Synthetic Biology Journal ›› 2024, Vol. 5 ›› Issue (4): 795-812.DOI: 10.12211/2096-8280.2023-106
• Invited Review • Previous Articles Next Articles
Integrated development of organoid technology and synthetic biology
CHEN Ziling, XIANG Yangfei
- School of Life Science and Technology,ShanghaiTech University,Shanghai 201210,China
-
Received:2023-12-09Revised:2024-02-23Online:2024-09-19Published:2024-08-31 -
Contact:XIANG Yangfei
类器官技术与合成生物学协同研究进展
陈子苓, 向阳飞
- 上海科技大学生命科学与技术学院,上海 201210
-
通讯作者:向阳飞 -
作者简介:陈子苓 (2000—),女,硕士研究生。研究方向为神经类器官。E-mail:chenzl2022@shanghaitech.edu.cn向阳飞 (1983—),男,博士,研究员。研究方向为干细胞与神经生物学。E-mail:xiangyf@shanghaitech.edu.cn -
基金资助:国家重点研发计划(2021YFF1200800);国家自然科学基金(32170836);中央引导地方项目(YDZX20233100001002)
CLC Number:
Cite this article
CHEN Ziling, XIANG Yangfei. Integrated development of organoid technology and synthetic biology[J]. Synthetic Biology Journal, 2024, 5(4): 795-812.
陈子苓, 向阳飞. 类器官技术与合成生物学协同研究进展[J]. 合成生物学, 2024, 5(4): 795-812.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2023-106
| 1 | LANCASTER M A, KNOBLICH J A. Generation of cerebral organoids from human pluripotent stem cells[J]. Nature Protocols, 2014, 9(10): 2329-2340. |
| 2 | XIANG Y F, CAKIR B, PARK I H. Generation of regionally specified human brain organoids resembling thalamus development[J]. STAR Protocols, 2020, 1(1): 100001. |
| 3 | WANG J Y, NIELSEN J, LIU Z H. Synthetic biology advanced natural product discovery[J]. Metabolites, 2021, 11(11): 785. |
| 4 | 李童, 杨劲树, 杨卫军. 肺成体干细胞体外培养模型的研究进展[J]. 生物工程学报, 2022, 38(9): 3255-3266. |
| LI T, YANG J S, YANG W J. Advances of in vitro culture models derived from lung adult stem cells[J]. Chinese Journal of Biotechnology, 2022, 38(9): 3255-3266. | |
| 5 | NIKOLAEV M, MITROFANOVA O, BROGUIERE N, et al. Homeostatic mini-intestines through scaffold-guided organoid morphogenesis[J]. Nature, 2020, 585(7826): 574-578. |
| 6 | CLEVERS H. Modeling development and disease with organoids[J]. Cell, 2016, 165(7): 1586-1597. |
| 7 | LI M, IZPISUA BELMONTE J C. Organoids - preclinical models of human disease[J]. The New England Journal of Medicine, 2019, 380(6): 569-579. |
| 8 | LANCASTER M A. Brain organoids: a new frontier of human neuroscience research[J]. Seminars in Cell & Developmental Biology, 2021, 111: 1-3. |
| 9 | MEIER A B, ZAWADA D, DE ANGELIS M T, et al. Epicardioid single-cell genomics uncovers principles of human epicardium biology in heart development and disease[J]. Nature Biotechnology, 2023, 41(12): 1787-1800. |
| 10 | UZQUIANO A, KEDAIGLE A J, PIGONI M, et al. Proper acquisition of cell class identity in organoids allows definition of fate specification programs of the human cerebral cortex[J]. Cell, 2022, 185(20): 3770-3788. |
| 11 | BOUFFI C, WIKENHEISER-BROKAMP K A, CHATURVEDI P, et al. In vivo development of immune tissue in human intestinal organoids transplanted into humanized mice[J]. Nature Biotechnology, 2023, 41(6): 824-831. |
| 12 | GABRIEL E, ALBANNA W, PASQUINI G, et al. Human brain organoids assemble functionally integrated bilateral optic vesicles[J]. Cell Stem Cell, 2021, 28(10): 1740-1757. |
| 13 | SCHMIDT C, DEYETT A, ILMER T, et al. Multi-chamber cardioids unravel human heart development and cardiac defects[J]. Cell, 2023, 186(25): 5587-5605. |
| 14 | FAUSTINO MARTINS J M, FISCHER C, URZI A, et al. Self-organizing 3D human trunk neuromuscular organoids[J]. Cell Stem Cell, 2020, 26(2): 172-186. |
| 15 | PELLEGRINI L, LANCASTER M A. Modeling neurodegeneration with mutant-tau organoids[J]. Cell, 2021, 184(17): 4377-4379. |
| 16 | JIANG S W, ZHAO H R, ZHANG W J, et al. An automated organoid platform with inter-organoid homogeneity and inter-patient heterogeneity[J]. Cell Reports Medicine, 2020, 1(9): 100161. |
| 17 | CORRÒ C, NOVELLASDEMUNT L, LI V S W. A brief history of organoids[J]. American Journal of Physiology Cell Physiology, 2020, 319(1): C151-C165. |
| 18 | BIJUKUMAR G, SOMVANSHI P R. Reverse engineering in biotechnology: the role of genetic engineering in synthetic biology[J]. Methods in Molecular Biology, 2024, 2719: 307-324. |
| 19 | CAMERON D E, BASHOR C J, COLLINS J J. A brief history of synthetic biology[J]. Nature Reviews Microbiology, 2014, 12(5): 381-390. |
| 20 | SMITH E, COCHRANE W J. Cystic organoid teratoma; report of a case[J]. Canadian Medical Association Journal, 1946, 55(2): 151. |
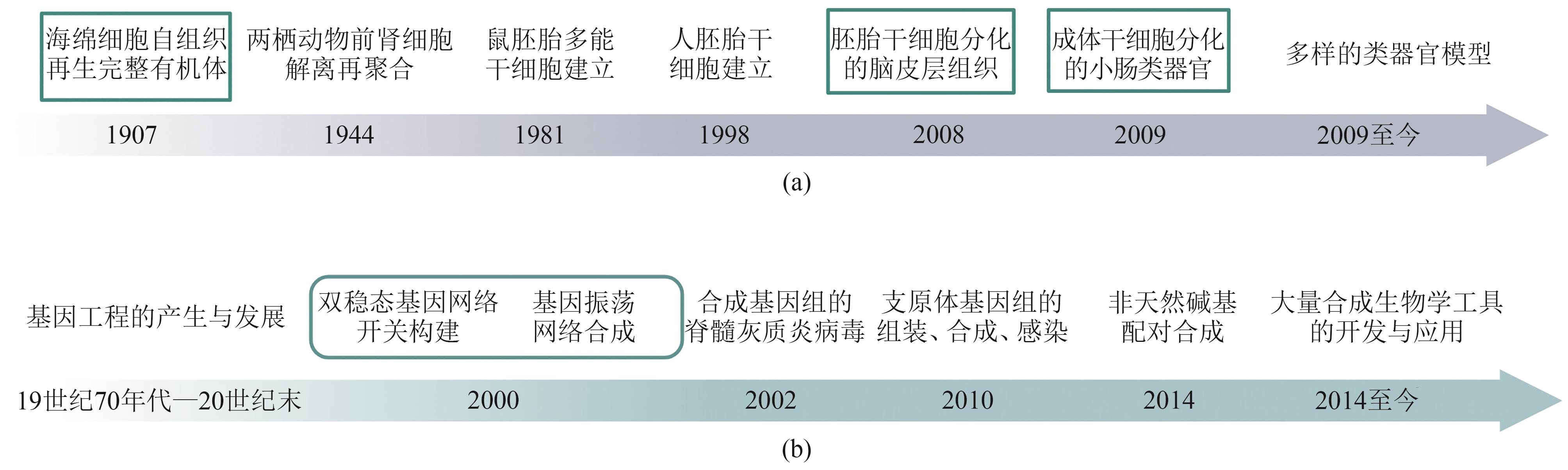
| 21 | WILSON H V. A new method by which sponges may be artificially reared[J]. Science, 1907, 25(649): 912-915. |
| 22 | TUNG T C, KÜ S H. Experimental studies on the development of the pronephric duct in anuran embryos[J]. Journal of Anatomy, 1944, 78(1/2): 52-57. |
| 23 | WEISS P, TAYLOR A C. Reconstitution of complete organs from single-cell suspensions of chick embryos in advanced stages of differentiation[J]. Proceedings of the National Academy of Sciences of the United States of America, 1960, 46(9): 1177-1185. |
| 24 | EIRAKU M, WATANABE K, MATSUO-TAKASAKI M, et al. Self-organized formation of polarized cortical tissues from ESCs and its active manipulation by extrinsic signals[J]. Cell Stem Cell, 2008, 3(5): 519-532. |
| 25 | SATO T, VRIES R G, SNIPPERT H J, et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche[J]. Nature, 2009, 459(7244): 262-265. |
| 26 | KIM T H, SHIVDASANI R A. Stomach development, stem cells and disease[J]. Development, 2016, 143(4): 554-565. |
| 27 | BROUTIER L, ANDERSSON-ROLF A, HINDLEY C J, et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation[J]. Nature Protocols, 2016, 11(9): 1724-1743. |
| 28 | HUCH M, GEHART H, VAN BOXTEL R, et al. Long-term culture of genome-stable bipotent stem cells from adult human liver[J]. Cell, 2015, 160(1/2): 299-312. |
| 29 | NARAYANAN S, SURETTE F A, HAHN Y S. The immune landscape in nonalcoholic steatohepatitis[J]. Immune Network, 2016, 16(3): 147-158. |
| 30 | TAKEBE T, SEKINE K, ENOMURA M, et al. Vascularized and functional human liver from an iPSC-derived organ bud transplant[J]. Nature, 2013, 499(7459): 481-484. |
| 31 | MORIZANE R, BONVENTRE J V. Generation of nephron progenitor cells and kidney organoids from human pluripotent stem cells[J]. Nature Protocols, 2017, 12(1): 195-207. |
| 32 | CHEN Y W, HUANG S X, DE CARVALHO A L R T, et al. A three-dimensional model of human lung development and disease from pluripotent stem cells[J]. Nature Cell Biology, 2017, 19(5): 542-549. |
| 33 | LANCASTER M A, RENNER M, MARTIN C A, et al. Cerebral organoids model human brain development and microcephaly[J]. Nature, 2013, 501(7467): 373-379. |
| 34 | QIAN X Y, NGUYEN H N, SONG M M, et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure[J]. Cell, 2016, 165(5): 1238-1254. |
| 35 | ASSAWACHANANONT J, MANDAI M, OKAMOTO S, et al. Transplantation of embryonic and induced pluripotent stem cell-derived 3D retinal sheets into retinal degenerative mice[J]. Stem Cell Reports, 2014, 2(5): 662-674. |
| 36 | KIM M S, MUN H M, SUNG C O, et al. Patient-derived lung cancer organoids as in vitro cancer models for therapeutic screening[J]. Nature Communications, 2019, 10(1): 3991. |
| 37 | QU M L, XIONG L, LÜ Y L, et al. Establishment of intestinal organoid cultures modeling injury-associated epithelial regeneration[J]. Cell Research, 2021, 31(3): 259-271. |
| 38 | LU X X, YANG J J, XIANG Y F. Modeling human neurodevelopmental diseases with brain organoids[J]. Cell Regeneration, 2022, 11(1): 1. |
| 39 | TANG X Y, WU S S, WANG D, et al. Human organoids in basic research and clinical applications[J]. Signal Transduction and Targeted Therapy, 2022, 7(1): 168. |
| 40 | CAKIR B, XIANG Y F, TANAKA Y, et al. Engineering of human brain organoids with a functional vascular-like system[J]. Nature Methods, 2019, 16(11): 1169-1175. |
| 41 | ZHAO Z X, CHEN X Y, DOWBAJ A M, et al. Organoids[J]. Nature Reviews Methods Primers, 2022, 2: 94. |
| 42 | TAO T T, DENG P W, WANG Y Q, et al. Microengineered multi-organoid system from hiPSCs to recapitulate human liver-islet axis in normal and type 2 diabetes[J]. Advanced Science, 2022, 9(5): e2103495. |
| 43 | ZHANG W J, LI J W, ZHOU J Q, et al. Translational organoid technology — the convergence of chemical, mechanical, and computational biology[J]. Trends in Biotechnology, 2022, 40(9): 1121-1135. |
| 44 | PARK S E, GEORGESCU A, HUH D. Organoids-on-a-chip[J]. Science, 2019, 364(6444): 960-965. |
| 45 | JI S Y, FENG L, FU Z L, et al. Pharmaco-proteogenomic characterization of liver cancer organoids for precision oncology[J]. Science Translational Medicine, 2023, 15(706): eadg3358. |
| 46 | LIN L, DEMARTINO J, WANG D S, et al. Unbiased transcription factor CRISPR screen identifies ZNF800 as master repressor of enteroendocrine differentiation[J]. Science, 2023, 382(6669): 451-458. |
| 47 | HENDRIKS D, BROUWERS J F, HAMER K, et al. Engineered human hepatocyte organoids enable CRISPR-based target discovery and drug screening for steatosis[J]. Nature Biotechnology, 2023, 41(11): 1567-1581. |
| 48 | HANSEN S L, LARSEN H L, PIKKUPEURA L M, et al. An organoid-based CRISPR-Cas9 screen for regulators of intestinal epithelial maturation and cell fate[J]. Science Advances, 2023, 9(28): eadg4055. |
| 49 | GARDNER T S, CANTOR C R, COLLINS J J. Construction of a genetic toggle switch in Escherichia coli [J]. Nature, 2000, 403(6767): 339-342. |
| 50 | ELOWITZ M B, LEIBLER S. A synthetic oscillatory network of transcriptional regulators[J]. Nature, 2000, 403(6767): 335-338. |
| 51 | MUELLER S, CAO X M, WELKER R, et al. Interaction of the poliovirus receptor CD155 with the dynein light chain Tctex-1 and its implication for poliovirus pathogenesis[J]. The Journal of Biological Chemistry, 2002, 277(10): 7897-7904. |
| 52 | GIBSON D G, GLASS J I, LARTIGUE C, et al. Creation of a bacterial cell controlled by a chemically synthesized genome[J]. Science, 2010, 329(5987): 52-56. |
| 53 | GIBSON D G, YOUNG L, CHUANG R Y, et al. Enzymatic assembly of DNA molecules up to several hundred kilobases[J]. Nature Methods, 2009, 6(5): 343-345. |
| 54 | LI L J, DEGARDIN M, LAVERGNE T, et al. Natural-like replication of an unnatural base pair for the expansion of the genetic alphabet and biotechnology applications[J]. Journal of the American Chemical Society, 2014, 136(3): 826-829. |
| 55 | SHI Y C, WU Q, WANG X Q. Modeling brain development and diseases with human cerebral organoids[J]. Current Opinion in Neurobiology, 2021, 66: 103-115. |
| 56 | MENG F K, ELLIS T. The second decade of synthetic biology: 2010—2020[J]. Nature Communications, 2020, 11(1): 5174. |
| 57 | ANDREWS L B, NIELSEN A A K, VOIGT C A. Cellular checkpoint control using programmable sequential logic[J]. Science, 2018, 361(6408): eaap8987. |
| 58 | MIURA Y, LI M Y, BIREY F, et al. Generation of human striatal organoids and cortico-striatal assembloids from human pluripotent stem cells[J]. Nature Biotechnology, 2020, 38(12): 1421-1430. |
| 59 | TODA S, BLAUCH L R, TANG S K Y, et al. Programming self-organizing multicellular structures with synthetic cell-cell signaling[J]. Science, 2018, 361(6398): 156-162. |
| 60 | MCCRACKEN K W, CATÁ E M, CRAWFORD C M, et al. Modelling human development and disease in pluripotent stem-cell-derived gastric organoids[J]. Nature, 2014, 516(7531): 400-404. |
| 61 | EIRAKU M, TAKATA N, ISHIBASHI H, et al. Self-organizing optic-cup morphogenesis in three-dimensional culture[J]. Nature, 2011, 472(7341): 51-56. |
| 62 | CRUZ N M, SONG X W, CZERNIECKI S M, et al. Organoid cystogenesis reveals a critical role of microenvironment in human polycystic kidney disease[J]. Nature Materials, 2017, 16(11): 1112-1119. |
| 63 | CEDERQUIST G Y, ASCIOLLA J J, TCHIEU J, et al. Specification of positional identity in forebrain organoids[J]. Nature Biotechnology, 2019, 37(4): 436-444. |
| 64 | DEMERS C J, SOUNDARARAJAN P, CHENNAMPALLY P, et al. Development-on-chip: in vitro neural tube patterning with a microfluidic device[J]. Development, 2016, 143(11): 1884-1892. |
| 65 | RIFES P, ISAKSSON M, RATHORE G S, et al. Modeling neural tube development by differentiation of human embryonic stem cells in a microfluidic WNT gradient[J]. Nature Biotechnology, 2020, 38(11): 1265-1273. |
| 66 | LEE K K, MCCAULEY H A, BRODA T R, et al. Human stomach-on-a-chip with luminal flow and peristaltic-like motility[J]. Lab on a Chip, 2018, 18(20): 3079-3085. |
| 67 | HOMAN K A, GUPTA N, KROLL K T, et al. Flow-enhanced vascularization and maturation of kidney organoids in vitro [J]. Nature Methods, 2019, 16(3): 255-262. |
| 68 | WANG Y Q, WANG H, DENG P W, et al. In situ differentiation and generation of functional liver organoids from human iPSCs in a 3D perfusable chip system[J]. Lab on a Chip, 2018, 18(23): 3606-3616. |
| 69 | TAO T T, WANG Y Q, CHEN W W, et al. Engineering human islet organoids from iPSCs using an organ-on-chip platform[J]. Lab on a Chip, 2019, 19(6): 948-958. |
| 70 | LIU X, CHENG Y L, ABRAHAM J M, et al. Modeling Wnt signaling by CRISPR-Cas9 genome editing recapitulates neoplasia in human Barrett epithelial organoids[J]. Cancer Letters, 2018, 436: 109-118. |
| 71 | ZHAO H, CHENG Y L, KALRA A, et al. Generation and multiomic profiling of a TP53/CDKN2A double-knockout gastroesophageal junction organoid model[J]. Science Translational Medicine, 2022, 14(673): eabq6146. |
| 72 | TRUJILLO C A, RICE E S, SCHAEFER N K, et al. Reintroduction of the archaic variant of NOVA1 in cortical organoids alters neurodevelopment[J]. Science, 2021, 371(6530): eaax2537. |
| 73 | ARTEGIANI B, HENDRIKS D, BEUMER J, et al. Fast and efficient generation of knock-in human organoids using homology-independent CRISPR-Cas9 precision genome editing[J]. Nature Cell Biology, 2020, 22(3): 321-331. |
| 74 | ESK C, LINDENHOFER D, HAENDELER S, et al. A human tissue screen identifies a regulator of ER secretion as a brain-size determinant[J]. Science, 2020, 370(6519): 935-941. |
| 75 | BORTOLOMAI I, SANDRI M, DRAGHICI E, et al. Gene modification and three-dimensional scaffolds as novel tools to allow the use of postnatal thymic epithelial cells for Thymus regeneration approaches[J]. Stem Cells Translational Medicine, 2019, 8(10): 1107-1122. |
| 76 | ROTHENBÜCHER T S P, GÜRBÜZ H, PEREIRA M P, et al. Next generation human brain models: engineered flat brain organoids featuring gyrification[J]. Biofabrication, 2021, 13(1): 011001. |
| 77 | CHEN S S, CHEN X, GENG Z, et al. The horizon of bone organoid: a perspective on construction and application[J]. Bioactive Materials, 2022, 18: 15-25. |
| 78 | Jamaluddin M F B, Ghosh A, Ingle A, et al. Bovine and human endometrium-derived hydrogels support organoid culture from healthy and cancerous tissues. Proceedings of the National Academy of Sciences of the United States of America, 2022, 119(44): e2208040119 |
| 79 | XIANG Y F, TANAKA Y, PATTERSON B, et al. Fusion of regionally specified hPSC-derived organoids models human brain development and interneuron migration[J]. Cell Stem Cell, 2017, 21(3): 383-398. e7. |
| 80 | ANDERSEN J, REVAH O, MIURA Y, et al. Generation of functional human 3D cortico-motor assembloids[J]. Cell, 2020, 183(7): 1913-1929. |
| 81 | LIND J U, BUSBEE T A, VALENTINE A D, et al. Instrumented cardiac microphysiological devices via multimaterial three-dimensional printing[J]. Nature Materials, 2017, 16(3): 303-308. |
| 82 | ZHANG Y S, ALEMAN J, SHIN S R, et al. Multisensor-integrated organs-on-chips platform for automated and continual in situ monitoring of organoid behaviors[J]. Proceedings of the National Academy of Sciences of the United States of America, 2017, 114(12): E2293-E2302. |
| 83 | STEPHENSON A, LASTRA L, NGUYEN B, et al. Physical laboratory automation in synthetic biology[J]. ACS Synthetic Biology, 2023, 12(11): 3156-3169. |
| 84 | CHIA N, LEE S Y, TONG Y J. Optogenetic tools for microbial synthetic biology[J]. Biotechnology Advances, 2022, 59: 107953. |
| 85 | REPINA N A, JOHNSON H J, BAO X P, et al. Optogenetic control of Wnt signaling models cell-intrinsic embryogenic patterning using 2D human pluripotent stem cell culture[J]. Development, 2023, 150(14): dev201386. |
| 86 | LEGNINI I, EMMENEGGER L, ZAPPULO A, et al. Spatiotemporal, optogenetic control of gene expression in organoids[J]. Nature Methods, 2023, 20(10): 1544-1552. |
| 87 | MANHAS J, EDELSTEIN H I, LEONARD J N, et al. The evolution of synthetic receptor systems[J]. Nature Chemical Biology, 2022, 18(3): 244-255. |
| 88 | WANG H F, ZANG C Z, LIU X S, et al. The role of Notch receptors in transcriptional regulation[J]. Journal of Cellular Physiology, 2015, 230(5): 982-988. |
| 89 | MORSUT L, ROYBAL K T, XIONG X, et al. Engineering customized cell sensing and response behaviors using synthetic Notch receptors[J]. Cell, 2016, 164(4): 780-791. |
| 90 | MALAGUTI M, PORTERO MIGUELES R, ANNOH J, et al. SyNPL: synthetic Notch pluripotent cell lines to monitor and manipulate cell interactions in vitro and in vivo [J]. Development, 2022, 149(12): dev200226. |
| 91 | TODA S, MCKEITHAN W L, HAKKINEN T J, et al. Engineering synthetic morphogen systems that can program multicellular patterning[J]. Science, 2020, 370(6514): 327-331. |
| 92 | FOTY R A, STEINBERG M S. The differential adhesion hypothesis: a direct evaluation[J]. Developmental Biology, 2005, 278(1): 255-263. |
| 93 | STEVENS A J, HARRIS A R, GERDTS J, et al. Programming multicellular assembly with synthetic cell adhesion molecules[J]. Nature, 2023, 614(7946): 144-152. |
| 94 | TRENTESAUX C, YAMADA T, KLEIN O D, et al. Harnessing synthetic biology to engineer organoids and tissues[J]. Cell Stem Cell, 2023, 30(1): 10-19. |
| 95 | ZHU J W, CHU P, FU X F. Unbalanced response to growth variations reshapes the cell fate decision landscape[J]. Nature Chemical Biology, 2023, 19(9): 1097-1104. |
| 96 | CAMP J G, BADSHA F, FLORIO M, et al. Human cerebral organoids recapitulate gene expression programs of fetal neocortex development[J]. Proceedings of the National Academy of Sciences of the United States of America, 2015, 112(51): 15672-15677. |
| 97 | PAULSEN B, VELASCO S, KEDAIGLE A J, et al. Autism genes converge on asynchronous development of shared neuron classes[J]. Nature, 2022, 602(7896): 268-273. |
| 98 | AMIN N D, KELLEY K W, HAO J, et al. Generating human neural diversity with a multiplexed morphogen screen in organoids[EB/OL]. bioRxiv, 2023: 2023.05.31.541819[2023-12-01]. . |
| 99 | HE Z S, DONY L, Fleck J S, et al. An integrated transcriptomic cell atlas of human neural organoids[EB/OL]. bioRxiv, 2023: 2023.10.05.561097[2023-12-01]. . |
| 100 | MENG X L, YAO D, IMAIZUMI K, et al. Assembloid CRISPR screens reveal impact of disease genes in human neurodevelopment[J]. Nature, 2023, 622(7982): 359-366. |
| [1] | GAO Ge, BIAN Qi, WANG Baojun. Synthetic genetic circuit engineering: principles, advances and prospects [J]. Synthetic Biology Journal, 2025, 6(1): 45-64. |
| [2] | DONG Ying, MA Mengdan, HUANG Weiren. Progress in the miniaturization of CRISPR-Cas systems [J]. Synthetic Biology Journal, 2025, 6(1): 105-117. |
| [3] | LI Jiyuan, WU Guosheng. Two hypothesises for the origins of organisms from the synthetic biology perspective [J]. Synthetic Biology Journal, 2025, 6(1): 190-202. |
| [4] | JIAO Hongtao, QI Meng, SHAO Bin, JIANG Jinsong. Legal issues for the storage of DNA data [J]. Synthetic Biology Journal, 2025, 6(1): 177-189. |
| [5] | TANG Xinghua, LU Qianneng, HU Yilin. Philosophical reflections on synthetic biology in the Anthropocene [J]. Synthetic Biology Journal, 2025, 6(1): 203-212. |
| [6] | XU Huaisheng, SHI Xiaolong, LIU Xiaoguang, XU Miaomiao. Key technologies for DNA storage: encoding, error correction, random access, and security [J]. Synthetic Biology Journal, 2025, 6(1): 157-176. |
| [7] | SHI Ting, SONG Zhan, SONG Shiyi, ZHANG Yi-Heng P. Job. In vitro BioTransformation (ivBT): a new frontier of industrial biomanufacturing [J]. Synthetic Biology Journal, 2024, 5(6): 1437-1460. |
| [8] | CHAI Meng, WANG Fengqing, WEI Dongzhi. Synthesis of organic acids from lignocellulose by biotransformation [J]. Synthetic Biology Journal, 2024, 5(6): 1242-1263. |
| [9] | SHAO Mingwei, SUN Simian, YANG Shimao, CHEN Guoqiang. Bioproduction based on extremophiles [J]. Synthetic Biology Journal, 2024, 5(6): 1419-1436. |
| [10] | CHEN Yu, ZHANG Kang, QIU Yijing, CHENG Caiyun, YIN Jingjing, SONG Tianshun, XIE Jingjing. Progress of microbial electrosynthesis for conversion of CO2 [J]. Synthetic Biology Journal, 2024, 5(5): 1142-1168. |
| [11] | ZHENG Haotian, LI Chaofeng, LIU Liangxu, WANG Jiawei, LI Hengrun, NI Jun. Design, optimization and application of synthetic carbon-negative phototrophic community [J]. Synthetic Biology Journal, 2024, 5(5): 1189-1210. |
| [12] | AI Zongyong, ZHANG Chengting, NIU Baohua, YIN Yu, YANG Jie, LI Tianqing. Early human embryo development and stem cells [J]. Synthetic Biology Journal, 2024, 5(4): 700-718. |
| [13] | Ke’er HU, Hanqi WANG, Ruqi HUANG, Canyang ZHANG, Xinhui XING, Shaohua MA. Integrated design strategies for engineered organoids and organ-on-a-chip technologies [J]. Synthetic Biology Journal, 2024, 5(4): 883-897. |
| [14] | Shikai LI, Dong′ao ZENG, Fangzhou DU, Jingzhong ZHANG, Shuang YU. The construction approaches and biomaterials for vascularized organoids [J]. Synthetic Biology Journal, 2024, 5(4): 851-866. |
| [15] | ZHANG Bohang, QI Xiaoxuan, YUAN Yan. Advancements in testicular organoids for in vitro spermatogenesis [J]. Synthetic Biology Journal, 2024, 5(4): 770-781. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||