Synthetic Biology Journal ›› 2022, Vol. 3 ›› Issue (6): 1201-1217.DOI: 10.12211/2096-8280.2022-041
• Invited Review • Previous Articles Next Articles
Recent progress in the molecular genetic modification tools of Clostridium
LIU Jiaxin1, CHENG Chi1,2, LI Xinqi1, WANG Chaojun2, ZHANG Ying2, XUE Chuang1,2
- 1.Engineering Research Center of Application and Transformation for Synthetic Biology Dalian,School of Bioengineering,Dalian University of Technology,Dalian 116024,Liaoning,China
2.NingBo Institute of Dalian University of Technology,Ningbo 315016,Zhejiang,China
-
Received:2022-07-18Revised:2022-08-30Online:2023-01-17Published:2022-12-31 -
Contact:CHENG Chi, XUE Chuang
梭菌分子遗传改造工具研究进展
刘佳昕1, 程驰1,2, 李欣启1, 汪超俊2, 张颖2, 薛闯1,2
- 1.大连理工大学生物工程学院,大连市合成生物学应用转化工程技术研究中心,辽宁 大连 116024
2.大连理工大学宁波研究院,浙江 宁波 315016
-
通讯作者:程驰,薛闯 -
作者简介:刘佳昕 (1998—),女,硕士研究生。主要进行生物学研究。E-mail:ljx123456@mail.dlut.edu.cn程驰 (1990—),女,副教授,研究生导师。主要研究领域为:①能源微生物遗传操作工具开发及代谢改造;②化学-生物耦合的CO2固定。E-mail:cheng.chi@dlut.edu.cn薛闯 (1982—),男,教授,博士生导师。主要从事生物质新能源的生产、分离纯化以及高效代谢菌株构建的研究:①生物法生产燃料乙醇及酵母菌的代谢网络研究;②微生物发酵法生产丁醇;③生物基化学品的高效分离;④有机膜的制备及分离技术;⑤先进能源生产菌株的构建及代谢途径信号转导;⑥激酶的生物信息分析。E-mail:xue.1@dlut.edu.cn -
基金资助:国家重点研发计划(2021YFC2102500);国家自然科学基金(21878035);大连市杰出青年科技人才支持计划(2021RJ03);大连理工大学基本科研业务费(DUT21RC(3)003, DUT22ZD102);辽宁省“百千万人才工程”经费
Cite this article
LIU Jiaxin, CHENG Chi, LI Xinqi, WANG Chaojun, ZHANG Ying, XUE Chuang. Recent progress in the molecular genetic modification tools of Clostridium[J]. Synthetic Biology Journal, 2022, 3(6): 1201-1217.
刘佳昕, 程驰, 李欣启, 汪超俊, 张颖, 薛闯. 梭菌分子遗传改造工具研究进展[J]. 合成生物学, 2022, 3(6): 1201-1217.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2022-041
| 遗传操作工具 | 优点 | 缺点 | 梭菌菌株 | 基因 | 文献 |
|---|---|---|---|---|---|
| 基于游离质粒的基因过表达 | 设计简单,易于操作 | 质粒不稳定,需靠抗生素维持 | C. acetobutylicum ATCC 824 C. acetobutylicum DSM 792 C. paraputrificum M-21 C. tyrobutyricum JM1 C. tyrobutyricum ATCC 25755 C. perfringens SM101 C. cellulolyticum H10 | spo0A, hydA, adhE2, adC, groESL, txeR, tcdC, pdc-adhII | [ |
| 反义RNA技术 | 致死率低,易于筛选 | 仅在转录水平调控 | C. acetobutylicum ATCC 824 | ptb, buk, CoAT | [ |
| 转座子突变技术 | 可用于建立突变体库 | 难以控制插入位点 | C. difficile CD37 C. perfringens Strain 13 | random | [ |
| 二型内含子技术 | 操作简单,几乎适用于所有梭菌 | 脱靶概率较大,存在极性效应 | C. acetobutylicum ATCC 824 C. beijerinckii NCIMB 8052 | glcG, cbei2385, xylR, bdhA, bdhB, ptb, ack, adc | [ |
| 同源重组 | 可精确进行基因编辑 | 同源重组效率很低 | C. acetobutylicum NCIMB 8052 | gutD, spo0A | [ |
| 反筛标记介导的等位基因替换 | 相比纯粹的等位基因替换,效率有所提升 | 受制于第一次单交换效率 | C. acetobutylicum ATCC 824 | adh | [ |
| Ⅰ-SceⅠ归巢内切酶介导的等位基因替换 | 适用范围广,在革兰氏阳性和阴性菌中均可使用 | 设计复杂,操作烦琐,周期长 | C. acetobutylicum ATCC 824 C. beijerinckii NCIMB 8052 | adc, glcG, xylR | [ |
| 噬菌体丝氨酸重组酶介导的位点特异性基因编辑技术 | 适用于大片段DNA快速整合到宿主染色体上 | 受限于附着位点attb/p,应用范围小 | C. ljungdahlii DSM 13528 | thl, crt, bcd, etfB, etfA, hbd, ptb, buk | [ |
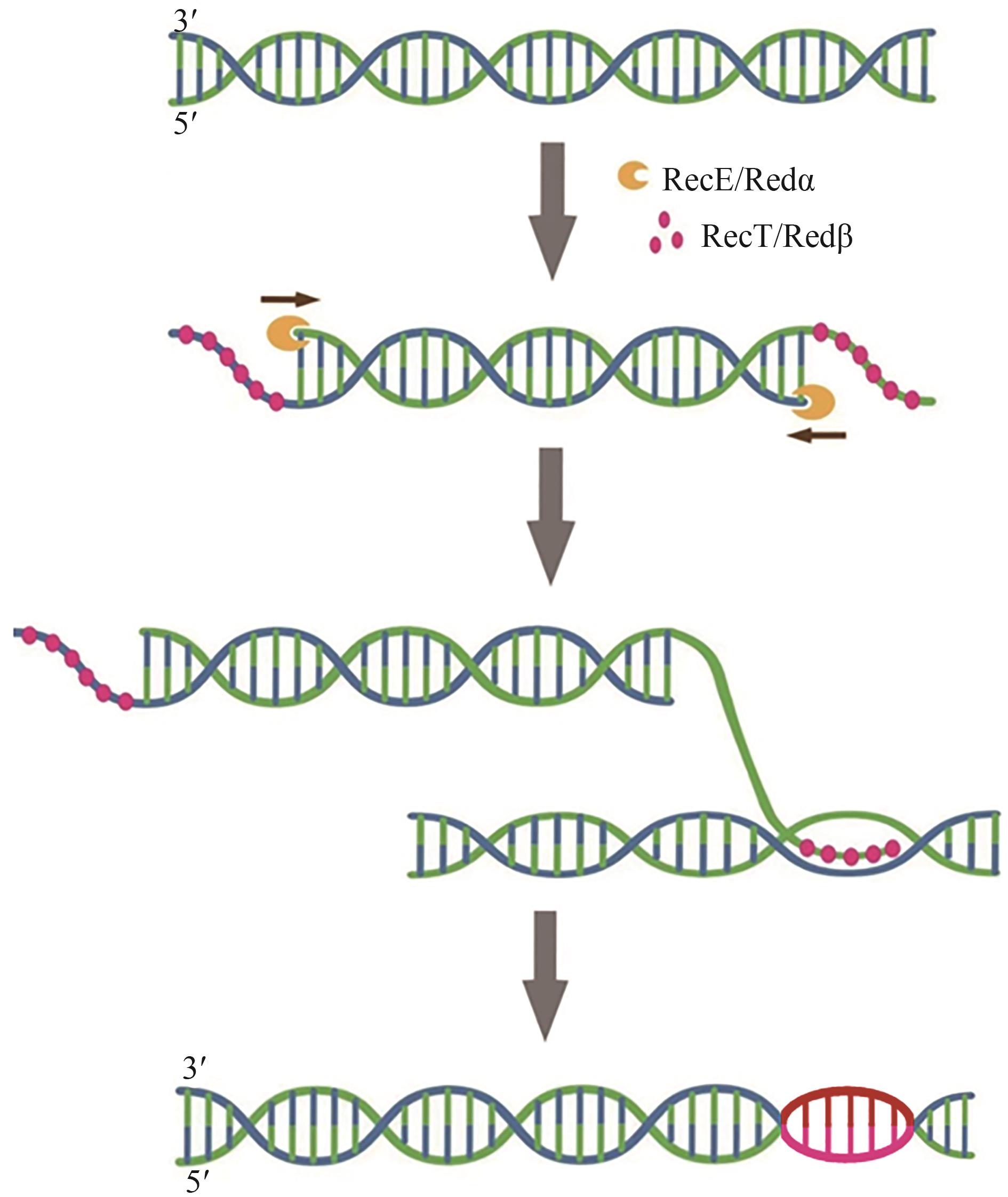
| Red/ET重组酶介导的同源重组 | 所需同源臂较短、不受限制性酶切位点限制 | 尚不完善 | C. acetobutylicum SMB009 | ermC | [ |
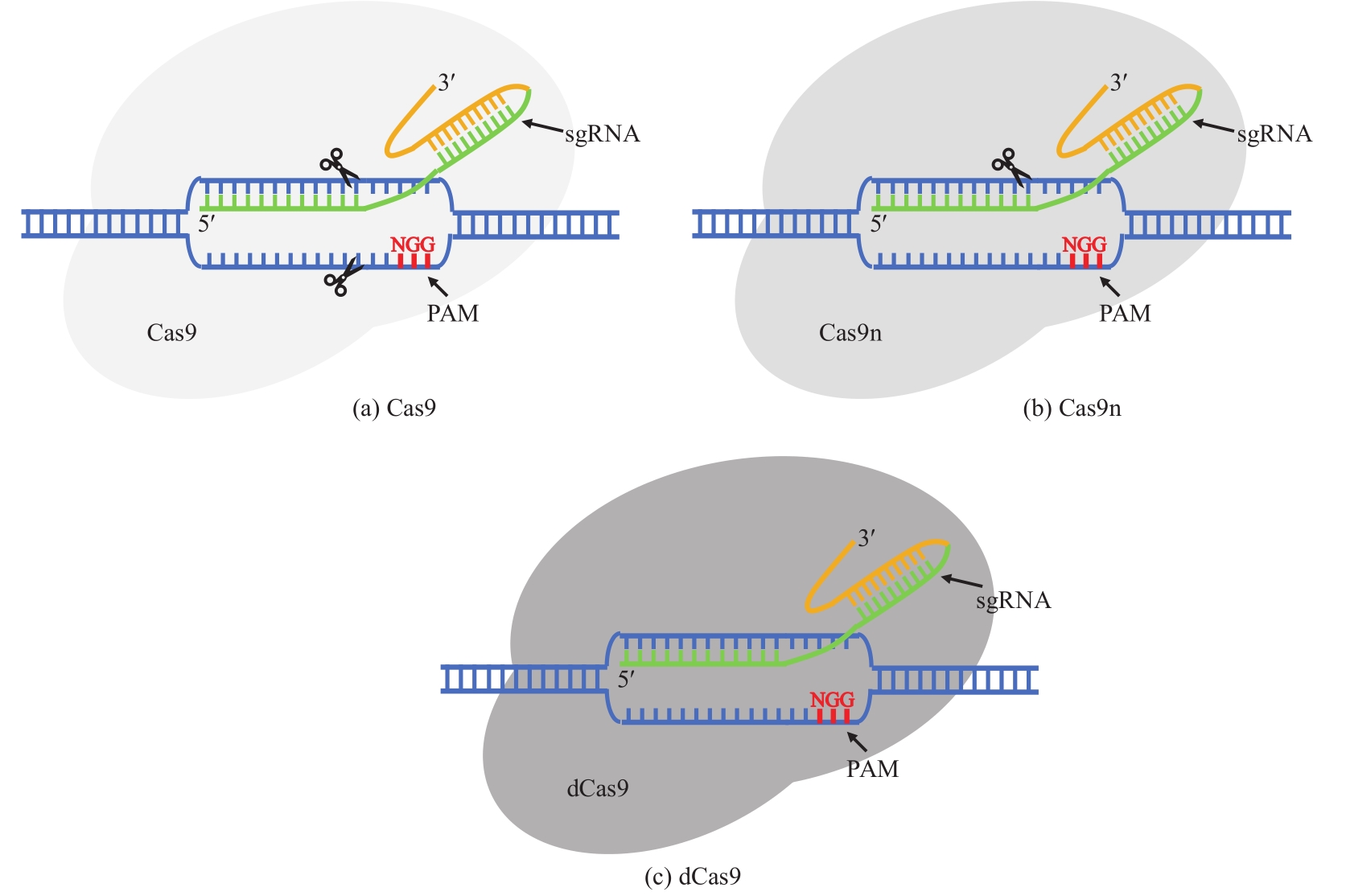
| CRISPR/Cas9系统介导的基因编辑 | 提高同源重组效率,可实现精确编辑 | 毒性大,很难获得转化子 | C. acetobutylicum DSM792 C. saccharoperbutylacetonicum N1-4 C. cellulovorans C. autoethanogenum DSM10061 | hprK, cac1502, pta, buk, clocel2243, adhE1, ctf,pyrE, 2,3-bdh | [ |
| CRISPR/Cas9n系统介导的基因编辑 | 相比于CRISPR/Cas9毒性有所降低,转化子数目增加 | 仍有毒性,转化子依然偏少 | C. cellulolyticum H10 C. acetobutylicum ATCC 824 C. beijerinckii NCIMB 8052 | pyrF, spo0A | [ |
| CRISPR/dCas9系统介导的基因表达下调 | 相比于CRISPR/Cas9毒性大大降低,易于得到转化子 | 依赖于sgRNA和基因;仅在转录水平调控 | C. pasteurianum ATCC 6013 C. acetobutylicum DSM792 | hprK, glpX | [ |
Tab. 1 Comparison of different genetic tools applicable in Clostridium
| 遗传操作工具 | 优点 | 缺点 | 梭菌菌株 | 基因 | 文献 |
|---|---|---|---|---|---|
| 基于游离质粒的基因过表达 | 设计简单,易于操作 | 质粒不稳定,需靠抗生素维持 | C. acetobutylicum ATCC 824 C. acetobutylicum DSM 792 C. paraputrificum M-21 C. tyrobutyricum JM1 C. tyrobutyricum ATCC 25755 C. perfringens SM101 C. cellulolyticum H10 | spo0A, hydA, adhE2, adC, groESL, txeR, tcdC, pdc-adhII | [ |
| 反义RNA技术 | 致死率低,易于筛选 | 仅在转录水平调控 | C. acetobutylicum ATCC 824 | ptb, buk, CoAT | [ |
| 转座子突变技术 | 可用于建立突变体库 | 难以控制插入位点 | C. difficile CD37 C. perfringens Strain 13 | random | [ |
| 二型内含子技术 | 操作简单,几乎适用于所有梭菌 | 脱靶概率较大,存在极性效应 | C. acetobutylicum ATCC 824 C. beijerinckii NCIMB 8052 | glcG, cbei2385, xylR, bdhA, bdhB, ptb, ack, adc | [ |
| 同源重组 | 可精确进行基因编辑 | 同源重组效率很低 | C. acetobutylicum NCIMB 8052 | gutD, spo0A | [ |
| 反筛标记介导的等位基因替换 | 相比纯粹的等位基因替换,效率有所提升 | 受制于第一次单交换效率 | C. acetobutylicum ATCC 824 | adh | [ |
| Ⅰ-SceⅠ归巢内切酶介导的等位基因替换 | 适用范围广,在革兰氏阳性和阴性菌中均可使用 | 设计复杂,操作烦琐,周期长 | C. acetobutylicum ATCC 824 C. beijerinckii NCIMB 8052 | adc, glcG, xylR | [ |
| 噬菌体丝氨酸重组酶介导的位点特异性基因编辑技术 | 适用于大片段DNA快速整合到宿主染色体上 | 受限于附着位点attb/p,应用范围小 | C. ljungdahlii DSM 13528 | thl, crt, bcd, etfB, etfA, hbd, ptb, buk | [ |
| Red/ET重组酶介导的同源重组 | 所需同源臂较短、不受限制性酶切位点限制 | 尚不完善 | C. acetobutylicum SMB009 | ermC | [ |
| CRISPR/Cas9系统介导的基因编辑 | 提高同源重组效率,可实现精确编辑 | 毒性大,很难获得转化子 | C. acetobutylicum DSM792 C. saccharoperbutylacetonicum N1-4 C. cellulovorans C. autoethanogenum DSM10061 | hprK, cac1502, pta, buk, clocel2243, adhE1, ctf,pyrE, 2,3-bdh | [ |
| CRISPR/Cas9n系统介导的基因编辑 | 相比于CRISPR/Cas9毒性有所降低,转化子数目增加 | 仍有毒性,转化子依然偏少 | C. cellulolyticum H10 C. acetobutylicum ATCC 824 C. beijerinckii NCIMB 8052 | pyrF, spo0A | [ |
| CRISPR/dCas9系统介导的基因表达下调 | 相比于CRISPR/Cas9毒性大大降低,易于得到转化子 | 依赖于sgRNA和基因;仅在转录水平调控 | C. pasteurianum ATCC 6013 C. acetobutylicum DSM792 | hprK, glpX | [ |
| 1 | 刘朝全, 姜学峰, 戴家权, 等. 疫情促变局 转型谋发展——2020年国内外油气行业发展概述及2021年展望[J]. 国际石油经济, 2021, 29(1): 28-37. |
| LIU C Q, JIANG X F, DAI J Q, et al. Overview of the domestic and foreign oil and gas industry development in 2020 and outlook for 2021[J]. International Petroleum Economics, 2021, 29(1): 28-37. | |
| 2 | 史硕博, 孟琼宇, 乔玮博, 等. 塑造低碳经济的第三代固碳生物炼制[J]. 合成生物学, 2020, 1(1): 44-59. |
| SHI S B, MENG Q Y, QIAO W B, et al. Establishing carbon dioxide-based third-generation biorefinery for a sustainable low-carbon economy[J]. Synthetic Biology Journal, 2020, 1(1): 44-59. | |
| 3 | PAPOUTSAKIS E T. Engineering solventogenic clostridia[J]. Current Opinion in Biotechnology, 2008, 19(5): 420-429. |
| 4 | PYNE M E, BRUDER M, MOO-YOUNG M, et al. Technical guide for genetic advancement of underdeveloped and intractable Clostridium [J]. Biotechnology Advances, 2014, 32(3): 623-641. |
| 5 | JONES D T, WOODS D R. Acetone-butanol fermentation revisited[J]. Microbiological Reviews, 1986, 50(4): 484-524. |
| 6 | TRACY B P, JONES S W, FAST A G, et al. Clostridia: the importance of their exceptional substrate and metabolite diversity for biofuel and biorefinery applications[J]. Current Opinion in Biotechnology, 2012, 23(3): 364-381. |
| 7 | VAN MELLAERT L, BARBÉ S, ANNÉ J. Clostridium spores as anti-tumour agents[J]. Trends in Microbiology, 2006, 14(4): 190-196. |
| 8 | ARORA R, BEHERA S, SHARMA N K, et al. Evaluating the pathway for co-fermentation of glucose and xylose for enhanced bioethanol production using flux balance analysis[J]. Biotechnology and Bioprocess Engineering, 2019, 24(6): 924-933. |
| 9 | BANERJEE S, MISHRA G, ROY A. Metabolic engineering of bacteria for renewable bioethanol production from cellulosic biomass[J]. Biotechnology and Bioprocess Engineering, 2019, 24(5): 713-733. |
| 10 | CHOI Y Y, HONG M E, CHANG W S, et al. Autotrophic biodiesel production from the thermotolerant microalga Chlorella sorokiniana by enhancing the carbon availability with temperature adjustment[J]. Biotechnology & Bioprocess Engineering, 2019, 24(1): 223-231. |
| 11 | KWON S W, PAARI K A, MALAVIYA A, et al. Synthetic biology tools for genome and transcriptome engineering of solventogenic Clostridium [J]. Frontiers in Bioengineering and Biotechnology, 2020, 8: 282. |
| 12 | ALEXANDRO E T, ZHANG F. Genome editing using Cas9 nickases[M]. Methods in Enzymology. Volume 546. Elsevier, 2014: 161-174. |
| 13 | 刘家宇, 杨智晗, 杨蕾, 等. 合成生物技术驱动酪丁酸梭菌细胞工厂开发的研究进展[J].合成生物学, 2022, 3(6): 1174. |
| LIU J Y, YANG Z H, YANG L, et al. Advances in the development of Clostridium tyrobutyricum cell factories driven by synthetic biotechnologies[J]. Synthetic Biology Journal, 2022, 3(6): 1174. | |
| 14 | PYNE M E, MOO-YOUNG M, CHUNG D A, et al. Development of an electrotransformation protocol for genetic manipulation of Clostridium pasteurianum [J]. Biotechnology for Biofuels, 2013, 6(1): 50. |
| 15 | MERMELSTEIN L D, WELKER N E, BENNETT G N, et al. Expression of cloned homologous fermentative genes in Clostridium acetobutylicum ATCC 824[J]. Nature Biotechnology, 1992, 10(2): 190-195. |
| 16 | JENNERT K C B, TARDIF C, YOUNG D I, et al. Gene transfer to Clostridium cellulolyticum ATCC 35319[J]. Microbiology, 2000, 146 (12): 3071-3080. |
| 17 | PURDY D, O'KEEFFE T A T, ELMORE M, et al. Conjugative transfer of clostridial shuttle vectors from Escherichia coli to Clostridium difficile through circumvention of the restriction barrier[J]. Molecular Microbiology, 2002, 46(2): 439-452. |
| 18 | RICHARDS D F, LINNETT P E, OULTRAM J D, et al. Restriction endonucleases in Clostridium pasteurianum ATCC 6013 and Clostridium thermohydrosulfuricum DSM 568[J]. Journal of general microbiology, 1988, 134(12): 3151-3157. |
| 19 | KLAPATCH T R, DEMAIN A L, LYND L R. Restriction endonuclease activity in Clostridium thermocellum and Clostridium thermosaccharolyticum [J]. Applied Microbiology & Biotechnology, 1996, 45(1/2): 127-131. |
| 20 | XUE C, CHENG C. Butanol production by Clostridium [M].Advances in Bioenergy. Volume 4. Amsterdam: Elsevier, 2019: 35-77. |
| 21 | MERMELSTEIN L D, PAPOUTSAKIS E T. In vivo methylation in Escherichia coli by the Bacillus subtilis phage-phi-3T-I methyltransferase to protect plasmids from restriction upon transformation of Clostridium acetobutylicum ATCC 824[J]. Applied & Environmental Microbiology, 1993, 59(4): 1077-1081. |
| 22 | MOLITOR B, KIRCHNER K, HENRICH A W, et al. Expanding the molecular toolkit for the homoacetogen Clostridium ljungdahlii [J]. Scientific Reports, 2016, 6: 31518. |
| 23 | DONG H J, ZHANG Y P, DAI Z J, et al. Engineering Clostridium strain to accept unmethylated DNA[J]. PLoS One, 2010, 5(2): e9038. |
| 24 | GONZÁLEZ-PAJUELO M, MEYNIAL-SALLES I, MENDES F, et al. Microbial conversion of glycerol to 1,3-propanediol: physiological comparison of a natural producer, Clostridium butyricum VPI 3266, and an engineered strain, Clostridium acetobutylicum DG1(pSPD5)[J]. Applied and Environmental Microbiology, 2006, 72(1): 96-101. |
| 25 | HEAP J T, PENNINGTON O J, CARTMAN S T, et al. The ClosTron: a universal gene knock-out system for the genus Clostridium [J]. Journal of Microbiological Methods, 2007, 70(3): 452-464. |
| 26 | LÜTKE-EVERSLOH T. Application of new metabolic engineering tools for Clostridium acetobutylicum [J]. Applied Microbiology and Biotechnology, 2014, 98(13): 5823-5837. |
| 27 | SILLERS R, CHOW A, TRACY B, et al. Metabolic engineering of the non-sporulating, non-solventogenic Clostridium acetobutylicum strain M5 to produce butanol without acetone demonstrate the robustness of the acid-formation pathways and the importance of the electron balance[J]. Metabolic Engineering, 2008, 10(6): 321-332. |
| 28 | ALSAKER K V, SPITZER T R, PAPOUTSAKIS E T. Transcriptional analysis of spo0A overexpression in Clostridium acetobutylicum and its effect on the cell's response to butanol stress[J]. Journal of Bacteriology, 2004, 186(7): 1959-1971. |
| 29 | GIRBAL L, VON ABENDROTH G, WINKLER M, et al. Homologous and heterologous overexpression in Clostridium acetobutylicum and characterization of purified clostridial and algal Fe-only hydrogenases with high specific activities[J]. Applied and Environmental Microbiology, 2005, 71(5): 2777-2781. |
| 30 | TOMAS C A, WELKER N E, PAPOUTSAKIS E T. Overexpression of groESL in Clostridium acetobutylicum results in increased solvent production and tolerance, prolonged metabolism, and changes in the cell's transcriptional program[J]. Applied and Environmental Microbiology, 2003, 69(8): 4951-4965. |
| 31 | YU M R, ZHANG Y L, TANG I C, et al. Metabolic engineering of Clostridium tyrobutyricum for n-butanol production[J]. Metabolic Engineering, 2011, 13(4): 373-382. |
| 32 | KOVÁCS K, WILLSON B J, SCHWARZ K, et al. Secretion and assembly of functional mini-cellulosomes from synthetic chromosomal operons in Clostridium acetobutylicum ATCC 824[J]. Biotechnology for Biofuels, 2013, 6(1): 117. |
| 33 | YU M R, DU Y M, JIANG W Y, et al. Effects of different replicons in conjugative plasmids on transformation efficiency, plasmid stability, gene expression and n-butanol biosynthesis in Clostridium tyrobutyricum [J]. Applied Microbiology and Biotechnology, 2012, 93(2): 881-889. |
| 34 | MANI N, DUPUY B. Regulation of toxin synthesis in Clostridium difficile by an alternative RNA polymerase sigma factor[J]. Proceedings of the National Academy of Sciences of the United States of America, 2001, 98(10): 5844-5849. |
| 35 | MANI N, LYRAS D, BARROSO L, et al. Environmental response and autoregulation of Clostridium difficile TxeR, a sigma factor for toxin gene expression[J]. Journal of Bacteriology, 2002, 184(21): 5971-5978. |
| 36 | LEE J Y, JANG Y S, LEE J, et al. Metabolic engineering of Clostridium acetobutylicum M5 for highly selective butanol production[J]. Biotechnology Journal, 2009, 4(10): 1432-1440. |
| 37 | MATAMOUROS S, ENGLAND P, DUPUY B. Clostridium difficile toxin expression is inhibited by the novel regulator TcdC [J]. Molecular Microbiology, 2007, 64(5): 1274-1288. |
| 38 | GUEDON E, DESVAUX M, PETITDEMANGE H. Improvement of cellulolytic properties of Clostridium cellulolyticum by metabolic engineering[J]. Applied and Environmental Microbiology, 2002, 68(1): 53-58. |
| 39 | DESAI R P, PAPOUTSAKIS E T. Antisense RNA strategies for metabolic engineering of Clostridium acetobutylicum [J]. Applied and Environmental Microbiology, 1999, 65(3): 936-945. |
| 40 | TUMMALA S B, WELKER N E, PAPOUTSAKIS E T. Design of antisense RNA constructs for downregulation of the acetone formation pathway of Clostridium acetobutylicum [J]. Journal of Bacteriology, 2003, 185(6): 1923-1934. |
| 41 | SILLERS R, AL-HINAI M A, PAPOUTSAKIS E T. Aldehyde-alcohol dehydrogenase and/or thiolase overexpression coupled with CoA transferase downregulation lead to higher alcohol titers and selectivity in Clostridium acetobutylicumfermentations[J]. Biotechnology and Bioengineering, 2009, 102(1): 38-49. |
| 42 | VIDAL J E, CHEN J M, LI J H, et al. Use of an EZ-Tn5-based random mutagenesis system to identify a novel toxin regulatory locus in Clostridium perfringens strain 13[J]. PLoS One, 2009, 4(7): e6232. |
| 43 | MULLANY P, WILKS M, TABAQCHALI S. Transfer of Tn916 and Tn916 ΔE into Clostridium difficile: demonstration of a hot-spot for these elements in the C. difficile genome[J]. FEMS Microbiology Letters, 1991, 63(2/3): 191-194. |
| 44 | XIAO H, GU Y, NING Y Y, et al. Confirmation and elimination of xylose metabolism bottlenecks in glucose phosphoenolpyruvate-dependent phosphotransferase system-deficient Clostridium acetobutylicum for simultaneous utilization of glucose, xylose, and arabinose[J]. Applied and Environmental Microbiology, 2011, 77(22): 7886-7895. |
| 45 | XIAO H, LI Z L, JIANG Y, et al. Metabolic engineering of D-xylose pathway in Clostridium beijerinckii to optimize solvent production from xylose mother liquid[J]. Metabolic Engineering, 2012, 14(5): 569-578. |
| 46 | GU Y, DING Y, REN C, et al. Reconstruction of xylose utilization pathway and regulons in firmicutes[J]. BMC Genomics, 2010, 11: 255. |
| 47 | COOKSLEY C M, ZHANG Y, WANG H Z, et al. Targeted mutagenesis of the Clostridium acetobutylicum acetone-butanol-ethanol fermentation pathway[J]. Metabolic Engineering, 2012, 14(6): 630-641. |
| 48 | WILKINSON S R, YOUNG M. Targeted integration of genes into the Clostridium acetobutylicum chromosome[J]. Microbiology, 1994, 140(1): 89-95. |
| 49 | HEAP J T, EHSAAN M, COOKSLEY C M, et al. Integration of DNA into bacterial chromosomes from plasmids without a counter-selection marker[J]. Nucleic Acids Research, 2012, 40(8): e59. |
| 50 | ZHANG N, SHAO L J, JIANG Y, et al. I-SceI-mediated scarless gene modification via allelic exchange in Clostridium [J]. Journal of Microbiological Methods, 2015, 108: 49-60. |
| 51 | HUANG H, CHAI C S, YANG S, et al. Phage serine integrase-mediated genome engineering for efficient expression of chemical biosynthetic pathway in gas-fermenting Clostridium ljungdahlii [J]. Metabolic Engineering, 2019, 52: 293-302. |
| 52 | DONG H J, TAO W W, GONG F Y, et al. A functional recT gene for recombineering of Clostridium [J]. Journal of Biotechnology, 2014, 173: 65-67. |
| 53 | BRUDER M R, PYNE M E, MOO-YOUNG M, et al. Extending CRISPR-Cas9 technology from genome editing to transcriptional engineering in the genus Clostridium [J]. Applied and Environmental Microbiology, 2016, 82(20): 6109-6119. |
| 54 | WANG S H, DONG S, WANG P X, et al. Genome editing in Clostridium saccharoperbutylacetonicum N1-4 with the CRISPR-Cas9 system[J]. Applied & Environmental Microbiology, 2017, 83(10): e00233-e00217. |
| 55 | WEN Z Q, MINTON N P, ZHANG Y, et al. Enhanced solvent production by metabolic engineering of a twin-clostridial consortium[J]. Metabolic Engineering, 2017, 39: 38-48. |
| 56 | HUANG H, CHAI C S, LI N, et al. CRISPR/Cas9-based efficient genome editing in Clostridium ljungdahlii, an autotrophic gas-fermenting bacterium[J]. ACS Synthetic Biology, 2016, 5(12): 1355-1361. |
| 57 | NAGARAJU S, DAVIES N K, WALKER D J F, et al. Genome editing of Clostridium autoethanogenum using CRISPR/Cas9[J]. Biotechnology for Biofuels, 2016, 9: 219. |
| 58 | LI Q, CHEN J, MINTON N P, et al. CRISPR-based genome editing and expression control systems in Clostridium acetobutylicum and Clostridium beijerinckii [J]. Biotechnology Journal, 2016, 11(7): 961-972. |
| 59 | XU T, LI Y C, SHI Z, et al. Efficient genome editing in Clostridium cellulolyticum via CRISPR-Cas9 nickase[J]. Applied and Environmental Microbiology, 2015, 81(13): 4423-4431. |
| 60 | DORNENBURG J E, DEVITA A M, PALUMBO M J, et al. Widespread antisense transcription in Escherichia coli [J]. mBio, 2010, 1(1): e00024-e00010. |
| 61 | 马欣, 高学文. 转座子随机突变芽孢杆菌的研究进展[J]. 中国生物防治学报, 2015, 31(3): 394-403. |
| MA X, GAO X W. Research progress of random mutagenesis of bacillus by use of transposon[J]. Chinese Journal of Biological Control, 2015, 31(3): 394-403. | |
| 62 | 顾阳, 杨晟, 姜卫红. 产溶剂梭菌分子遗传操作技术研究进展[J]. 生物工程学报, 2013, 29(8): 1133-1145. |
| GU Y, YANG S, JIANG W H. Development in molecular genetic manipulation of solventogenic clostridia[J]. Chinese Journal of Biotechnology, 2013, 29(8): 1133-1145. | |
| 63 | LIU H L, BOUILLAUT L, SONENSHEIN A L, et al. Use of a mariner-based transposon mutagenesis system to isolate Clostridium perfringens mutants deficient in gliding motility[J]. Journal of Bacteriology, 2013, 195(3): 629-636. |
| 64 | LANCKRIET A, TIMBERMONT L, HAPPONEN L J, et al. Generation of single-copy transposon insertions in Clostridium perfringens by electroporation of phage mu DNA transposition complexes[J]. Applied and Environmental Microbiology, 2009, 75(9): 2638-2642. |
| 65 | CARTMAN S T, MINTON N P. A mariner-based transposon system for in vivo random mutagenesis of Clostridium difficile [J]. Applied and Environmental Microbiology, 2010, 76(4): 1103-1109. |
| 66 | WANG H, ROBERTS A P, LYRAS D, et al. Characterization of the ends and target sites of the novel conjugative transposon Tn5397 from Clostridium difficile: excision and circularization is mediated by the large resolvase, TndX[J]. Journal of Bacteriology, 2000, 182(13): 3775-3783. |
| 67 | ZHANG Y, XU S, CHAI C S, et al. Development of an inducible transposon system for efficient random mutagenesis in Clostridium acetobutylicum [J]. FEMS Microbiology Letters, 2016, 363(8): fnw065. |
| 68 | KUEHNE S A, MINTON N P. ClosTron-mediated engineering of Clostridium [J]. Bioengineered, 2012, 3(4): 247-254. |
| 69 | SHAO L J, HU S Y, YANG Y, et al. Targeted gene disruption by use of a group II intron (targetron) vector in Clostridium acetobutylicum [J]. Cell Research, 2007, 17(11): 963-965. |
| 70 | 闻志强, 孙小曼, 汪庆卓, 等. 梭菌正丁醇代谢工程研究进展[J]. 合成生物学, 2021, 2(2): 194-221. |
| WEN Z Q, SUN X M, WANG Q Z, et al. Recent advances in metabolic engineering of clostridia for n-butanol production[J]. Synthetic Biology Journal, 2021, 2(2): 194-221. | |
| 71 | HÖNICKE D, LÜTKE-EVERSLOH T, LIU Z Y, et al. Chemostat cultivation and transcriptional analyses of Clostridium acetobutylicum mutants with defects in the acid and acetone biosynthetic pathways[J]. Applied Microbiology and Biotechnology, 2014, 98(23): 9777-9794. |
| 72 | LIU J, GUO T, WANG D, et al. Enhanced butanol production by increasing NADH and ATP levels in Clostridium beijerinckii NCIMB 8052 by insertional inactivation of Cbei_4110 [J]. Applied Microbiology and Biotechnology, 2016, 100(11): 4985-4996. |
| 73 | JANG Y S, IM J A, CHOI S Y, et al. Metabolic engineering of Clostridium acetobutylicum for butyric acid production with high butyric acid selectivity[J]. Metabolic Engineering, 2014, 23: 165-174. |
| 74 | WEN Z Q, LU M R, LEDESMA-AMARO R, et al. TargeTron technology applicable in solventogenic clostridia: revisiting 12 years' advances[J]. Biotechnology Journal, 2020, 15(1): 1900284. |
| 75 | LAMBOWITZ A M, ZIMMERLY S. Group II introns: mobile ribozymes that invade DNA[J]. Cold Spring Harbor Perspectives in Biology, 2011, 3(8): a003616. |
| 76 | HEAP J T, KUEHNE S A, EHSAAN M, et al. The ClosTron: mutagenesis in Clostridium refined and streamlined[J]. Journal of Microbiological Methods, 2010, 80(1): 49-55. |
| 77 | ARGYROS D A, TRIPATHI S A, BARRETT T F, et al. High ethanol titers from cellulose by using metabolically engineered thermophilic, anaerobic microbes[J]. Applied and Environmental Microbiology, 2011, 77(23): 8288-8294. |
| 78 | KOHJI K, NORIKO K T, HIROSHI Y, et al. Involvement of RecE exonuclease and RecT annealing protein in DNA double-strand break repair by homologous recombination[J]. Gene, 1994, 138(1/2): 17-25. |
| 79 | HORVATH P, BARRANGOU R. CRISPR/Cas, the immune system of bacteria and archaea[J]. Science, 2010, 327(5962): 167-170. |
| 80 | JANSEN R, VAN EMBDEN J D A, GAASTRA W, et al. Identification of genes that are associated with DNA repeats in prokaryotes[J]. Molecular Microbiology, 2002, 43(6): 1565-1575. |
| 81 | MALI P, YANG L H, ESVELT K M, et al. RNA-guided human genome engineering via Cas9[J]. Science, 2013, 339(6121): 823-826. |
| 82 | CONG L, RAN F A, COX D, et al. Multiplex genome engineering using CRISPR/Cas systems[J]. Science, 2013, 339(6121): 819-823. |
| 83 | WANG Y, ZHANG Z T, SEO S O, et al. Markerless chromosomal gene deletion in Clostridium beijerinckii using CRISPR/Cas9 system[J]. Journal of Biotechnology, 2015, 200: 1-5. |
| 84 | WANG Y, ZHANG Z T, SEO S O, et al. Bacterial genome editing with CRISPR-Cas9: deletion, integration, single nucleotide modification, and desirable "clean" mutant selection in Clostridium beijerinckii as an example[J]. ACS Synthetic Biology, 2016, 5(7): 721-732. |
| 85 | BERNHEIM A, CALVO-VILLAMAÑÁN A, BASIER C, et al. Inhibition of NHEJ repair by type II-A CRISPR-Cas systems in bacteria[J]. Nature Communications, 2017, 8: 2094. |
| 86 | CUI L, BIKARD D. Consequences of Cas9 cleavage in the chromosome of Escherichia coli [J]. Nucleic Acids Research, 2016, 44(9): 4243-4251. |
| 87 | SHUMAN S, GLICKMAN M S. Bacterial DNA repair by non-homologous end joining[J]. Nature Reviews Microbiology, 2007, 5(11): 852-861. |
| 88 | XU T, LI Y C, HE Z L, et al. Cas9 nickase-assisted RNA repression enables stable and efficient manipulation of essential metabolic genes in Clostridium cellulolyticum [J]. Frontiers in Microbiology, 2017, 8: 1744. |
| 89 | STRECKER J, LADHA A, GARDNER Z, et al. RNA-guided DNA insertion with CRISPR-associated transposases[J]. Science, 2019, 365(6448): 48-53. |
| 90 | PYNE M E, BRUDER M R, MOO-YOUNG M, et al. Harnessing heterologous and endogenous CRISPR-Cas machineries for efficient markerless genome editing in Clostridium [J]. Scientific Reports, 2016, 6: 25666. |
| 91 | ZHANG J, ZONG W M, HONG W, et al. Exploiting endogenous CRISPR-Cas system for multiplex genome editing in Clostridium tyrobutyricum and engineer the strain for high-level butanol production[J]. Metabolic Engineering, 2018, 47: 49-59. |
| 92 | ZHOU X Q, WANG X L, LUO H Y, et al. Exploiting heterologous and endogenous CRISPR-Cas systems for genome editing in the probiotic Clostridium butyricum [J]. Biotechnology and Bioengineering, 2021, 118(7): 2448-2459. |
| 93 | WENTZ T G, TREMBLAY B J M, BRADSHAW M, et al. Endogenous CRISPR-Cas systems in group I Clostridium botulinum and Clostridium sporogenes do not directly target the botulinum neurotoxin gene cluster[J]. Frontiers in Microbiology, 2021, 12: 787726. |
| 94 | WALKER J E, LANAHAN A A, ZHENG T Y, et al. Development of both type I-B and type II CRISPR/Cas genome editing systems in the cellulolytic bacterium Clostridium thermocellum [J]. Metabolic Engineering Communications, 2019, 10: eoo116. |
| 95 | HONG W, ZHANG J, CUI G Z, et al. Multiplexed CRISPR-Cpf1-mediated genome editing in Clostridium difficile toward the understanding of pathogenesis of C. difficile infection[J]. ACS Synthetic Biology, 2018, 7(6): 1588-1600. |
| 96 | MERRICK C A, ZHAO J, ROSSER S J. Serine integrases: advancing synthetic biology[J]. ACS Synthetic Biology, 2018, 7(2): 299-310. |
| 97 | BAI H, SUN M X, GHOSH P, et al. Single-molecule analysis reveals the molecular bearing mechanism of DNA strand exchange by a serine recombinase[J]. Proceedings of the National Academy of Sciences of the United States of America, 2011, 108(18): 7419-7424. |
| 98 | 田雨顺, 罗鹏, 刘秋婷, 等. 细菌重组系统及其应用研究进展[J]. 微生物学通报, 2017, 44(2): 473-482. |
| TIAN Y S, LUO P, LIU Q T, et al. Progress on bacterial recombination systems and their application[J]. Microbiology China, 2017, 44(2): 473-482. | |
| 99 | IYER L M, KOONIN E V, ARAVIND L. Classification and evolutionary history of the single-strand annealing proteins, RecT, Redβ, ERF and RAD52[J]. BMC Genomics, 2002, 3: 8. |
| 100 | 王雪. 伯克氏菌重组系统的研究与应用[D]. 济南:山东大学, 2018. |
| WANG X. The research and application of recombineering system in Burkholderiales species[D]. Jinan: Shandong University, 2018. | |
| 101 | SHARAN S K, THOMASON L C, KUZNETSOV S G, et al. Recombineering: a homologous recombination-based method of genetic engineering[J]. Nature Protocols, 2009, 4(2): 206-223. |
| 102 | REISCH C R, PRATHER K L J. The no-SCAR (Scarless Cas9 Assisted Recombineering) system for genome editing in Escherichia coli [J]. Scientific Reports, 2015, 5: 15096. |
| 103 | WANG H H, ISAACS F J, CARR P A, et al. Programming cells by multiplex genome engineering and accelerated evolution[J]. Nature, 2009, 460(7257): 894-898. |
| 104 | WARNER J R, REEDER P J, KARIMPOUR-FARD A, et al. Rapid profiling of a microbial genome using mixtures of barcoded oligonucleotides[J]. Nature Biotechnology, 2010, 28(8): 856-862. |
| 105 | VAN KESSEL J C, HATFULL G F. Recombineering in Mycobacterium tuberculosis [J]. Nature Methods, 2007, 4(2): 147-152. |
| 106 | VAN KESSEL J C, HATFULL G F. Efficient point mutagenesis in mycobacteria using single-stranded DNA recombineering: characterization of antimycobacterial drug targets[J]. Molecular Microbiology, 2008, 67(5): 1094-1107. |
| 107 | VAN PIJKEREN J P, BRITTON R A. High efficiency recombineering in lactic acid bacteria[J]. Nucleic Acids Research, 2012, 40(10): e76. |
| [1] | DONG Ying, MA Mengdan, HUANG Weiren. Progress in the miniaturization of CRISPR-Cas systems [J]. Synthetic Biology Journal, 2025, 6(1): 105-117. |
| [2] | CHEN Ziling, XIANG Yangfei. Integrated development of organoid technology and synthetic biology [J]. Synthetic Biology Journal, 2024, 5(4): 795-812. |
| [3] | CHEN Yingying, LIU Yang, SHI Junjie, MA Junying, JU Jianhua. CRISPR/Cas systems and their applications in gene editing with filamentous fungi [J]. Synthetic Biology Journal, 2024, 5(3): 672-693. |
| [4] | HUI Zhen, TANG Xiaoyu. Applications of the CRISPR/Cas9 editing system in the study of microbial natural products [J]. Synthetic Biology Journal, 2024, 5(3): 658-671. |
| [5] | DU Yao, GAO Hongdan, LIU Jiakun, LIU Xiaorong, XING Zhihao, ZHANG Tao, MA Dongli. Research progress of the CRISPR-Cas system in the detecting pathogen nucleic acids [J]. Synthetic Biology Journal, 2024, 5(1): 202-216. |
| [6] | CHEN Yaru, CAO Yingxiu, SONG Hao. Advances and applications of gene editing and transcriptional regulation in electroactive microorganisms [J]. Synthetic Biology Journal, 2023, 4(6): 1281-1299. |
| [7] | WANG Tiantian, ZHU Hong, YANG Chen. Development of CRISPRa for metabolic engineering applications in cyanobacteria [J]. Synthetic Biology Journal, 2023, 4(4): 824-839. |
| [8] | MA Mengdan, SHANG Mengyu, LIU Yuchen. Application and prospect of CRISPR-Cas9 system in tumor biology [J]. Synthetic Biology Journal, 2023, 4(4): 703-719. |
| [9] | LIN Jicong, ZOU Gen, LIU Hongmin, WEI Yongjun. Application of CRISPR/Cas genome editing technology in the synthesis of secondary metabolites of filamentous fungi [J]. Synthetic Biology Journal, 2023, 4(4): 738-755. |
| [10] | MA Mengdan, LIU Yuchen. Potential application of synthetic biology in disease information recording and real-time monitoring [J]. Synthetic Biology Journal, 2023, 4(2): 301-317. |
| [11] | LV Hailong, WANG Jian, LV Hao, WANG Jin, XU Yong, GU Dayong. Synthetic biology for next-generation genetic diagnostics [J]. Synthetic Biology Journal, 2023, 4(2): 318-332. |
| [12] | LIU Ke, LIN Guihong, LIU Kun, ZHOU Wei, WANG Fengqing, WEI Dongzhi. Mining, engineering and functional expansion of CRISPR/Cas systems [J]. Synthetic Biology Journal, 2023, 4(1): 47-66. |
| [13] | PAN Yingjia, XIA Siyang, DONG Chang, CAI Jin, LIAN Jiazhang. Mutator-driven continuous genome evolution of Saccharomyces cerevisiae [J]. Synthetic Biology Journal, 2023, 4(1): 225-240. |
| [14] | TENG Xiaolong, SHI Shuobo. Optimization and development of CRISPR/Cas9 systems for genome editing [J]. Synthetic Biology Journal, 2023, 4(1): 67-85. |
| [15] | LIANG Liya, LIU Rongming. Protein engineering of DNA targeting type Ⅱ CRISPR/Cas systems [J]. Synthetic Biology Journal, 2023, 4(1): 86-101. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||