
| [14] | 吕靖伟, 邓子新, 张琪, 丁伟. 基于深度学习识别RiPPs前体肽及裂解位点[J]. 合成生物学, 2022, 3(6): 1262-1276. |
| [15] | 董佳钰, 李敏, 肖宗华, 胡明, 松田侑大, 汪伟光. 米曲霉异源表达天然产物研究进展[J]. 合成生物学, 2022, 3(6): 1126-1149. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||