Synthetic Biology Journal ›› 2024, Vol. 5 ›› Issue (5): 1189-1210.DOI: 10.12211/2096-8280.2024-001
• Invited Review • Previous Articles Next Articles
Design, optimization and application of synthetic carbon-negative phototrophic community
ZHENG Haotian1,2, LI Chaofeng1,2, LIU Liangxu1,2, WANG Jiawei1,2, LI Hengrun1,2, NI Jun1,2
- 1.State Key Laboratory of Microbial Metabolism,School of Life Science and Technology,Shanghai Jiaotong University,Shanghai 200240,China
2.Zhangjiang Institute for Advanced Study,Shanghai Jiaotong University,Shanghai 201203,China
-
Received:2024-01-02Revised:2024-04-16Online:2024-11-20Published:2024-10-31 -
Contact:NI Jun
负碳人工光合群落的设计、优化与应用
郑皓天1,2, 李朝风1,2, 刘良叙1,2, 王嘉伟1,2, 李恒润1,2, 倪俊1,2
- 1.上海交通大学生命科学技术学院,微生物代谢国家重点实验室,上海 200240
2.上海交通大学张江高等研究院,上海 201203
-
通讯作者:倪俊 -
作者简介:郑皓天 (2000—),男,硕士研究生。研究方向为人工光合群落和光合细胞工厂的设计与优化。 E-mail:zhenghtjames@sjtu.edu.cn倪俊 (1987—),男,副教授,博士生导师。研究方向为合成生物学;人工光合群落;多酶级联组装;负碳生物合成等。 E-mail:tearroad@sjtu.edu.cn -
基金资助:国家自然科学基金(32071418);国家重点研发计划(2019YFA0904603)
CLC Number:
Cite this article
ZHENG Haotian, LI Chaofeng, LIU Liangxu, WANG Jiawei, LI Hengrun, NI Jun. Design, optimization and application of synthetic carbon-negative phototrophic community[J]. Synthetic Biology Journal, 2024, 5(5): 1189-1210.
郑皓天, 李朝风, 刘良叙, 王嘉伟, 李恒润, 倪俊. 负碳人工光合群落的设计、优化与应用[J]. 合成生物学, 2024, 5(5): 1189-1210.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2024-001
| 自养微生物 | 异养微生物 | 中间碳 | 产物 | 滴度/(mg/L) | 产率/[mg/(L·d)] | 参考文献 |
|---|---|---|---|---|---|---|
| S. elongatus PCC 7942 | S. cerevisiae | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | A. vinelandii | 蔗糖 | PHB | < 40 | < 8 | [ |
| S. elongatus PCC 7942 | H. boliviensis | 蔗糖 | PHB | 未指明 | 28.3 | [ |
| S. elongatus PCC 7942 | R. glutinis | 蔗糖 | 游离脂肪酸 | 39 | 1.2 | [ |
| S. elongatus PCC 7942 | P. putida | 蔗糖 | PHA | 156 | 9.8 | [ |
| S. elongatus PCC 7942 | S. cerevisiae | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | PHB | < 1 | < 0.15 | [ |
| S. elongatus PCC 7942 | B. subtilis | 蔗糖 | α-淀粉酶 | 未指明 | 未指明 | [ |
| S. elongatus PCC 7942 | P. putida | 蔗糖 | PHA;降解2,4-DNT | 5.1 | 5.1 | [ |
| S. elongatus PCC 7942 | P. putida | 蔗糖 | HMF转化为FDCA | 约750 | 约250 | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | Y. lipolytica | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | B. subtilis | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 异戊二烯 | 400 | 15.4 | [ |
| S. elongatus PCC 7942 | P. putida | 蔗糖 | PHA | 393 | 42.1 | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 乙烯 | 1.2 | 0.6 | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 异戊二烯 | 0.051 | 0.026 | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | V. natriegens | 蔗糖 | 乳酸 | 313.3 | 52.2 | [ |
| S. elongatus PCC 7942 | V. natriegens | 蔗糖 | 2,3-丁二醇 | 137.3 | 22.9 | [ |
| S. elongatus PCC 7942 | V. natriegens | 蔗糖 | 对香豆酸 | 24.7 | 4.1 | [ |
| S. elongatus PCC 7942 | V. natriegens | 蔗糖 | 黑色素 | 9.1 | 1.5 | [ |
| S. elongatus PCC 7942 | A. nidulans | 蔗糖 | Neosartoricin B | 0.2 | 0.05 | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 构建方法开发 | — | — | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 匹配度预测 | — | — | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 群体感应工具箱开发 | — | — | [ |
| S. elongatus UTEX 2973 | E. coli | 蔗糖 | 3-羟基丙酸 | 68.3 | 9.8 | [ |
| S. elongatus UTEX 2973 | P. putida | 蔗糖 | 天然蓝色素 | 7500 | 1250 | [ |
| S. elongatus UTEX 2973 | Y. lipolytica | 蔗糖 | β-胡萝卜素 | 1300 | 260 | [ |
| Synechococcus sp. WH7803 | R. pomeroyi | 光合产物 | 促进生长 | — | — | [ |
| Synechococcus sp. PCC 7002 | S. putrefaciens | 光合产物 | 促进生长 | — | — | [ |
| Synechococcus sp. PCC 7002 | M. alcaliphilum | 光合产物 | 甲烷降解 | — | — | [ |
| T. elongates PKUAC-SCTE542 | E. coli | 蔗糖 | 乙烯 | 1.5 | 0.74 | [ |
| T. elongates PKUAC-SCTE542 | E. coli | 蔗糖 | 异戊二烯 | 0.027 | 0.013 | [ |
| T. elongatus BP-1 | M. ruber | 光合产物 | 促进生长 | — | — | [ |
| Nostoc sp. PCC 6720 | A. nidulans | 光合产物 | 促进生长 | — | — | [ |
| Nostoc sp. PCC 7413 | A. niger | 光合产物 | 促进生长 | — | — | [ |
| Nostocaceae sp. SAB-B866 | P. cypripedii | 光合产物 | 促进植物生长 | — | — | [ |
| Nostocaceae sp. SAB-B866 | P. putida | 光合产物 | 促进植物生长 | — | — | [ |
| S. hyalinum | 废水内源微生物 | 光合产物 | 废水处理 | — | — | [ |
| P. keutzingium | 活性污泥微生物 | 光合产物 | 氢气;废水处理 | 89.9 | 14.9 | [ |
| T. obliquus IS2 | V. paradoxus | 光合产物 | 废水处理 | — | — | [ |
| Chlorella sp. GY-H4 | S. cerevisiae | 光合产物 | 苯乙醇 | 2130 | 710 | [ |
Table 1 Synthetic consortia consisting of phototroph/heterotroph pairs
| 自养微生物 | 异养微生物 | 中间碳 | 产物 | 滴度/(mg/L) | 产率/[mg/(L·d)] | 参考文献 |
|---|---|---|---|---|---|---|
| S. elongatus PCC 7942 | S. cerevisiae | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | A. vinelandii | 蔗糖 | PHB | < 40 | < 8 | [ |
| S. elongatus PCC 7942 | H. boliviensis | 蔗糖 | PHB | 未指明 | 28.3 | [ |
| S. elongatus PCC 7942 | R. glutinis | 蔗糖 | 游离脂肪酸 | 39 | 1.2 | [ |
| S. elongatus PCC 7942 | P. putida | 蔗糖 | PHA | 156 | 9.8 | [ |
| S. elongatus PCC 7942 | S. cerevisiae | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | PHB | < 1 | < 0.15 | [ |
| S. elongatus PCC 7942 | B. subtilis | 蔗糖 | α-淀粉酶 | 未指明 | 未指明 | [ |
| S. elongatus PCC 7942 | P. putida | 蔗糖 | PHA;降解2,4-DNT | 5.1 | 5.1 | [ |
| S. elongatus PCC 7942 | P. putida | 蔗糖 | HMF转化为FDCA | 约750 | 约250 | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | Y. lipolytica | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | B. subtilis | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 异戊二烯 | 400 | 15.4 | [ |
| S. elongatus PCC 7942 | P. putida | 蔗糖 | PHA | 393 | 42.1 | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 乙烯 | 1.2 | 0.6 | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 异戊二烯 | 0.051 | 0.026 | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 促进生长 | — | — | [ |
| S. elongatus PCC 7942 | V. natriegens | 蔗糖 | 乳酸 | 313.3 | 52.2 | [ |
| S. elongatus PCC 7942 | V. natriegens | 蔗糖 | 2,3-丁二醇 | 137.3 | 22.9 | [ |
| S. elongatus PCC 7942 | V. natriegens | 蔗糖 | 对香豆酸 | 24.7 | 4.1 | [ |
| S. elongatus PCC 7942 | V. natriegens | 蔗糖 | 黑色素 | 9.1 | 1.5 | [ |
| S. elongatus PCC 7942 | A. nidulans | 蔗糖 | Neosartoricin B | 0.2 | 0.05 | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 构建方法开发 | — | — | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 匹配度预测 | — | — | [ |
| S. elongatus PCC 7942 | E. coli | 蔗糖 | 群体感应工具箱开发 | — | — | [ |
| S. elongatus UTEX 2973 | E. coli | 蔗糖 | 3-羟基丙酸 | 68.3 | 9.8 | [ |
| S. elongatus UTEX 2973 | P. putida | 蔗糖 | 天然蓝色素 | 7500 | 1250 | [ |
| S. elongatus UTEX 2973 | Y. lipolytica | 蔗糖 | β-胡萝卜素 | 1300 | 260 | [ |
| Synechococcus sp. WH7803 | R. pomeroyi | 光合产物 | 促进生长 | — | — | [ |
| Synechococcus sp. PCC 7002 | S. putrefaciens | 光合产物 | 促进生长 | — | — | [ |
| Synechococcus sp. PCC 7002 | M. alcaliphilum | 光合产物 | 甲烷降解 | — | — | [ |
| T. elongates PKUAC-SCTE542 | E. coli | 蔗糖 | 乙烯 | 1.5 | 0.74 | [ |
| T. elongates PKUAC-SCTE542 | E. coli | 蔗糖 | 异戊二烯 | 0.027 | 0.013 | [ |
| T. elongatus BP-1 | M. ruber | 光合产物 | 促进生长 | — | — | [ |
| Nostoc sp. PCC 6720 | A. nidulans | 光合产物 | 促进生长 | — | — | [ |
| Nostoc sp. PCC 7413 | A. niger | 光合产物 | 促进生长 | — | — | [ |
| Nostocaceae sp. SAB-B866 | P. cypripedii | 光合产物 | 促进植物生长 | — | — | [ |
| Nostocaceae sp. SAB-B866 | P. putida | 光合产物 | 促进植物生长 | — | — | [ |
| S. hyalinum | 废水内源微生物 | 光合产物 | 废水处理 | — | — | [ |
| P. keutzingium | 活性污泥微生物 | 光合产物 | 氢气;废水处理 | 89.9 | 14.9 | [ |
| T. obliquus IS2 | V. paradoxus | 光合产物 | 废水处理 | — | — | [ |
| Chlorella sp. GY-H4 | S. cerevisiae | 光合产物 | 苯乙醇 | 2130 | 710 | [ |
| 1 | CESTELLOS-BLANCO S, ZHANG H, KIM J M, et al. Photosynthetic semiconductor biohybrids for solar-driven biocatalysis[J]. Nature Catalysis, 2020, 3: 245-255. |
| 2 | LIAO J C, MI L, PONTRELLI S, et al. Fuelling the future: microbial engineering for the production of sustainable biofuels[J]. Nature Reviews Microbiology, 2016, 14(5): 288-304. |
| 3 | LIU Z H, WANG K, CHEN Y, et al. Third-generation biorefineries as the means to produce fuels and chemicals from CO2 [J]. Nature Catalysis, 2020, 3: 274-288. |
| 4 | CASE A E, ATSUMI S. Cyanobacterial chemical production[J]. Journal of Biotechnology, 2016, 231: 106-114. |
| 5 | GLEIZER S, BEN-NISSAN R, BAR-ON Y M, et al. Conversion of Escherichia coli to generate all biomass carbon from CO2 [J]. Cell, 2019, 179(6): 1255-1263.e12. |
| 6 | VENKATA MOHAN S, MODESTRA J A, AMULYA K, et al. A circular bioeconomy with biobased products from CO2 sequestration[J]. Trends in Biotechnology, 2016, 34(6): 506-519. |
| 7 | KASHYAP M, CHAKRABORTY S, KUMARI A, et al. Strategies and challenges to enhance commercial viability of algal biorefineries for biofuel production[J]. Bioresource Technology, 2023, 387: 129551. |
| 8 | AHIRWAR A, DAS S, DAS S, et al. Photosynthetic microbial fuel cell for bioenergy and valuable production: a review of circular bio-economy approach[J]. Algal Research, 2023, 70: 102973. |
| 9 | ERDEM E, MALIHAN-YAP L, ASSIL-COMPANIONI L, et al. Photobiocatalytic oxyfunctionalization with high reaction rate using a Baeyer-Villiger monooxygenase from Burkholderia xenovorans in metabolically engineered cyanobacteria[J]. ACS Catalysis, 2022, 12(1): 66-72. |
| 10 | MASCIA F, PEREIRA S B, PACHECO C C, et al. Light-driven hydroxylation of testosterone by Synechocystis sp. PCC 6803 expressing the heterologous CYP450 monooxygenase CYP110D1[J]. Green Chemistry, 2022, 24(16): 6156-6167. |
| 11 | TAN C L, TAO F, XU P. Direct carbon capture for the production of high-performance biodegradable plastics by cyanobacterial cell factories[J]. Green Chemistry, 2022, 24(11): 4470-4483. |
| 12 | WANG Q K, YU Z Y, WEI D, et al. Mixotrophic Chlorella pyrenoidosa as cell factory for ultrahigh-efficient removal of ammonium from catalyzer wastewater with valuable algal biomass coproduction through short-time acclimation[J]. Bioresource Technology, 2021, 333: 125151. |
| 13 | ZHU B J, WEI D, POHNERT G. The thermoacidophilic red alga Galdieria sulphuraria is a highly efficient cell factory for ammonium recovery from ultrahigh-NH4+ industrial effluent with co-production of high-protein biomass by photo-fermentation[J]. Chemical Engineering Journal, 2022, 438: 135598. |
| 14 | BAI W, RANAIVOARISOA T O, SINGH R, et al. n-Butanol production by Rhodopseudomonas palustris TIE-1[J]. Communications Biology, 2021, 4(1): 1257. |
| 15 | LI M J, XIA Q Q, LV S Z, et al. Enhanced CO2 capture for photosynthetic lycopene production in engineered Rhodopseudomonas palustris, a purple nonsulfur bacterium[J]. Green Chemistry, 2022, 24(19): 7500-7518. |
| 16 | DEMAY J, BERNARD C, REINHARDT A, et al. Natural products from cyanobacteria: focus on beneficial activities[J]. Marine Drugs, 2019, 17(6): 320. |
| 17 | TAN C L, XU P, TAO F. Carbon-negative synthetic biology: challenges and emerging trends of cyanobacterial technology[J]. Trends in Biotechnology, 2022, 40(12): 1488-1502. |
| 18 | KNOOT C J, UNGERER J, WANGIKAR P P, et al. Cyanobacteria: promising biocatalysts for sustainable chemical production[J]. Journal of Biological Chemistry, 2018, 293(14): 5044-5052. |
| 19 | OPEL F, SIEBERT N A, KLATT S, et al. Generation of synthetic shuttle vectors enabling modular genetic engineering of cyanobacteria[J]. ACS Synthetic Biology, 2022, 11(5): 1758-1771. |
| 20 | DERUYCK B, NGUYEN K H THI, DECAESTECKER E, et al. Modeling the impact of rotifer contamination on microalgal production in open pond, photobioreactor and thin layer cultivation systems[J]. Algal Research, 2019, 38: 101398. |
| 21 | STUART R K, MAYALI X, LEE J Z, et al. Cyanobacterial reuse of extracellular organic carbon in microbial mats[J]. The ISME Journal, 2016, 10(5): 1240-1251. |
| 22 | SENEVIRATNE G, INDRASENA I K. Nitrogen fixation in lichens is important for improved rock weathering[J]. Journal of Biosciences, 2006, 31(5): 639-643. |
| 23 | ZUÑIGA C, LI C T, YU G, et al. Environmental stimuli drive a transition from cooperation to competition in synthetic phototrophic communities[J]. Nature Microbiology, 2019, 4(12): 2184-2191. |
| 24 | KRATZL F, KREMLING A, PFLÜGER-GRAU K. Streamlining of a synthetic co-culture towards an individually controllable one-pot process for polyhydroxyalkanoate production from light and CO2 [J]. Engineering in Life Sciences, 2022, 23(1): e2100156. |
| 25 | WEISS T L, YOUNG E J, DUCAT D C. A synthetic, light-driven consortium of cyanobacteria and heterotrophic bacteria enables stable polyhydroxybutyrate production[J]. Metabolic Engineering, 2017, 44: 236-245. |
| 26 | ZHANG L, CHEN L, DIAO J J, et al. Construction and analysis of an artificial consortium based on the fast-growing cyanobacterium Synechococcus elongatus UTEX 2973 to produce the platform chemical 3-hydroxypropionic acid from CO2 [J]. Biotechnology for Biofuels, 2020, 13: 82. |
| 27 | WANG L, ZHANG X, TANG C W, et al. Engineering consortia by polymeric microbial swarmbots[J]. Nature Communications, 2022, 13(1): 3879. |
| 28 | LI C F, WANG R Y, WANG J W, et al. A highly compatible phototrophic community for carbon-negative biosynthesis[J]. Angewandte Chemie International Edition, 2023, 62(2): e202215013. |
| 29 | ZHAO R Y, SENGUPTA A, TAN A X, et al. Photobiological production of high-value pigments via compartmentalized co-cultures using Ca-alginate hydrogels[J]. Scientific Reports, 2022, 12(1): 22163. |
| 30 | SHU W S, HUANG L N. Microbial diversity in extreme environments[J]. Nature Reviews Microbiology, 2022, 20(4): 219-235. |
| 31 | SCHIPPERS A, NERETIN L N, KALLMEYER J, et al. Prokaryotic cells of the deep sub-seafloor biosphere identified as living bacteria[J]. Nature, 2005, 433(7028): 861-864. |
| 32 | CASTENHOLZ R W. The effect of sulfide on the blue-green algae of hot springsⅡ. Yellowstone National Park[J]. Microbial Ecology, 1977, 3(2): 79-105. |
| 33 | THOMAS D N, DIECKMANN G S. Antarctic Sea ice: a habitat for extremophiles[J]. Science, 2002, 295(5555): 641-644. |
| 34 | TANG Y Z, KOCH F, GOBLER C J. Most harmful algal bloom species are vitamin B1 and B12 auxotrophs[J]. Proceedings of the National Academy of Sciences of the United States of America, 2010, 107(48): 20756-20761. |
| 35 | XIONG Q, HU L X, LIU Y S, et al. Microalgae-based technology for antibiotics removal: from mechanisms to application of innovational hybrid systems[J]. Environment International, 2021, 155: 106594. |
| 36 | KUMARI M, GHOSH P, SWATI, et al. Development of artificial consortia of microalgae and bacteria for efficient biodegradation and detoxification of lindane[J]. Bioresource Technology Reports, 2020, 10: 100415. |
| 37 | MOHSENPOUR S F, HENNIGE S, WILLOUGHBY N, et al. Integrating micro-algae into wastewater treatment: a review[J]. Science of the Total Environment, 2021, 752: 142168. |
| 38 | SANTOS C A, REIS A. Microalgal symbiosis in biotechnology[J]. Applied Microbiology and Biotechnology, 2014, 98(13): 5839-5846. |
| 39 | PARK Y, JE K W, LEE K, et al. Growth promotion of Chlorella ellipsoidea by co-inoculation with Brevundimonas sp. isolated from the microalga[J]. Hydrobiologia, 2008, 598(1): 219-228. |
| 40 | BUCHAN A, LECLEIR G R, GULVIK C A, et al. Master recyclers: features and functions of bacteria associated with phytoplankton blooms[J]. Nature Reviews Microbiology, 2014, 12(10): 686-698. |
| 41 | JAMES C C, BARTON A D, ALLEN L Z, et al. Influence of nutrient supply on plankton microbiome biodiversity and distribution in a coastal upwelling region[J]. Nature Communications, 2022, 13(1): 2448. |
| 42 | SEYMOUR J R, AMIN S A, RAINA J B, et al. Zooming in on the phycosphere: the ecological interface for phytoplankton-bacteria relationships[J]. Nature Microbiology, 2017, 2: 17065. |
| 43 | MISHRA A, KAVITA K, JHA B. Characterization of extracellular polymeric substances produced by micro-algae Dunaliella salina [J]. Carbohydrate Polymers, 2011, 83(2): 852-857. |
| 44 | VAN OOSTENDE N, MOERDIJK-POORTVLIET T C W, BOSCHKER H T S, et al. Release of dissolved carbohydrates by Emiliania huxleyi and formation of transparent exopolymer particles depend on algal life cycle and bacterial activity[J]. Environmental Microbiology, 2013, 15(5): 1514-1531. |
| 45 | WICKER R J, DANESHVAR E, KUMAR GUPTA A, et al. Hybrid planktonic-biofilm cultivation of a Nordic mixed-species photosynthetic consortium: a pilot study on carbon capture and nutrient removal[J]. Chemical Engineering Journal, 2023, 471: 144585. |
| 46 | COLE J J. Interactions between bacteria and algae in aquatic ecosystems[J]. Annual Review of Ecology and Systematics, 1982, 13: 291-314. |
| 47 | ŽITNIK M, ŠUNTA U, GODIČ TORKAR K, et al. The study of interactions and removal efficiency of Escherichia coli in raw blackwater treated by microalgae Chlorella vulgaris [J]. Journal of Cleaner Production, 2019, 238: 117865. |
| 48 | FOSTER R A, KUYPERS M M M, VAGNER T, et al. Nitrogen fixation and transfer in open ocean diatom-cyanobacterial symbioses[J]. The ISME Journal, 2011, 5(9): 1484-1493. |
| 49 | THOMPSON A W, FOSTER R A, KRUPKE A, et al. Unicellular cyanobacterium symbiotic with a single-celled eukaryotic alga[J]. Science, 2012, 337(6101): 1546-1550. |
| 50 | SYLVAN J B, DORTCH Q, NELSON D M, et al. Phosphorus limits phytoplankton growth on the Louisiana shelf during the period of hypoxia formation[J]. Environmental Science & Technology, 2006, 40(24): 7548-7553. |
| 51 | CARRIÓN O, LI C Y, PENG M, et al. DMSOP-cleaving enzymes are diverse and widely distributed in marine microorganisms[J]. Nature Microbiology, 2023, 8(12): 2326-2337. |
| 52 | BRINKHOFF T, BACH G, HEIDORN T, et al. Antibiotic production by a Roseobacter clade-affiliated species from the German Wadden Sea and its antagonistic effects on indigenous isolates[J]. Applied and Environmental Microbiology, 2004, 70(4): 2560-2565. |
| 53 | SEYEDSAYAMDOST M R, CARR G, KOLTER R, et al. Roseobacticides: small molecule modulators of an algal-bacterial symbiosis[J]. Journal of the American Chemical Society, 2011, 133(45): 18343-18349. |
| 54 | LIU H L, ZHOU Y Y, XIAO W J, et al. Shifting nutrient-mediated interactions between algae and bacteria in a microcosm: evidence from alkaline phosphatase assay[J]. Microbiological Research, 2012, 167(5): 292-298. |
| 55 | GURUNG T B, URABE J, NAKANISHI M. Regulation of the relationship between phytoplankton Scenedesmus acutus and heterotrophic bacteria by the balance of light and nutrients[J]. Aquatic Microbial Ecology, 1999, 17: 27-35. |
| 56 | LIU L, HALL G, CHAMPAGNE P. Effects of environmental factors on the disinfection performance of a wastewater stabilization pond operated in a temperate climate[J]. Water, 2015, 8(1): 5. |
| 57 | VASKER B, BEN-ZION M, KINEL-TAHAN Y, et al. Computerized optimization of microalgal photosynthesis and growth[J]. Applied Phycology, 2021, 2(1): 22-30. |
| 58 | CHO K H, WOLNY J, KASE J A, et al. Interactions of E. coli with algae and aquatic vegetation in natural waters[J]. Water Research, 2022, 209: 117952. |
| 59 | CROFT M T, LAWRENCE A D, RAUX-DEERY E, et al. Algae acquire vitamin B12 through a symbiotic relationship with bacteria[J]. Nature, 2005, 438(7064): 90-93. |
| 60 | AMIN S A, GREEN D H, HART M C, et al. Photolysis of iron-siderophore chelates promotes bacterial-algal mutualism[J]. Proceedings of the National Academy of Sciences of the United States of America, 2009, 106(40): 17071-17076. |
| 61 | DURHAM B P, SHARMA S, LUO H W, et al. Cryptic carbon and sulfur cycling between surface ocean plankton[J]. Proceedings of the National Academy of Sciences of the United States of America, 2015, 112(2): 453-457. |
| 62 | PAERL R W, BOUGET F Y, LOZANO J C, et al. Use of plankton-derived vitamin B1 precursors, especially thiazole-related precursor, by key marine picoeukaryotic phytoplankton[J]. The ISME Journal, 2017, 11(3): 753-765. |
| 63 | TORTELL P D, MALDONADO M T, GRANGER J, et al. Marine bacteria and biogeochemical cycling of iron in the oceans[J]. FEMS Microbiology Ecology, 1999, 29(1): 1-11. |
| 64 | YAO S, LYU S, AN Y, et al. Microalgae-bacteria symbiosis in microalgal growth and biofuel production: a review[J]. Journal of Applied Microbiology, 2019, 126(2): 359-368. |
| 65 | HOPKINSON B M, MOREL F M M. The role of siderophores in iron acquisition by photosynthetic marine microorganisms[J]. BioMetals, 2009, 22(4): 659-669. |
| 66 | RAINA J B, LAMBERT B S, PARKS D H, et al. Chemotaxis shapes the microscale organization of the ocean’s microbiome[J]. Nature, 2022, 605(7908): 132-138. |
| 67 | CHEN S S, CHEN J, ZHANG L L, et al. Biophotoelectrochemical process co-driven by dead microalgae and live bacteria[J]. The ISME Journal, 2023, 17(5): 712-719. |
| 68 | MUKHERJEE S, BASSLER B L. Bacterial quorum sensing in complex and dynamically changing environments[J]. Nature Reviews Microbiology, 2019, 17(6): 371-382. |
| 69 | FEDERLE M J. Autoinducer-2-based chemical communication in bacteria: complexities of interspecies signaling[M/OL]// COLLIN M, SCHUCH R. Contributions to microbiology: bacterial sensing and signaling. Basel: Karger, 2009, 16: 18-32 [2023-12-01]. . |
| 70 | ZHOU J, LYU Y H, RICHLEN M, et al. Quorum sensing is a language of chemical signals and plays an ecological role in algal-bacterial interactions[J]. Critical Reviews in Plant Sciences, 2016, 35(2): 81-105. |
| 71 | JI X Y, JIANG M Q, ZHANG J B, et al. The interactions of algae-bacteria symbiotic system and its effects on nutrients removal from synthetic wastewater[J]. Bioresource Technology, 2018, 247: 44-50. |
| 72 | ZHOU D D, ZHANG C F, FU L, et al. Responses of the microalga Chlorophyta sp. to bacterial quorum sensing molecules (N-acylhomoserine lactones): aromatic protein-induced self-aggregation[J]. Environmental Science & Technology, 2017, 51(6): 3490-3498. |
| 73 | PAPENFORT K, BASSLER B L. Quorum sensing signal-response systems in Gram-negative bacteria[J]. Nature Reviews Microbiology, 2016, 14(9): 576-588. |
| 74 | CHOI H, MASCUCH S J, VILLA F A, et al. Honaucins A-C, potent inhibitors of inflammation and bacterial quorum sensing: synthetic derivatives and structure-activity relationships[J]. Chemistry & Biology, 2012, 19(5): 589-598. |
| 75 | BORGES A, SIMÕES M. Quorum sensing inhibition by marine bacteria[J]. Marine Drugs, 2019, 17(7): 427. |
| 76 | ZHANG B, LI W, GUO Y, et al. Microalgal-bacterial consortia: from interspecies interactions to biotechnological applications[J]. Renewable and Sustainable Energy Reviews, 2020, 118: 109563. |
| 77 | KANAGASABHAPATHY M, YAMAZAKI G, ISHIDA A, et al. Presence of quorum-sensing inhibitor-like compounds from bacteria isolated from the brown alga Colpomenia sinuosa [J]. Letters in Applied Microbiology, 2009, 49(5): 573-579. |
| 78 | KOKARAKIS E J, RILLEMA R, DUCAT D C, et al. Developing cyanobacterial quorum sensing toolkits: toward interspecies coordination in mixed autotroph/heterotroph communities[J]. ACS Synthetic Biology, 2023, 12(1): 265-276. |
| 79 | RIQUELME C E, ISHIDA Y. Chemotaxis of bacteria to extracellular products of marine bloom algae[J]. The Journal of General and Applied Microbiology, 1988, 34(5): 417-423. |
| 80 | LI X J, CAI F S, LUAN T G, et al. Pyrene metabolites by bacterium enhancing cell division of green alga Selenastrum capricornutum [J]. Science of the Total Environment, 2019, 689: 287-294. |
| 81 | ALLEN A E, DUPONT C L, OBORNÍK M, et al. Evolution and metabolic significance of the urea cycle in photosynthetic diatoms[J]. Nature, 2011, 473(7346): 203-207. |
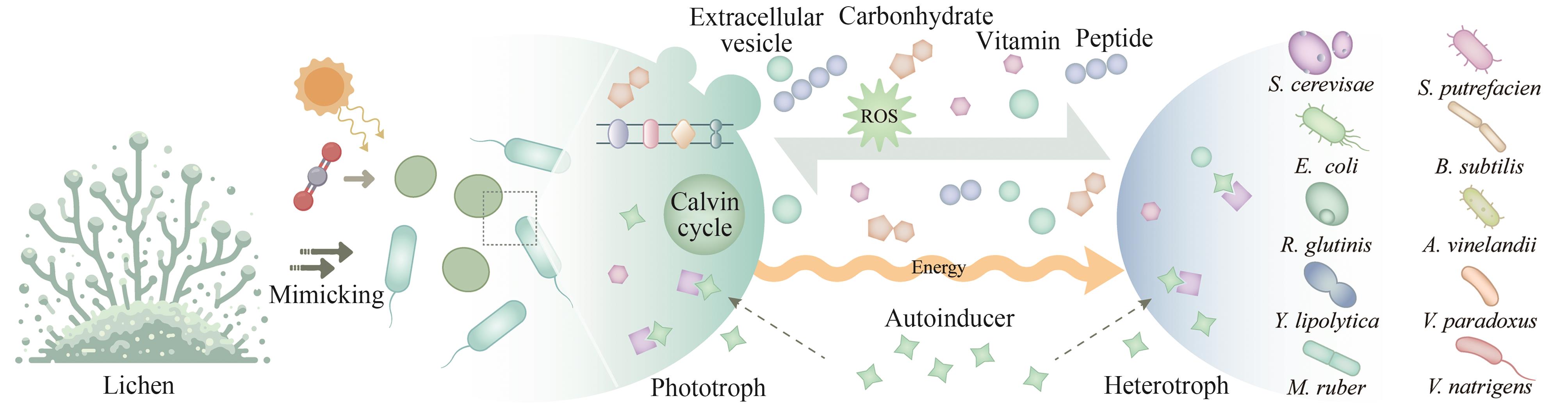
| 82 | LI T T, LI C T, BUTLER K, et al. Mimicking lichens: incorporation of yeast strains together with sucrose-secreting cyanobacteria improves survival, growth, ROS removal, and lipid production in a stable mutualistic co-culture production platform[J]. Biotechnology for Biofuels, 2017, 10: 55. |
| 83 | FEDESON D T, SAAKE P, CALERO P, et al. Biotransformation of 2,4-dinitrotoluene in a phototrophic co-culture of engineered Synechococcus elongatus and Pseudomonas putida [J]. Microbial Biotechnology, 2020, 13(4): 997-1011. |
| 84 | CUI Y X, RASUL F, JIANG Y, et al. Construction of an artificial consortium of Escherichia coli and cyanobacteria for clean indirect production of volatile platform hydrocarbons from CO2 [J]. Frontiers in Microbiology, 2022, 13: 965968. |
| 85 | KRATZL F, URBAN M, PANDHAL J, et al. Pseudomonas putida as saviour for troubled Synechococcus elongatus in a synthetic co-culture-interaction studies based on a multi-OMICs approach[J]. Communications Biology, 2024, 7: 452. |
| 86 | 国陶红, 宋馨宇, 陈磊, 等. 人工微生物混菌系统机制解析中的组学应用及进展[J]. 生物工程学报, 2022, 38(2): 460-477. |
| GUO T H, SONG X Y, CHEN L, et al. Using OMICS technologies to analyze the mechanisms of synthetic microbial co-culture systems: a review[J]. Chinese Journal of Biotechnology, 2022, 38(2): 460-477. | |
| 87 | LIU H, CAO Y J, GUO J, et al. Study on the isoprene-producing co-culture system of Synechococcus elongates-Escherichia coli through omics analysis[J]. Microbial Cell Factories, 2021, 20(1): 6. |
| 88 | NAIR S, ZHANG Z H, LI H M, et al. Inherent tendency of Synechococcus and heterotrophic bacteria for mutualism on long-term coexistence despite environmental interference[J]. Science Advances, 2022, 8(39): eabf4792. |
| 89 | ZHANG Z H, NAIR S, TANG L L, et al. Long-term survival of Synechococcus and heterotrophic bacteria without external nutrient supply after changes in their relationship from antagonism to mutualism[J]. mBio, 2021, 12(4): e0161421. |
| 90 | MA J J, GUO T H, REN M J, et al. Cross-feeding between cyanobacterium Synechococcus and Escherichia coli in an artificial autotrophic-heterotrophic coculture system revealed by integrated omics analysis[J]. Biotechnology for Biofuels and Bioproducts, 2022, 15(1): 69. |
| 91 | LIU H, XIAN M, CAO Y J, et al. Omics integration for in-depth understanding of the low-carbon co-culture platform system of Chlorella vulgaris-Escherichia coli [J]. Algal Research, 2023, 75: 103252. |
| 92 | BECKER S A, FEIST A M, MO M L, et al. Quantitative prediction of cellular metabolism with constraint-based models: the COBRA Toolbox[J]. Nature Protocols, 2007, 2(3): 727-738. |
| 93 | KUMAR M, JI B Y, ZENGLER K, et al. Modelling approaches for studying the microbiome[J]. Nature Microbiology, 2019, 4(8): 1253-1267. |
| 94 | ANTONAKOUDIS A, BARBOSA R, KOTIDIS P, et al. The era of big data: genome-scale modelling meets machine learning[J]. Computational and Structural Biotechnology Journal, 2020, 18: 3287-3300. |
| 95 | CHIU H C, LEVY R, BORENSTEIN E. Emergent biosynthetic capacity in simple microbial communities[J]. PLoS Computational Biology, 2014, 10(7): e1003695. |
| 96 | ZUÑIGA C, LI T T, GUARNIERI M T, et al. Synthetic microbial communities of heterotrophs and phototrophs facilitate sustainable growth[J]. Nature Communications, 2020, 11(1): 3803. |
| 97 | LLOYD C J, KING Z A, SANDBERG T E, et al. The genetic basis for adaptation of model-designed syntrophic co-cultures[J]. PLoS Computational Biology, 2019, 15(3): e1006213. |
| 98 | GARCÍA-JIMÉNEZ B, GARCÍA J L, NOGALES J. FLYCOP: metabolic modeling-based analysis and engineering microbial communities[J]. Bioinformatics, 2018, 34(17): i954-i963. |
| 99 | THOMMES M, WANG T Y, ZHAO Q, et al. Designing metabolic division of labor in microbial communities[J]. mSystems, 2019, 4(2): e00263-18. |
| 100 | KARKARIA B D, FEDOREC A J H, BARNES C P. Automated design of synthetic microbial communities[J]. Nature Communications, 2021, 12(1): 672. |
| 101 | SAKKOS J K, SANTOS-MERINO M, KOKARAKIS E J, et al. Predicting partner fitness based on spatial structuring in a light-driven microbial community[J]. PLoS Computational Biology, 2023, 19(5): e1011045. |
| 102 | DUCAT D C, AVELAR-RIVAS J A, WAY J C, et al. Rerouting carbon flux to enhance photosynthetic productivity[J]. Applied and Environmental Microbiology, 2012, 78(8): 2660-2668. |
| 103 | SMITH M J, FRANCIS M B. A designed A. vinelandii-S. elongatus coculture for chemical photoproduction from air, water, phosphate, and trace metals[J]. ACS Synthetic Biology, 2016, 5(9): 955-961. |
| 104 | LÖWE H, HOBMEIER K, MOOS M, et al. Photoautotrophic production of polyhydroxyalkanoates in a synthetic mixed culture of Synechococcus elongatus cscB and Pseudomonas putida cscAB [J]. Biotechnology for Biofuels, 2017, 10: 190. |
| 105 | HAYS S G, YAN L L W, SILVER P A, et al. Synthetic photosynthetic consortia define interactions leading to robustness and photoproduction[J]. Journal of Biological Engineering, 2017, 11: 4. |
| 106 | LIN T Y, WEN R C, SHEN C R, et al. Biotransformation of 5-hydroxymethylfurfural to 2,5-furandicarboxylic acid by a syntrophic consortium of engineered Synechococcus elongatus and Pseudomonas putida [J]. Biotechnology Journal, 2020, 15(6): e1900357. |
| 107 | FENG J, LI J W, LIU D X, et al. Generation and comprehensive analysis of Synechococcus elongatus-Aspergillus nidulans co-culture system for polyketide production[J]. Biotechnology for Biofuels and Bioproducts, 2023, 16(1): 32. |
| 108 | SINGH A K, DUCAT D C. Generation of stable, light-driven co-cultures of cyanobacteria with heterotrophic microbes[M/OL]//ZURBRIGGEN M D. Methods in molecular biology: plant synthetic biology. New York: Humana, 2022, 2379: 277-291 [2023-12-01]. . |
| 109 | CHRISTIE-OLEZA J A, SOUSONI D, LLOYD M, et al. Nutrient recycling facilitates long-term stability of marine microbial phototroph-heterotroph interactions[J]. Nature Microbiology, 2017, 2: 17100. |
| 110 | BELIAEV A S, ROMINE M F, SERRES M, et al. Inference of interactions in cyanobacterial-heterotrophic co-cultures via transcriptome sequencing[J]. The ISME Journal, 2014, 8(11): 2243-2255. |
| 111 | HILL E A, CHRISLER W B, BELIAEV A S, et al. A flexible microbial co-culture platform for simultaneous utilization of methane and carbon dioxide from gas feedstocks[J]. Bioresource Technology, 2017, 228: 250-256. |
| 112 | BERNSTEIN H C, MCCLURE R S, THIEL V, et al. Indirect interspecies regulation: transcriptional and physiological responses of a Cyanobacterium to heterotrophic partnership[J]. mSystems, 2017, 2(2): e00181-16. |
| 113 | JIANG L Q, LI T T, JENKINS J, et al. Evidence for a mutualistic relationship between the cyanobacteria Nostoc and fungi Aspergilli in different environments[J]. Applied Microbiology and Biotechnology, 2020, 104(14): 6413-6426. |
| 114 | LI T T, JIANG L Q, HU Y F, et al. Creating a synthetic lichen: mutualistic co-culture of fungi and extracellular polysaccharide-secreting cyanobacterium Nostoc PCC 7413[J]. Algal Research, 2020, 45: 101755. |
| 115 | TORIBIO A J, SUÁREZ-ESTRELLA F, JURADO M M, et al. Design and validation of cyanobacteria-rhizobacteria consortia for tomato seedlings growth promotion[J]. Scientific Reports, 2022, 12(1): 13150. |
| 116 | WU L, QUAN L H, DENG Z K, et al. Performance of a biocrust cyanobacteria-indigenous bacteria (BCIB) co-culture system for nutrient capture and transfer in municipal wastewater[J]. The Science of the Total Environment, 2023, 888: 164236. |
| 117 | MOHAMMED AL NUAIMI M, JAVED M A, EL-TARABILY K A, et al. Biohydrogen production of a halophytic cyanobacteria Phormidium keutzingium and activated sludge co-culture using different carbon substrates and saline concentrations[J]. Energy Conversion and Management: X, 2023, 20: 100487. |
| 118 | PERERA I A, ABINANDAN S, PANNEERSELVAN L, et al. Co-culturing of microalgae and bacteria in real wastewaters alters indigenous bacterial communities enhancing effluent bioremediation[J]. Algal Research, 2022, 64: 102705. |
| 119 | GAO H, WANG H X, ZHANG Y Q, et al. Design and optimization of artificial light-driven microbial consortia for the sustainable growth and biosynthesis of 2-phenylethanol[J]. Chemical Engineering Journal, 2023, 466: 143050. |
| 120 | RUFFING A M, KALLAS T. Editorial: cyanobacteria: the green E. coli [J]. Frontiers in Bioengineering and Biotechnology, 2016, 4: 7. |
| 121 | HU Q, SOMMERFELD M, JARVIS E, et al. Microalgal triacylglycerols as feedstocks for biofuel production: perspectives and advances[J]. The Plant Journal, 2008, 54(4): 621-639. |
| 122 | LIN P C, ZHANG F Z, PAKRASI H B. Enhanced production of sucrose in the fast-growing cyanobacterium Synechococcus elongatus UTEX 2973[J]. Scientific Reports, 2020, 10(1): 390. |
| 123 | YU J J, LIBERTON M, CLIFTEN P F, et al. Synechococcus elongatus UTEX 2973, a fast growing cyanobacterial chassis for biosynthesis using light and CO2 [J]. Scientific Reports, 2015, 5: 8132. |
| 124 | LI S B, SUN T, XU C X, et al. Development and optimization of genetic toolboxes for a fast-growing cyanobacterium Synechococcus elongatus UTEX 2973[J]. Metabolic Engineering, 2018, 48: 163-174. |
| 125 | TAN X M, HOU S W, SONG K, et al. The primary transcriptome of the fast-growing Cyanobacterium Synechococcus elongatus UTEX 2973[J]. Biotechnology for Biofuels, 2018, 11: 218. |
| 126 | LÖWE H, SCHMAUDER L, HOBMEIER K, et al. Metabolic engineering to expand the substrate spectrum of Pseudomonas putida toward sucrose[J]. Microbiology Open, 2017, 6(4): e00473. |
| 127 | LIU J Z, WU Y H, WU C X, et al. Advanced nutrient removal from surface water by a consortium of attached microalgae and bacteria: a review[J]. Bioresource Technology, 2017, 241: 1127-1137. |
| 128 | YU W, ZENG Y, WANG Z H, et al. Solar-powered multi-organism symbiont mimic system for beyond natural synthesis of polypeptides from CO2 and N2 [J]. Science Advances, 2023, 9(11): eadf6772. |
| 129 | ALNAHHAS R N, SADEGHPOUR M, CHEN Y, et al. Majority sensing in synthetic microbial consortia[J]. Nature Communications, 2020, 11(1): 3659. |
| 130 | KONG W T, MELDGIN D R, COLLINS J J, et al. Designing microbial consortia with defined social interactions[J]. Nature Chemical Biology, 2018, 14(8): 821-829. |
| 131 | KYLILIS N, TUZA Z A, STAN G B, et al. Tools for engineering coordinated system behaviour in synthetic microbial consortia[J]. Nature Communications, 2018, 9(1): 2677. |
| 132 | KIM J K, CHEN Y, HIRNING A J, et al. Long-range temporal coordination of gene expression in synthetic microbial consortia[J]. Nature Chemical Biology, 2019, 15(11): 1102-1109. |
| 133 | ROELL G W, ZHA J, CARR R R, et al. Engineering microbial consortia by division of labor[J]. Microbial Cell Factories, 2019, 18(1): 35. |
| 134 | LI Z H, WANG X N, ZHANG H R. Balancing the non-linear rosmarinic acid biosynthetic pathway by modular co-culture engineering[J]. Metabolic Engineering, 2019, 54: 1-11. |
| 135 | WANG X, LIU W, XIN C P, et al. Enhanced limonene production in cyanobacteria reveals photosynthesis limitations[J]. Proceedings of the National Academy of Sciences of the United States of America, 2016, 113(50): 14225-14230. |
| 136 | RICHARDSON K N, BLACK W B, LI H. Aldehyde production in crude lysate- and whole cell-based biotransformation using a noncanonical redox cofactor system[J]. ACS Catalysis, 2020, 10(15): 8898-8903. |
| 137 | LI C F, YIN L J, WANG J W, et al. Light-driven biosynthesis of volatile, unstable and photosensitive chemicals from CO2 [J]. Nature Synthesis, 2023, 2: 960-971. |
| 138 | WANG H, ZHANG W, CHEN L, et al. The contamination and control of biological pollutants in mass cultivation of microalgae[J]. Bioresource Technology, 2013, 128: 745-750. |
| 139 | FUCHS G. Alternative pathways of carbon dioxide fixation: insights into the early evolution of life?[J]. Annual Review of Microbiology, 2011, 65: 631-658. |
| 140 | KUMAR M, SUNDARAM S, GNANSOUNOU E, et al. Carbon dioxide capture, storage and production of biofuel and biomaterials by bacteria: a review[J]. Bioresource Technology, 2018, 247: 1059-1068. |
| 141 | WŁODARCZYK A, SELÃO T T, NORLING B, et al. Newly discovered Synechococcus sp. PCC 11901 is a robust cyanobacterial strain for high biomass production[J]. Communications Biology, 2020, 3(1): 215. |
| 142 | ZHANG S S, SUN J H, FENG D D, et al. Unlocking the potentials of cyanobacterial photosynthesis for directly converting carbon dioxide into glucose[J]. Nature Communications, 2023, 14(1): 3425. |
| 143 | ZHU Z, JIANG J H, FA Y. Overcoming the biological contamination in microalgae and cyanobacteria mass cultivations for photosynthetic biofuel production[J]. Molecules, 2020, 25(22): 5220. |
| 144 | SING S FON, ISDEPSKY A, BOROWITZKA M A, et al. Pilot-scale continuous recycling of growth medium for the mass culture of a halotolerant Tetraselmis sp. in raceway ponds under increasing salinity: a novel protocol for commercial microalgal biomass production[J]. Bioresource Technology, 2014, 161: 47-54. |
| 145 | BACELLAR MENDES L B, VERMELHO A B. Allelopathy as a potential strategy to improve microalgae cultivation[J]. Biotechnology for Biofuels, 2013, 6(1): 152. |
| 146 | SINGH J S, KUMAR A, RAI A N, et al. Cyanobacteria: a precious bio-resource in agriculture, ecosystem, and environmental sustainability[J]. Frontiers in Microbiology, 2016, 7: 529. |
| 147 | SELÃO T T, WŁODARCZYK A, NIXON P J, et al. Growth and selection of the cyanobacterium Synechococcus sp. PCC 7002 using alternative nitrogen and phosphorus sources[J]. Metabolic Engineering, 2019, 54: 255-263. |
| 148 | GIFUNI I, POLLIO A, SAFI C, et al. Current bottlenecks and challenges of the microalgal biorefinery[J]. Trends in Biotechnology, 2019, 37(3): 242-252. |
| 149 | HOBISCH M, SPASIC J, MALIHAN-YAP L, et al. Internal illumination to overcome the cell density limitation in the scale-up of whole-cell photobiocatalysis[J]. ChemSusChem, 2021, 14(15): 3219-3225. |
| 150 | PRABHA S, VIJAY A K, PAUL R R, et al. Cyanobacterial biorefinery: towards economic feasibility through the maximum valorization of biomass[J]. Science of the Total Environment, 2022, 814: 152795. |
| 151 | JODLBAUER J, ROHR T, SPADIUT O, et al. Biocatalysis in green and blue: cyanobacteria[J]. Trends in Biotechnology, 2021, 39(9): 875-889. |
| 152 | LIN Y L, SHI J Y, FENG W, et al. Periplasmic biomineralization for semi-artificial photosynthesis[J]. Science Advances, 2023, 9(29): eadg5858. |
| 153 | PI S S, YANG W J, FENG W, et al. Solar-driven waste-to-chemical conversion by wastewater-derived semiconductor biohybrids[J]. Nature Sustainability, 2023, 6: 1673-1684. |
| 154 | HU Q S, HU H T, CUI L, et al. Ultrafast electron transfer in Au-cyanobacteria hybrid for solar to chemical production[J]. ACS Energy Letters, 2023, 8(1): 677-684. |
| 155 | LIANG J, CHEN Z, YIN P Q, et al. Efficient semi-artificial photosynthesis of ethylene by a self-assembled InP-cyanobacterial biohybrid system[J]. ChemSusChem, 2023, 16(20): e202300773. |
| [1] | GAO Ge, BIAN Qi, WANG Baojun. Synthetic genetic circuit engineering: principles, advances and prospects [J]. Synthetic Biology Journal, 2025, 6(1): 45-64. |
| [2] | LI Jiyuan, WU Guosheng. Two hypothesises for the origins of organisms from the synthetic biology perspective [J]. Synthetic Biology Journal, 2025, 6(1): 190-202. |
| [3] | JIAO Hongtao, QI Meng, SHAO Bin, JIANG Jinsong. Legal issues for the storage of DNA data [J]. Synthetic Biology Journal, 2025, 6(1): 177-189. |
| [4] | TANG Xinghua, LU Qianneng, HU Yilin. Philosophical reflections on synthetic biology in the Anthropocene [J]. Synthetic Biology Journal, 2025, 6(1): 203-212. |
| [5] | XU Huaisheng, SHI Xiaolong, LIU Xiaoguang, XU Miaomiao. Key technologies for DNA storage: encoding, error correction, random access, and security [J]. Synthetic Biology Journal, 2025, 6(1): 157-176. |
| [6] | SHI Ting, SONG Zhan, SONG Shiyi, ZHANG Yi-Heng P. Job. In vitro BioTransformation (ivBT): a new frontier of industrial biomanufacturing [J]. Synthetic Biology Journal, 2024, 5(6): 1437-1460. |
| [7] | CHAI Meng, WANG Fengqing, WEI Dongzhi. Synthesis of organic acids from lignocellulose by biotransformation [J]. Synthetic Biology Journal, 2024, 5(6): 1242-1263. |
| [8] | SHAO Mingwei, SUN Simian, YANG Shimao, CHEN Guoqiang. Bioproduction based on extremophiles [J]. Synthetic Biology Journal, 2024, 5(6): 1419-1436. |
| [9] | CHEN Yu, ZHANG Kang, QIU Yijing, CHENG Caiyun, YIN Jingjing, SONG Tianshun, XIE Jingjing. Progress of microbial electrosynthesis for conversion of CO2 [J]. Synthetic Biology Journal, 2024, 5(5): 1142-1168. |
| [10] | CHEN Ziling, XIANG Yangfei. Integrated development of organoid technology and synthetic biology [J]. Synthetic Biology Journal, 2024, 5(4): 795-812. |
| [11] | CAI Bingyu, TAN Xiangtian, LI Wei. Advances in synthetic biology for engineering stem cell [J]. Synthetic Biology Journal, 2024, 5(4): 782-794. |
| [12] | XIE Huang, ZHENG Yilei, SU Yiting, RUAN Jingyi, LI Yongquan. An overview on reconstructing the biosynthetic system of actinomycetes for polyketides production [J]. Synthetic Biology Journal, 2024, 5(3): 612-630. |
| [13] | ZHA Wenlong, BU Lan, ZI Jiachen. Advances in synthetic biology for producing potent pharmaceutical ingredients of traditional Chinese medicine [J]. Synthetic Biology Journal, 2024, 5(3): 631-657. |
| [14] | HUI Zhen, TANG Xiaoyu. Applications of the CRISPR/Cas9 editing system in the study of microbial natural products [J]. Synthetic Biology Journal, 2024, 5(3): 658-671. |
| [15] | LIU Xiaonan, LI Jing, ZHU Xiaoxi, XU Zishuo, QI Jian, JIANG Huifeng. Research advances on paclitaxel biosynthesis [J]. Synthetic Biology Journal, 2024, 5(3): 527-547. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||