Synthetic Biology Journal ›› 2022, Vol. 3 ›› Issue (4): 658-675.DOI: 10.12211/2096-8280.2021-069
• Invited Review • Previous Articles Next Articles
Application of cell-free synthesis strategy in biomaterial research
JI Botao, QIAN Zhigang, XIA Xiaoxia
- State Key Laboratory of Microbial Metabolism,Joint International Research Laboratory of Metabolic and Developmental Sciences,School of Life Science & Biotechnology,Shanghai Jiao Tong University,Shanghai 200240,China
-
Received:2021-06-30Revised:2021-07-22Online:2022-09-08Published:2022-08-31 -
Contact:XIA Xiaoxia
无细胞合成策略在生物材料研究中的应用
吉博涛, 钱志刚, 夏小霞
- 上海交通大学生命科学技术学院,微生物代谢国家重点实验室,教育部代谢与发育科学国际合作联合实验室,上海 200240
-
通讯作者:夏小霞 -
作者简介:吉博涛 (1997—),男,博士研究生。研究方向为无细胞表达系统合成生物材料。E-mail: botao-ji@sjtu.edu.cn夏小霞 (1978—),女,教授,博士生导师。研究方向为蛋白材料合成生物学、人工细胞器、生物大分子自组装等。E-mail: xiaoxiaxia@sjtu.edu.cn -
基金资助:国家重点研发计划(2020YFA0907702);国家自然科学基金(32071414);上海市自然科学基金(21ZR1432100)
CLC Number:
Cite this article
JI Botao, QIAN Zhigang, XIA Xiaoxia. Application of cell-free synthesis strategy in biomaterial research[J]. Synthetic Biology Journal, 2022, 3(4): 658-675.
吉博涛, 钱志刚, 夏小霞. 无细胞合成策略在生物材料研究中的应用[J]. 合成生物学, 2022, 3(4): 658-675.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2021-069
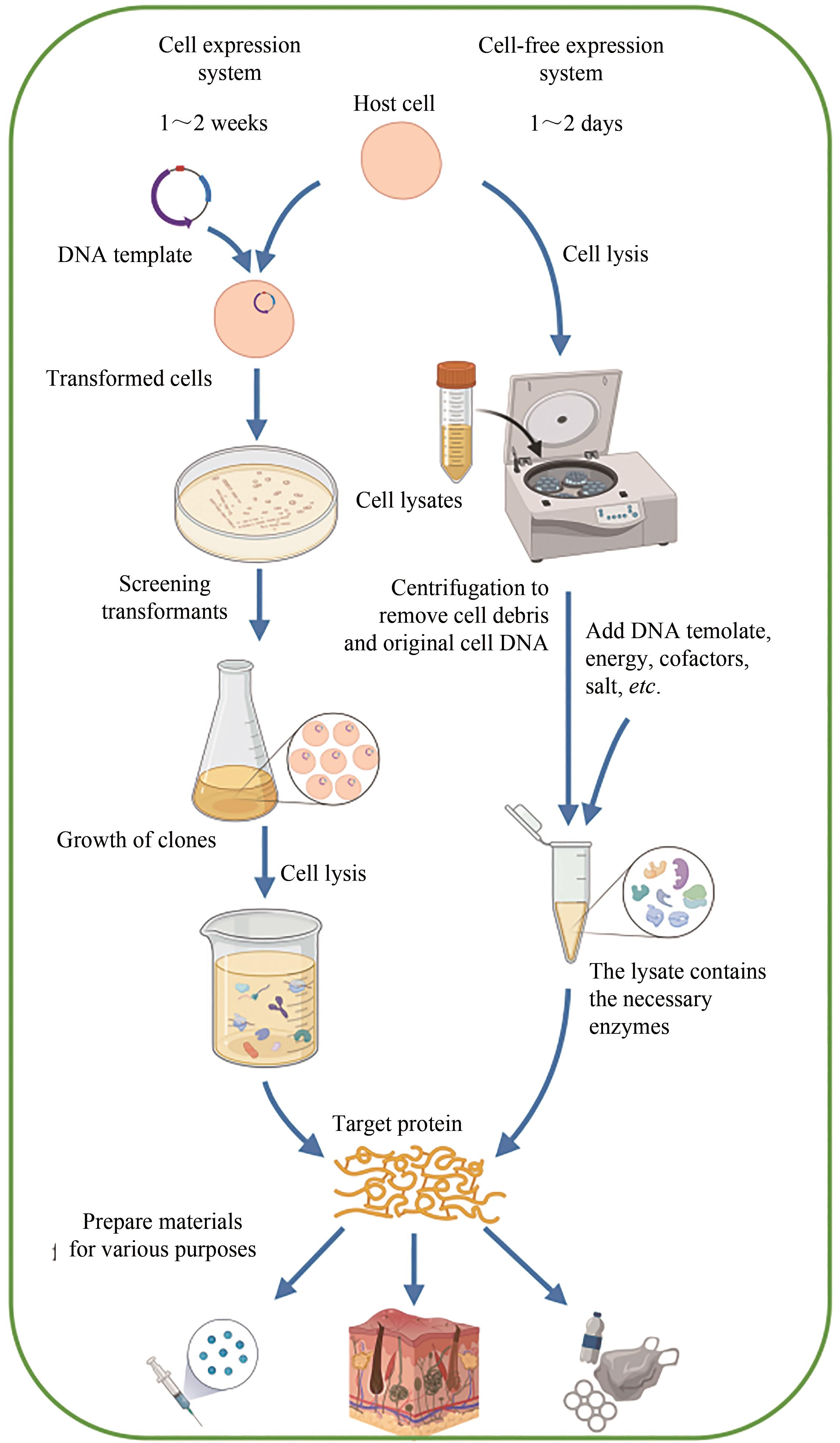
| 特征 | 细胞表达系统 | 无细胞表达系统 |
|---|---|---|
| 转录和翻译 | -具有细胞膜,细胞内表征和产物释放可能具有挑战性 -内源性调控难以预测和修改 +能够使用定向进化技术 | +开放的环境易于实时操作和控制 +直接添加底物或收获产物 +易于使用来自多种生物体的酶途径 |
| 翻译后修饰 | +简单 | -困难 |
| 自我复制 | +简单 | -困难 |
| DNA模板 | -质粒(或者整合到基因组的DNA) | +质粒或 PCR 产物 |
| 引入非天然氨基酸 | -困难 | +简单 |
| 膜蛋白和复合蛋白的合成 | -由于细胞内环境有限而难以合成 | +通过添加表面活性剂或调整系统环境易于合成 |
| 对毒性的耐受力 | -低,难以合成毒性蛋白 -若中间产物有毒性,则可行性受到限制 | +高,可以合成毒性蛋白 +无限制(如某些途径酶使用体积分数为12%异丁醇) |
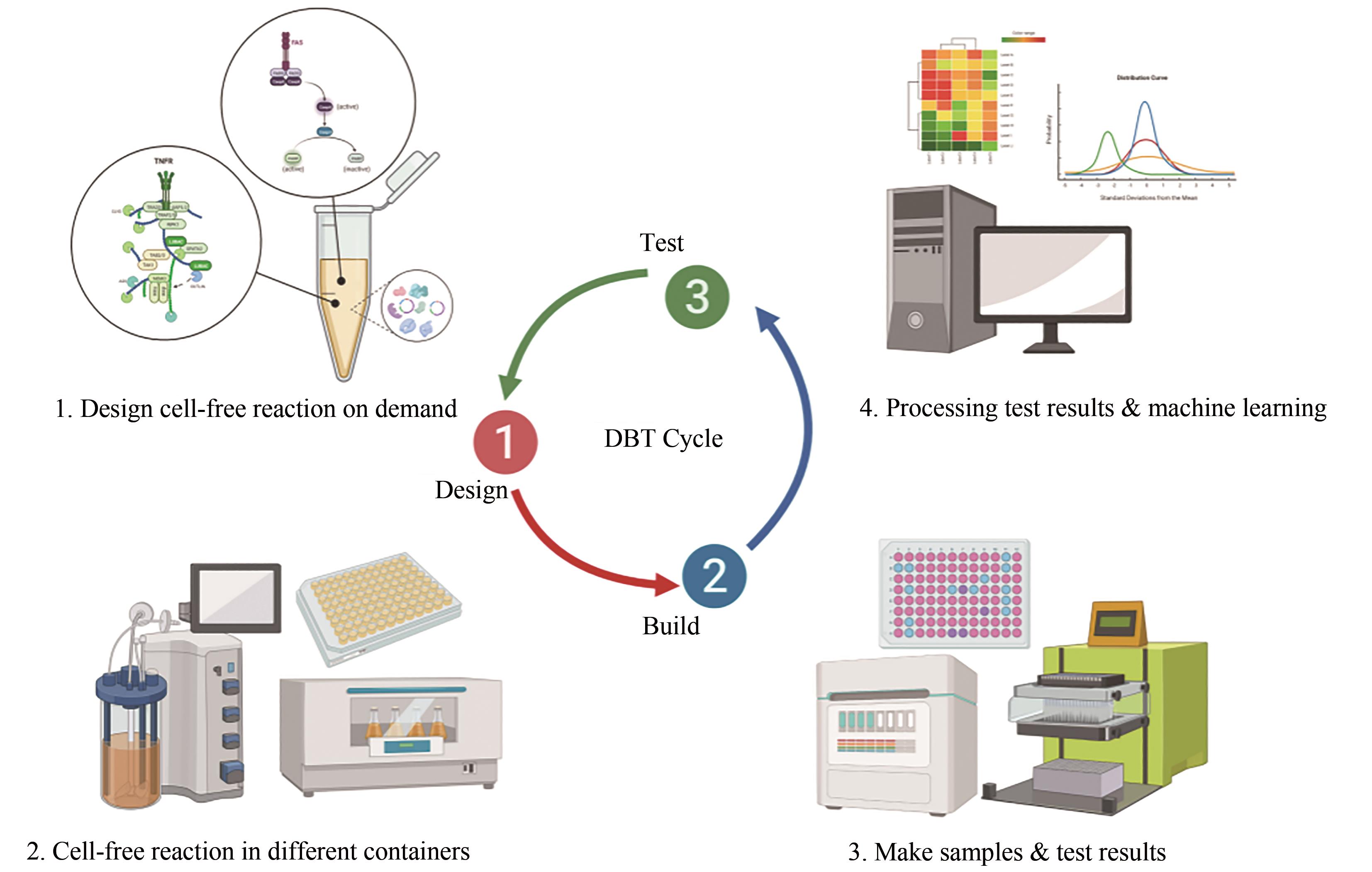
| 设计-构建-测试(DBT)循环 | -慢,1~2周 | +快,1~2天 |
| 资源利用率 | -较低,由于复杂的细胞代谢而将资源转向细胞维护和副产品 | +较高,所有资源都可以直接用于生产产品 |
| 与材料整合的能力 | -困难 | +简单 |
| 生物制造 | -适中的产率和产量 -纯化前需再次裂解细胞 -不易放大生产,工业规模发酵条件是不同的 -菌种污染可能会是灾难性的 +多年实践经验;完善的方法 | +较高的产率和产量 +无须细胞裂解的简单的纯化过程 +易于放大生产,具有线性放大的可能 -较为年轻,最近成立 |
| 成本 | +低到适中 | -中高,主要为酶和辅因子成本 |
| 稳定性 | -通过发酵条件影响细胞环境 | +由工程师直接控制反应条件 |
Tab. 1 The advantages and disadvantages of cell-free expression systems
| 特征 | 细胞表达系统 | 无细胞表达系统 |
|---|---|---|
| 转录和翻译 | -具有细胞膜,细胞内表征和产物释放可能具有挑战性 -内源性调控难以预测和修改 +能够使用定向进化技术 | +开放的环境易于实时操作和控制 +直接添加底物或收获产物 +易于使用来自多种生物体的酶途径 |
| 翻译后修饰 | +简单 | -困难 |
| 自我复制 | +简单 | -困难 |
| DNA模板 | -质粒(或者整合到基因组的DNA) | +质粒或 PCR 产物 |
| 引入非天然氨基酸 | -困难 | +简单 |
| 膜蛋白和复合蛋白的合成 | -由于细胞内环境有限而难以合成 | +通过添加表面活性剂或调整系统环境易于合成 |
| 对毒性的耐受力 | -低,难以合成毒性蛋白 -若中间产物有毒性,则可行性受到限制 | +高,可以合成毒性蛋白 +无限制(如某些途径酶使用体积分数为12%异丁醇) |
| 设计-构建-测试(DBT)循环 | -慢,1~2周 | +快,1~2天 |
| 资源利用率 | -较低,由于复杂的细胞代谢而将资源转向细胞维护和副产品 | +较高,所有资源都可以直接用于生产产品 |
| 与材料整合的能力 | -困难 | +简单 |
| 生物制造 | -适中的产率和产量 -纯化前需再次裂解细胞 -不易放大生产,工业规模发酵条件是不同的 -菌种污染可能会是灾难性的 +多年实践经验;完善的方法 | +较高的产率和产量 +无须细胞裂解的简单的纯化过程 +易于放大生产,具有线性放大的可能 -较为年轻,最近成立 |
| 成本 | +低到适中 | -中高,主要为酶和辅因子成本 |
| 稳定性 | -通过发酵条件影响细胞环境 | +由工程师直接控制反应条件 |
| 组成生物材料的特殊蛋白质 | 使CFE系统提高生产力和/或生物活性的方法 |
|---|---|
| 对生产宿主有毒的蛋白质 | 额外的tRNA补充[ 选择不同的提取物(例如昆虫)[ 连续交换无细胞表达系统[ 纯化标签以提高溶解度[ |
| 具有氨基酸组成偏好的蛋白质 | 稀有tRNA的添加[ 选择不同的提取物(例如小麦胚芽)[ 用同义密码子替换初始密码子[ |
| 需要分子伴侣或特定化学环境合成的蛋白质 | 伴侣蛋白,蛋白质二硫键异构酶(PDI),还原和氧化的谷胱甘肽(glutathione)调整[ 前导肽和微粒体囊泡[ 对于膜蛋白,加入洗涤剂/表面活性剂[ |
| 含有非天然氨基酸的蛋白质 | 通过基因组工程消除提取物中的负面效应物,如释放因子1(RF1)的竞争限制[ 去除同源tRNA[ |
| 需要糖基化修饰的蛋白质 | 内切糖苷酶介导的N-糖基化蛋白的制备[ 原核寡糖基转移酶(OST)介导的无细胞糖蛋白生物合成[ 糖基转移酶(GT)介导的蛋白质糖基化和聚糖加工[ |
| 需要磷酸化的蛋白质 | 基因组重新编码的大肠杆菌菌株[ |
Tab. 2 Corresponding methods for special protein synthesis using CFE system
| 组成生物材料的特殊蛋白质 | 使CFE系统提高生产力和/或生物活性的方法 |
|---|---|
| 对生产宿主有毒的蛋白质 | 额外的tRNA补充[ 选择不同的提取物(例如昆虫)[ 连续交换无细胞表达系统[ 纯化标签以提高溶解度[ |
| 具有氨基酸组成偏好的蛋白质 | 稀有tRNA的添加[ 选择不同的提取物(例如小麦胚芽)[ 用同义密码子替换初始密码子[ |
| 需要分子伴侣或特定化学环境合成的蛋白质 | 伴侣蛋白,蛋白质二硫键异构酶(PDI),还原和氧化的谷胱甘肽(glutathione)调整[ 前导肽和微粒体囊泡[ 对于膜蛋白,加入洗涤剂/表面活性剂[ |
| 含有非天然氨基酸的蛋白质 | 通过基因组工程消除提取物中的负面效应物,如释放因子1(RF1)的竞争限制[ 去除同源tRNA[ |
| 需要糖基化修饰的蛋白质 | 内切糖苷酶介导的N-糖基化蛋白的制备[ 原核寡糖基转移酶(OST)介导的无细胞糖蛋白生物合成[ 糖基转移酶(GT)介导的蛋白质糖基化和聚糖加工[ |
| 需要磷酸化的蛋白质 | 基因组重新编码的大肠杆菌菌株[ |
| 73 | TSUBOI T, TAKEO S, ARUMUGAM T U, et al. The wheat germ cell-free protein synthesis system: a key tool for novel malaria vaccine candidate discovery[J]. Acta Tropica, 2010, 114(3): 171-176. |
| 74 | AHN J H, KEUM J W, KIM D M. High-throughput, combinatorial engineering of initial codons for tunable expression of recombinant proteins[J]. Journal of Proteome Research, 2008, 7(5): 2107-2113. |
| 75 | PARK Y J, LEE K H, BAEK M S, et al. High-throughput engineering of initial coding regions for maximized production of recombinant proteins[J]. Biotechnology and Bioprocess Engineering, 2017, 22(5): 497-503. |
| 76 | STECH M, KUBICK S. Cell-free synthesis meets antibody production: a review[J]. Antibodies, 2015, 4(1): 12-33. |
| 77 | STECH M, NIKOLAEVA O, THORING L, et al. Cell-free synthesis of functional antibodies using a coupled in vitro transcription-translation system based on CHO cell lysates[J]. Scientific Reports, 2017, 7: 12030. |
| 78 | SACHSE R, DONDAPATI S K, FENZ S F, et al. Membrane protein synthesis in cell-free systems: from bio-mimetic systems to bio-membranes[J]. FEBS Letters, 2014, 588(17): 2774-2781. |
| 79 | BÄCKLUND E, IGNATUSHCHENKO M, LARSSON G. Suppressing glucose uptake and acetic acid production increases membrane protein overexpression in Escherichia coli [J]. Microbial Cell Factories, 2011, 10: 35. |
| 80 | MICHEL-REYDELLET N, CALHOUN K, SWARTZ J. Amino acid stabilization for cell-free protein synthesis by modification of the Escherichia coli genome[J]. Metabolic Engineering, 2004, 6(3): 197-203. |
| 81 | GAN Q L, FAN C G. Increasing the fidelity of noncanonical amino acid incorporation in cell-free protein synthesis[J]. Biochimica et Biophysica Acta (BBA)-General Subjects, 2017, 1861(11): 3047-3052. |
| 82 | GIDDENS J P, LOMINO J V, AMIN M N, et al. Endo-F3 glycosynthase mutants enable chemoenzymatic synthesis of core-fucosylated triantennary complex type glycopeptides and glycoproteins[J]. Journal of Biological Chemistry, 2016, 291(17): 9356-9370. |
| 83 | FAN S Q, HUANG W, WANG L X. Remarkable transglycosylation activity of glycosynthase mutants of endo-D, an endo-β-N-acetylglucosaminidase from Streptococcus pneumoniae [J]. Journal of Biological Chemistry, 2012, 287(14): 11272-11281. |
| 84 | NATARAJAN A, JAROENTOMEECHAI T, CABRERA-SÁNCHEZ M, et al. Engineering orthogonal human O-linked glycoprotein biosynthesis in bacteria[J]. Nature Chemical Biology, 2020, 16(10): 1062-1070. |
| 85 | KIGHTLINGER W, DUNCKER K E, RAMESH A, et al. A cell-free biosynthesis platform for modular construction of protein glycosylation pathways[J]. Nature Communications, 2019, 10: 5404. |
| 86 | LIU F, VIJAYAKRISHNAN B, FARIDMOAYER A, et al. Rationally designed short polyisoprenol-linked PglB substrates for engineered polypeptide and protein N-glycosylation[J]. Journal of the American Chemical Society, 2014, 136(2): 566-569. |
| 87 | OZA J P, AERNI H R, PIRMAN N L, et al. Robust production of recombinant phosphoproteins using cell-free protein synthesis[J]. Nature Communications, 2015, 6: 8168. |
| 88 | JIN X, HONG S H. Cell-free protein synthesis for producing 'difficult-to-express' proteins[J]. Biochemical Engineering Journal, 2018, 138: 156-164. |
| 89 | DONDAPATI S K, STECH M, ZEMELLA A, et al. Cell-free protein synthesis: a promising option for future drug development[J]. BioDrugs, 2020, 34(3): 327-348. |
| 90 | BENCI J L, XU B H, QIU Y, et al. Tumor interferon signaling regulates a multigenic resistance program to immune checkpoint blockade[J]. Cell, 2016, 167(6): 1540-1554.e12. |
| 91 | XIE Q H, MATSUNAGA S, WEN Z S, et al. In vitro system for high-throughput screening of random peptide libraries for antimicrobial peptides that recognize bacterial membranes[J]. Journal of Peptide Science: an Official Publication of the European Peptide Society, 2006, 12(10): 643-652. |
| 92 | RAGHUNATH P. Roles of thermostable direct hemolysin (TDH) and TDH-related hemolysin (TRH) in vibrio parahaemolyticus[J]. Frontiers in Microbiology, 2015, 5: 805. |
| 93 | PARK N, KAHN J S, RICE E J, et al. High-yield cell-free protein production from P-gel[J]. Nature Protocols, 2009, 4(12): 1759-1770. |
| 94 | UM S H, LEE J B, PARK N, et al. Enzyme-catalysed assembly of DNA hydrogel[J]. Nature Materials, 2006, 5(10): 797-801. |
| 95 | PARK N, UM S H, FUNABASHI H, et al. A cell-free protein-producing gel[J]. Nature Materials, 2009, 8(5): 432-437. |
| 96 | RUIZ R C H, KIATWUTHINON P, KAHN J S, et al. Cell-free protein expression from DNA-based hydrogel (P-gel) droplets for scale-up production[J]. Industrial Biotechnology, 2012, 8(6): 372-377. |
| 1 | MARTIN J D. What's in a name change? [J]. Physics in Perspective, 2015, 17(1): 3-32. |
| 2 | CALLISTER JR W D, RETHWISCH D G. Fundamentals of materials science and engineering: an integrated approach[M]. John Wiley & Sons, 2020. |
| 3 | CROWTHER T W, GLICK H B, COVEY K R, et al. Mapping tree density at a global scale[J]. Nature, 2015, 525(7568): 201-205. |
| 4 | ARMSTRONG E. Voluntary greenhouse gas reporting[J]. Environmental Quality Management, 2011, 20(4): 29-42. |
| 5 | ERIKSEN M, LEBRETON L C M, CARSON H S, et al. Plastic pollution in the world's oceans: more than 5 trillion plastic pieces weighing over 250, 000 tons afloat at sea[J]. PLoS One, 2014, 9(12): e111913. |
| 6 | LENTON T M, ROCKSTRÖM J, GAFFNEY O, et al. Climate tipping points—too risky to bet against[J]. Nature, 2019, 575(7784): 592-595. |
| 7 | MENG F K, ELLIS T. The second decade of synthetic biology: 2010—2020[J]. Nature Communications, 2020, 11: 5174. |
| 8 | MAPLESTON P. New technologies for a greener industry[J]. Plastics Engineering, 2008, 64(1): 10-15. |
| 9 | COLWILL J, RAHIMIFARD S, CLEGG A. Eco-design tool to support the use of renewable polymers within packaging applications[C]// Glocalized Solutions for Sustainability in Manufacturing. Springer, 2011: 160-165. |
| 10 | AKSAKAL R, MERTENS C, SOETE M, et al. Applications of discrete synthetic macromolecules in life and materials science: recent and future trends[J]. Advanced Science, 2021, 8(6): 2004038. |
| 11 | YADAV P, YADAV H, SHAH V G, et al. Biomedical biopolymers, their origin and evolution in biomedical sciences: a systematic review[J]. Journal of Clinical and Diagnostic Research: JCDR, 2015, 9(9): ZE21-ZE25. |
| 12 | WEINER S, ADDADI L, WAGNER H D. Materials design in biology[J]. Materials Science and Engineering: C, 2000, 11(1): 1-8. |
| 97 | KAHN J S, RUIZ R C H, SUREKA S, et al. DNA microgels as a platform for cell-free protein expression and display[J]. Biomacromolecules, 2016, 17(6): 2019-2026. |
| 98 | CUI J H, WU D, SUN Q, et al. A PEGDA/DNA hybrid hydrogel for cell-free protein synthesis[J]. Frontiers in Chemistry, 2020, 8: 28. |
| 99 | YANG D Y, PENG S M, HARTMAN M R, et al. Enhanced transcription and translation in clay hydrogel and implications for early life evolution[J]. Scientific Reports, 2013, 3: 3165. |
| 100 | JIAO Y, LIU Y, LUO D, et al. Microfluidic-assisted fabrication of clay microgels for cell-free protein synthesis[J]. ACS Applied Materials & Interfaces, 2018, 10(35): 29308-29313. |
| 101 | LEE M S, HUNG C S, PHILLIPS D A, et al. Silk fibroin as an additive for cell-free protein synthesis[J]. Synthetic and Systems Biotechnology, 2020, 5(3): 145-154. |
| 102 | PARDEE K, GREEN A A, FERRANTE T, et al. Paper-based synthetic gene networks[J]. Cell, 2014, 159(4): 940-954. |
| 103 | PARDEE K. Perspective: solidifying the impact of cell-free synthetic biology through lyophilization[J]. Biochemical Engineering Journal, 2018, 138: 91-97. |
| 104 | BLUM S M, LEE M S, MGBOJI G E, et al. Impact of porous matrices and concentration by lyophilization on cell-free expression[J]. ACS Synthetic Biology, 2021, 10(5): 1116-1131. |
| 105 | CHO E, LU Y. Compartmentalizing cell-free systems: toward creating life-like artificial cells and beyond[J]. ACS Synthetic Biology, 2020, 9(11): 2881-2901. |
| 106 | PATRICK J F, ROBB M J, SOTTOS N R, et al. Polymers with autonomous life-cycle control[J]. Nature, 2016, 540(7633): 363-370. |
| 107 | SUCH G K, EVANS R A, DAVIS T P. Rapid photochromic switching in a rigid polymer matrix using living radical polymerization[J]. Macromolecules, 2006, 39(4): 1391-1396. |
| 108 | CASTANO L M, FLATAU A B. Smart fabric sensors and e-textile technologies: a review[J]. Smart Materials and Structures, 2014, 23(5): 053001. |
| 13 | SCHMALZ G. Determination of biocompatibility[M]//Biocompatibility of dental materials. Berlin, Heidelberg: Springer, 2009: 13-43. |
| 14 | GAD S C, GAD-MCDONALD S. Biomaterials, medical devices, and combination products: biocompatibility testing and safety assessment[M]. CRC Press, 2015. |
| 15 | ANDERSON J, STRELKOWA N, STAN G B, et al. Engineering and ethical perspectives in synthetic biology. Rigorous, robust and predictable designs, public engagement and a modern ethical framework are vital to the continued success of synthetic biology[J]. EMBO Reports, 2012, 13(7): 584-590. |
| 16 | CHURCH G M, ELOWITZ M B, SMOLKE C D, et al. Realizing the potential of synthetic biology[J]. Nature Reviews Molecular Cell Biology, 2014, 15(4): 289-294. |
| 17 | KELWICK R, MACDONALD J T, WEBB A J, et al. Developments in the tools and methodologies of synthetic biology[J]. Frontiers in Bioengineering and Biotechnology, 2014, 2: 60. |
| 18 | KHAMBHATI K, BHATTACHARJEE G, GOHIL N, et al. Exploring the potential of cell-free protein synthesis for extending the abilities of biological systems[J]. Frontiers in Bioengineering and Biotechnology, 2019, 7: 248. |
| 19 | MOORE S J, MACDONALD J T, WIENECKE S, et al. Rapid acquisition and model-based analysis of cell-free transcription-translation reactions from nonmodel bacteria[J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(19): E4340-E4349. |
| 20 | MARTIN R W, DES SOYE B J, KWON Y C, et al. Cell-free protein synthesis from genomically recoded bacteria enables multisite incorporation of noncanonical amino acids[J]. Nature Communications, 2018, 9(1): 1203. |
| 21 | BOWIE J U, SHERKHANOV S, KORMAN T P, et al. Synthetic biochemistry: the bio-inspired cell-free approach to commodity chemical production[J]. Trends in Biotechnology, 2020, 38(7): 766-778. |
| 22 | HODGMAN C E, JEWETT M C. Cell-free synthetic biology: thinking outside the cell[J]. Metabolic Engineering, 2012, 14(3): 261-269. |
| 23 | SMOLKE C D, SILVER P A. Informing biological design by integration of systems and synthetic biology[J]. Cell, 2011, 144(6): 855-859. |
| 24 | YUE K, ZHU Y Y, KAI L. Cell-free protein synthesis: chassis toward the minimal cell[J]. Cells, 2019, 8(4): 315. |
| 109 | HOARE T R, KOHANE D S. Hydrogels in drug delivery: progress and challenges[J]. Polymer, 2008, 49(8): 1993-2007. |
| 110 | AHN S K, KASI R M, KIM S C, et al. Stimuli-responsive polymer gels[J]. Soft Matter, 2008, 4(6): 1151. |
| 111 | QIAN Z G, ZHOU M L, SONG W W, et al. Dual thermosensitive hydrogels assembled from the conserved C‑terminal domain of spider dragline silk[J]. Biomacromolecules, 2015, 16(11): 3704-3711. |
| 112 | SONG W W, QIAN Z G, LIU H, et al. On-demand regulation of dual thermosensitive protein hydrogels[J]. ACS Macro Letters, 2021, 10(4): 395-400. |
| 113 | HU X, XIA X X, HUANG S C, et al. Development of adhesive and conductive resilin-based hydrogels for wearable sensors[J]. Biomacromolecules, 2019, 20(9): 3283-3293. |
| 114 | SARIKAYA M, TAMERLER C, JEN A K Y, et al. Molecular biomimetics: nanotechnology through biology[J]. Nature Materials, 2003, 2(9): 577-585. |
| 115 | WHITFIELD C J, BANKS A M, DURA G, et al. Cell-free genetic devices confer autonomic and adaptive properties to hydrogels[EB/OL]. bioRxiv, 2019. |
| 116 | KENNE L, GOHIL S, NILSSON E M, et al. Modification and cross-linking parameters in hyaluronic acid hydrogels — definitions and analytical methods[J]. Carbohydrate Polymers, 2013, 91(1): 410-418. |
| 117 | SMITH M T, BERKHEIMER S D, WERNER C J, et al. Lyophilized Escherichia coli-based cell-free systems for robust, high-density, long-term storage[J]. BioTechniques, 2014, 56(4): 186-193. |
| 118 | LEE K H, KIM D M. In vitro use of cellular synthetic machinery for biosensing applications[J]. Frontiers in Pharmacology, 2019, 10: 1166. |
| 119 | PARDEE K, GREEN A A, TAKAHASHI M K, et al. Rapid, low-cost detection of Zika virus using programmable biomolecular components[J]. Cell, 2016, 165(5): 1255-1266. |
| 120 | AHN D G, JEON I J, KIM J D, et al. RNA aptamer-based sensitive detection of SARS coronavirus nucleocapsid protein[J]. The Analyst, 2009, 134(9): 1896-1901. |
| 25 | BUCHNER E. Alkoholische gährung ohne hefezellen[J]. Berichte Der Deutschen Chemischen Gesellschaft, 1897, 30(1): 117-124. |
| 26 | NIRENBERG M W, MATTHAEI J H. The dependence of cell-free protein synthesis in E. coli upon naturally occurring or synthetic polyribonucleotides[J]. Proceedings of the National Academy of Sciences of the United States of America, 1961, 47(10): 1588-1602. |
| 27 | GREGORIO N E, LEVINE M Z, OZA J P. A user's guide to cell-free protein synthesis[J]. Methods and Protocols, 2019, 2(1): 24. |
| 28 | LIU W Q, ZHANG L K, CHEN M Z, et al. Cell-free protein synthesis: recent advances in bacterial extract sources and expanded applications[J]. Biochemical Engineering Journal, 2019, 141: 182-189. |
| 29 | GARENNE D, NOIREAUX V. Cell-free transcription-translation: engineering biology from the nanometer to the millimeter scale[J]. Current Opinion in Biotechnology, 2019, 58: 19-27. |
| 30 | SWARTZ J R. Expanding biological applications using cell-free metabolic engineering: an overview[J]. Metabolic Engineering, 2018, 50: 156-172. |
| 31 | WILDING K M, SCHINN S M, LONG E A, et al. The emerging impact of cell-free chemical biosynthesis[J]. Current Opinion in Biotechnology, 2018, 53: 115-121. |
| 32 | LU Y. Cell-free synthetic biology: engineering in an open world[J]. Synthetic and Systems Biotechnology, 2017, 2(1): 23-27. |
| 33 | DUDLEY Q M, KARIM A S, JEWETT M C. Cell-free metabolic engineering: biomanufacturing beyond the cell[J]. Biotechnology Journal, 2015, 10(1): 69-82. |
| 34 | GUTERL J K, GARBE D, CARSTEN J, et al. Cell-free metabolic engineering: production of chemicals by minimized reaction cascades[J]. ChemSusChem, 2012, 5(11): 2165-2172. |
| 35 | CARLSON E D, GAN R, HODGMAN C E, et al. Cell-free protein synthesis: applications come of age[J]. Biotechnology Advances, 2012, 30(5): 1185-1194. |
| 36 | SUN Z Z, HAYES C A, SHIN J, et al. Protocols for implementing an Escherichia coli based TX-TL cell-free expression system for synthetic biology[J]. Journal of Visualized Experiments: JoVE, 2013(79): e50762. |
| 37 | CALHOUN K A, SWARTZ J R. An economical method for cell-free protein synthesis using glucose and nucleoside monophosphates[J]. Biotechnology Progress, 2005, 21(4): 1146-1153. |
| 38 | LIU D V, ZAWADA J F, SWARTZ J R. Streamlining Escherichia coli S30 extract preparation for economical cell-free protein synthesis[J]. Biotechnology Progress, 2005, 21(2): 460-465. |
| 39 | JEWETT M C, CALHOUN K A, VOLOSHIN A, et al. An integrated cell-free metabolic platform for protein production and synthetic biology[J]. Molecular Systems Biology, 2008, 4(1): 220. |
| 40 | SMOLSKAYA S, LOGASHINA Y A, ANDREEV Y A. Escherichia coli extract-based cell-free expression system as an alternative for difficult-to-obtain protein biosynthesis[J]. International Journal of Molecular Sciences, 2020, 21(3): 928. |
| 41 | KWON Y C, JEWETT M C. High-throughput preparation methods of crude extract for robust cell-free protein synthesis[J]. Scientific Reports, 2015, 5: 8663. |
| 42 | DIDOVYK A, TONOOKA T, TSIMRING L, et al. Rapid and scalable preparation of bacterial lysates for cell-free gene expression[J]. ACS Synthetic Biology, 2017, 6(12): 2198-2208. |
| 43 | SILVERMAN A D, KARIM A S, JEWETT M C. Cell-free gene expression: an expanded repertoire of applications[J]. Nature Reviews Genetics, 2020, 21(3): 151-170. |
| 44 | GÜTTEL R. Chemical process technology. von J. A. Moulijn, M. Makkee, A. E. van Diepen[J]. Chemie Ingenieur Technik, 2014, 86(5): 585. |
| 45 | HÖÖK M, TANG X. Depletion of fossil fuels and anthropogenic climate change—a review[J]. Energy Policy, 2013, 52: 797-809. |
| 46 | CHAE T U, CHOI S Y, KIM J W, et al. Recent advances in systems metabolic engineering tools and strategies[J]. Current Opinion in Biotechnology, 2017, 47: 67-82. |
| 47 | CHUBUKOV V, MUKHOPADHYAY A, PETZOLD C J, et al. Synthetic and systems biology for microbial production of commodity chemicals[J]. NPJ Systems Biology and Applications, 2016, 2: 16009. |
| 48 | STEPHANOPOULOS G. Challenges in engineering microbes for biofuels production[J]. Science, 2007, 315(5813): 801-804. |
| 121 | LIU R, FU A S, DENG Z X, et al. Promising methods for detection of novel coronavirus SARS-CoV-2[J]. View, 2020, 1(1): e4. |
| 122 | RAMACHANDRAN N, RAPHAEL J V, HAINSWORTH E, et al. Next-generation high-density self-assembling functional protein arrays[J]. Nature Methods, 2008, 5(6): 535-538. |
| 123 | HE M Y, TAUSSIG M J. Single step generation of protein arrays from DNA by cell-free expression and in situ immobilisation (PISA method)[J]. Nucleic Acids Research, 2001, 29(15): e73. |
| 124 | BHIDE M, NATARAJAN S, HRESKO S, et al. Rapid in vitro protein synthesis pipeline: a promising tool for cost-effective protein array design[J]. Molecular BioSystems, 2014, 10(6): 1236-1245. |
| 125 | HE M Y, LIU H, TURNER M, et al. Detection of protein-protein interactions by ribosome display and protein in situ immobilisation[J]. New Biotechnology, 2009, 26(6): 277-281. |
| 126 | HE M Y, STOEVESANDT O, TAUSSIG M J. In situ synthesis of protein arrays[J]. Current Opinion in Biotechnology, 2008, 19(1): 4-9. |
| 127 | LEE K H, KWON Y C, YOO S J, et al. Ribosomal synthesis and in situ isolation of peptide molecules in a cell-free translation system[J]. Protein Expression and Purification, 2010, 71(1): 16-20. |
| 128 | LEE K H, LEE K Y, BYUN J Y, et al. On-bead expression of recombinant proteins in an agarose gel matrix coated on a glass slide[J]. Lab on a Chip, 2012, 12(9): 1605-1610. |
| 129 | BYUN J Y, LEE K H, LEE K Y, et al. In-gel expression and in situ immobilization of proteins for generation of three dimensional protein arrays in a hydrogel matrix[J]. Lab on a Chip, 2013, 13(5): 886-891. |
| 130 | KARIG D K, BESSLING S, THIELEN P, et al. Preservation of protein expression systems at elevated temperatures for portable therapeutic production[J]. Journal of the Royal Society, Interface, 2017, 14(129): 20161039. |
| 131 | BENÍTEZ-MATEOS A I, LLARENA I, SÁNCHEZ-IGLESIAS A, et al. Expanding one-pot cell-free protein synthesis and immobilization for on-demand manufacturing of biomaterials[J]. ACS Synthetic Biology, 2018, 7(3): 875-884. |
| 132 | MISIRLI G, NGUYEN T, MCLAUGHLIN J A, et al. A computational workflow for the automated generation of models of genetic designs[J]. ACS Synthetic Biology, 2019, 8(7): 1548-1559. |
| 49 | HASUNUMA T, OKAZAKI F, OKAI N, et al. A review of enzymes and microbes for lignocellulosic biorefinery and the possibility of their application to consolidated bioprocessing technology[J]. Bioresource Technology, 2013, 135: 513-522. |
| 50 | NIELSEN C, RAHMAN A, REHMAN A U, et al. Food waste conversion to microbial polyhydroxyalkanoates[J]. Microbial Biotechnology, 2017, 10(6): 1338-1352. |
| 51 | STRONG P J, LAYCOCK B, MAHAMUD S N S, et al. The opportunity for high-performance biomaterials from methane[J]. Microorganisms, 2016, 4(1): 11. |
| 52 | KELWICK R J R, WEBB A J, FREEMONT P S. Biological materials: the next frontier for cell-free synthetic biology[J]. Frontiers in Bioengineering and Biotechnology, 2020, 8: 399. |
| 53 | KELWICK R, RICCI L, CHEE S M, et al. Cell-free prototyping strategies for enhancing the sustainable production of polyhydroxyalkanoates bioplastics[J]. Synthetic Biology, 2018, 3(1): ysy016. |
| 54 | PETROLL K, KOPP D, CARE A, et al. Tools and strategies for constructing cell-free enzyme pathways[J]. Biotechnology Advances, 2019, 37(1): 91-108. |
| 55 | KOPP D, WILLOWS R D, SUNNA A. Cell-free enzymatic conversion of spent coffee grounds into the platform chemical lactic acid[J]. Frontiers in Bioengineering and Biotechnology, 2019, 7: 389. |
| 56 | CASTRO-AGUIRRE E, IÑIGUEZ-FRANCO F, SAMSUDIN H, et al. Poly(lactic acid)—mass production, processing, industrial applications, and end of life[J]. Advanced Drug Delivery Reviews, 2016, 107: 333-366. |
| 57 | OPGENORTH P H, KORMAN T P, BOWIE J U. A synthetic biochemistry module for production of bio-based chemicals from glucose[J]. Nature Chemical Biology, 2016, 12(6): 393-395. |
| 58 | ULLAH M W, UL-ISLAM M, KHAN S, et al. Innovative production of bio-cellulose using a cell-free system derived from a single cell line[J]. Carbohydrate Polymers, 2015, 132: 286-294. |
| 59 | GREEN E M. Fermentative production of butanol—the industrial perspective[J]. Current Opinion in Biotechnology, 2011, 22(3): 337-343. |
| 60 | ENDOH T, KANAI T, SATO Y T, et al. Cell-free protein synthesis at high temperatures using the lysate of a hyperthermophile[J]. Journal of Biotechnology, 2006, 126(2): 186-195. |
| 133 | LIAN J Z, JIN R, ZHAO H M. Construction of plasmids with tunable copy numbers in Saccharomyces cerevisiae and their applications in pathway optimization and multiplex genome integration[J]. Biotechnology and Bioengineering, 2016, 113(11): 2462-2473. |
| 134 | DU J, YUAN Y B, SI T, et al. Customized optimization of metabolic pathways by combinatorial transcriptional engineering[J]. Nucleic Acids Research, 2012, 40(18): e142. |
| 135 | JIANG L H, ZHAO J R, LIAN J Z, et al. Cell-free protein synthesis enabled rapid prototyping for metabolic engineering and synthetic biology[J]. Synthetic and Systems Biotechnology, 2018, 3(2): 90-96. |
| 136 | MOORE S J, MACDONALD J T, WIENECKE S, et al. Rapid acquisition and model-based analysis of cell-free transcription-translation reactions from nonmodel bacteria[J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(19): E4340-E4349. |
| 137 | KOPNICZKY M B, CANAVAN C, MCCLYMONT D W, et al. Cell-free protein synthesis as a prototyping platform for mammalian synthetic biology[J]. ACS Synthetic Biology, 2020, 9(1): 144-156. |
| 138 | SWANK Z, LAOHAKUNAKORN N, MAERKL S J. Cell-free gene-regulatory network engineering with synthetic transcription factors[J]. Proceedings of the National Academy of Sciences of the United States of America, 2019, 116(13): 5892-5901. |
| 139 | ZHANG Y, MINAGAWA Y, KIZOE H, et al. Accurate high-throughput screening based on digital protein synthesis in a massively parallel femtoliter droplet array[J]. Science Advances, 2019, 5(8): eaav8185. |
| 140 | SAWASAKI T, OGASAWARA T, MORISHITA R, et al. A cell-free protein synthesis system for high-throughput proteomics[J]. Proceedings of the National Academy of Sciences of the United States of America, 2002, 99(23): 14652-14657. |
| 141 | KANTER G, YANG J H, VOLOSHIN A, et al. Cell-free production of scFv fusion proteins: an efficient approach for personalized lymphoma vaccines[J]. Blood, 2007, 109(8): 3393-3399. |
| 142 | FAN J Z, VILLARREAL F, WEYERS B, et al. Multi-dimensional studies of synthetic genetic promoters enabled by microfluidic impact printing[J]. Lab on a Chip, 2017, 17(13): 2198-2207. |
| 143 | CONTRERAS-LLANO L E, TAN C. High-throughput screening of biomolecules using cell-free gene expression systems[J]. Synthetic Biology, 2018, 3(1): ysy012. |
| 144 | GUBERNATIS J E, LOOKMAN T. Machine learning in materials design and discovery: examples from the present and suggestions for the future[J]. Physical Review Materials, 2018, 2(12): 120301. |
| 61 | KRUTSAKORN B, HONDA K, YE X T, et al. In vitro production of n-butanol from glucose[J]. Metabolic Engineering, 2013, 20: 84-91. |
| 62 | YE X T, HONDA K, MORIMOTO Y, et al. Direct conversion of glucose to malate by synthetic metabolic engineering[J]. Journal of Biotechnology, 2013, 164(1): 34-40. |
| 63 | YOUNT N Y, YEAMAN M R. Immunocontinuum: perspectives in antimicrobial peptide mechanisms of action and resistance[J]. Protein and Peptide Letters, 2005, 12(1): 49-67. |
| 64 | XU M, LEWIS R V. Structure of a protein superfiber: spider dragline silk[J]. Proceedings of the National Academy of Sciences of the United States of America, 1990, 87(18): 7120-7124. |
| 65 | SAHIN KEHRIBAR E, ISILAK M E, BOZKURT E U, et al. Engineering of biofilms with a glycosylation circuit for biomaterial applications[J]. Biomaterials Science, 2021, 9(10): 3650-3661. |
| 66 | PEREZ J G, STARK J C, JEWETT M C. Cell-free synthetic biology: engineering beyond the cell[J]. Cold Spring Harbor Perspectives in Biology, 2016, 8(12): a023853. |
| 67 | CHONG S R. Overview of cell-free protein synthesis: historic landmarks, commercial systems, and expanding applications[J]. Current Protocols in Molecular Biology, 2014, 108: 16.30.1-16.3011. |
| 68 | SALEHI A S M, SMITH M T, BENNETT A M, et al. Cell-free protein synthesis of a cytotoxic cancer therapeutic: onconase production and a just-add-water cell-free system[J]. Biotechnology Journal, 2016, 11(2): 274-281. |
| 69 | ORTH J H C, SCHORCH B, BOUNDY S, et al. Cell-free synthesis and characterization of a novel cytotoxic pierisin-like protein from the cabbage butterfly Pieris rapae [J]. Toxicon, 2011, 57(2): 199-207. |
| 70 | MARTEMYANOV K A, SHIROKOV V A, KURNASOV O V, et al. Cell-free production of biologically active polypeptides: application to the synthesis of antibacterial peptide cecropin[J]. Protein Expression and Purification, 2001, 21(3): 456-461. |
| 71 | CHEN H Q, FAN L M, XU Z N, et al. Efficient production of soluble human beta-defensin-3-4 fusion proteins in Escherichia coli cell-free system[J]. Process Biochemistry, 2007, 42(3): 423-428. |
| 72 | MU J B, AWADALLA P, DUAN J H, et al. Genome-wide variation and identification of vaccine targets in the Plasmodium falciparum genome[J]. Nature Genetics, 2007, 39(1): 126-130. |
| 145 | GAO W, CHO E, LIU Y Y, et al. Advances and challenges in cell-free incorporation of unnatural amino acids into proteins[J]. Frontiers in Pharmacology, 2019, 10: 611. |
| 146 | HONG S H, NTAI I, HAIMOVICH A D, et al. Cell-free protein synthesis from a release factor 1 deficient Escherichia coli activates efficient and multiple site-specific nonstandard amino acid incorporation[J]. ACS Synthetic Biology, 2014, 3(6): 398-409. |
| 147 | MANKOWSKA S A, GATTI-LAFRANCONI P, CHODORGE M, et al. A shorter route to antibody binders via quantitative in vitro bead-display screening and consensus analysis[J]. Scientific Reports, 2016, 6: 36391. |
| 148 | ALBAYRAK C, SWARTZ J R. Direct polymerization of proteins[J]. ACS Synthetic Biology, 2014, 3(6): 353-362. |
| [1] | GAO Ge, BIAN Qi, WANG Baojun. Synthetic genetic circuit engineering: principles, advances and prospects [J]. Synthetic Biology Journal, 2025, 6(1): 45-64. |
| [2] | LI Jiyuan, WU Guosheng. Two hypothesises for the origins of organisms from the synthetic biology perspective [J]. Synthetic Biology Journal, 2025, 6(1): 190-202. |
| [3] | JIAO Hongtao, QI Meng, SHAO Bin, JIANG Jinsong. Legal issues for the storage of DNA data [J]. Synthetic Biology Journal, 2025, 6(1): 177-189. |
| [4] | TANG Xinghua, LU Qianneng, HU Yilin. Philosophical reflections on synthetic biology in the Anthropocene [J]. Synthetic Biology Journal, 2025, 6(1): 203-212. |
| [5] | XU Huaisheng, SHI Xiaolong, LIU Xiaoguang, XU Miaomiao. Key technologies for DNA storage: encoding, error correction, random access, and security [J]. Synthetic Biology Journal, 2025, 6(1): 157-176. |
| [6] | SHI Ting, SONG Zhan, SONG Shiyi, ZHANG Yi-Heng P. Job. In vitro BioTransformation (ivBT): a new frontier of industrial biomanufacturing [J]. Synthetic Biology Journal, 2024, 5(6): 1437-1460. |
| [7] | CHAI Meng, WANG Fengqing, WEI Dongzhi. Synthesis of organic acids from lignocellulose by biotransformation [J]. Synthetic Biology Journal, 2024, 5(6): 1242-1263. |
| [8] | SHAO Mingwei, SUN Simian, YANG Shimao, CHEN Guoqiang. Bioproduction based on extremophiles [J]. Synthetic Biology Journal, 2024, 5(6): 1419-1436. |
| [9] | CHEN Yu, ZHANG Kang, QIU Yijing, CHENG Caiyun, YIN Jingjing, SONG Tianshun, XIE Jingjing. Progress of microbial electrosynthesis for conversion of CO2 [J]. Synthetic Biology Journal, 2024, 5(5): 1142-1168. |
| [10] | ZHENG Haotian, LI Chaofeng, LIU Liangxu, WANG Jiawei, LI Hengrun, NI Jun. Design, optimization and application of synthetic carbon-negative phototrophic community [J]. Synthetic Biology Journal, 2024, 5(5): 1189-1210. |
| [11] | CHEN Ziling, XIANG Yangfei. Integrated development of organoid technology and synthetic biology [J]. Synthetic Biology Journal, 2024, 5(4): 795-812. |
| [12] | LI Shikai, ZENG Dong′ao, DU Fangzhou, ZHANG Jingzhong, YU Shuang. The construction approaches and biomaterials for vascularized organoids [J]. Synthetic Biology Journal, 2024, 5(4): 851-866. |
| [13] | CAI Bingyu, TAN Xiangtian, LI Wei. Advances in synthetic biology for engineering stem cell [J]. Synthetic Biology Journal, 2024, 5(4): 782-794. |
| [14] | XIE Huang, ZHENG Yilei, SU Yiting, RUAN Jingyi, LI Yongquan. An overview on reconstructing the biosynthetic system of actinomycetes for polyketides production [J]. Synthetic Biology Journal, 2024, 5(3): 612-630. |
| [15] | ZHA Wenlong, BU Lan, ZI Jiachen. Advances in synthetic biology for producing potent pharmaceutical ingredients of traditional Chinese medicine [J]. Synthetic Biology Journal, 2024, 5(3): 631-657. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||