Synthetic Biology Journal ›› 2020, Vol. 1 ›› Issue (6): 722-731.DOI: 10.12211/2096-8280.2020-020
• Invited Review • Previous Articles Next Articles
An overview on regulatory mechanism of daptomycin biosynthesis
FANG Jiaole1,2, LYU Zhongyuan1,2, SUN Chenfan1,2, LIU Yifan1,2, XU Weifeng1,2, MAO Xuming1,2, LI Yongquan1,2
- 1.Institute of Pharmaceutical Biotechnology,Zhejiang University,Hangzhou 310058,Zhejiang,China
2.Zhejiang Provincial Key Laboratory for Microbial Biochemistry and Metabolic Engineering,Hangzhou 310058,Zhejiang,China
-
Received:2020-03-10Revised:2020-11-05Online:2021-01-15Published:2020-12-31 -
Contact:MAO Xuming, LI Yongquan
达托霉素生物合成过程的调控机制研究进展
方教乐1,2, 吕中原1,2, 孙晨番1,2, 刘一帆1,2, 徐炜锋1,2, 毛旭明1,2, 李永泉1,2
- 1.浙江大学药物生物技术研究所,浙江 杭州 310058
2.浙江省微生物生化与代谢工程重点实验室,浙江 杭州 310058
-
通讯作者:毛旭明,李永泉 -
作者简介:方教乐(1991—),男,博士研究生,研究方向为微生物次级代谢产物调控,链霉菌隐性基因簇激活,表观遗传学研究。E-mail:fjl20@live.cn
毛旭明(1978—),男,博士,教授,研究方向为基于合成生物学的微生物药物开发、微生物药物生物合成的调控机制研究、基于多组学的新活性和新结构微生物天然产物挖掘、微生物天然产物生物合成的酶学机制和化学机制研究。E-mail:xmmao@zju.edu.cn
李永泉(1962—),男,博士,求是特聘教授,研究方向为微生物合成生物学、微生物次级代谢调控和微生物制药。E-mail:lyq@zju.edu.cn -
基金资助:国家新药创制重大专项(2018ZX09711001-006-013);国家自然科学基金(3173002)
CLC Number:
Cite this article
FANG Jiaole, LYU Zhongyuan, SUN Chenfan, LIU Yifan, XU Weifeng, MAO Xuming, LI Yongquan. An overview on regulatory mechanism of daptomycin biosynthesis[J]. Synthetic Biology Journal, 2020, 1(6): 722-731.
方教乐, 吕中原, 孙晨番, 刘一帆, 徐炜锋, 毛旭明, 李永泉. 达托霉素生物合成过程的调控机制研究进展[J]. 合成生物学, 2020, 1(6): 722-731.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2020-020
| 1 | HUBER F M, PIEPER R L, TIETZ A J. The synthesis of A21978C analogs by Streptomyces roseosporus cultivated under carbon limitation and fed fatty acids [J]. Journal of Biotechnology, 1990, 7: 283-292. |
| 2 | DOEKEL S, COËFFET-LE GAL M F, GU Jianqiao, et al. Non-ribosomal peptide synthetase module fusions to produce derivatives of daptomycin in Streptomyces roseosporus [J]. Microbiology, 2008, 154(9): 2872-2880. |
| 3 | SAUERMANN R, ROTHENBURGER M, GRANINGER W, et al. Daptomycin: a review 4 years after first approval [J]. Pharmacology, 2008, 81: 79-91. |
| 4 | SMITH J R, CLAEYS K C, ZASOWSKI E J, et al. Daptomycin resistance [M]// MAYERS D L, SOBEL J D, OUELLETTE M, et al. Antimicrobial drug resistance: mechanisms of drug resistance. Springer International Publishing; 2017: 307-317. |
| 5 | PERSECHINI A, MONCRIEF N D, KRETSINGER R H. The EF-hand family of calcium-modulated proteins [J]. Trends in Neurosciences, 1989, 12(11): 462-467. |
| 6 | ROBBEL L, MARAHIEL M A. Daptomycin, a bacterial lipopeptide synthesized by a nonribosomal machinery [J]. Journal of Biological Chemistry, 2010, 285(36): 27501-27508. |
| 7 | ROMERO-RODRÍGUEZ A, ROBLEDO-CASADOS I, SÁNCHEZ S. An overview on transcriptional regulators in Streptomyces [J]. Biochimica et Biophysica Acta (BBA)-Gene Regulatory Mechanisms, 2015, 1849(8): 1017-1039. |
| 8 | LIU Gang, CHATER K F, CHANDRA G, et al. Molecular regulation of antibiotic biosynthesis in Streptomyces [J]. Microbiology and Molecular Biology Reviews, 2013, 77(12): 112-143. |
| 9 | LUO Shuai, CHEN Xin'ai, MAO Xuming, et a. Transposon-based identification of a negative regulator for the antibiotic hyper-production in Streptomyces [J]. Applied Microbiology and Biotechnology, 2018, 102(15): 6581-6592. |
| 10 | MAO Xuming, LUO Shuai, ZHOU Richeng, et al. Transcriptional regulation of the daptomycin gene cluster in Streptomyces roseosporus by an autoregulator, AtrA [J]. Journal of Biological Chemistry, 2015, 290(12): 7992-8001. |
| 11 | YUAN Penghui, ZHOU Richeng, CHEN Xuepeng, et a. DepR1, a TetR family transcriptional regulator, positively regulates daptomycin production in an industrial producer, Streptomyces roseosporus SW0702 [J]. Applied and Environmental Microbiology, 2016, 82(6): 03002. |
| 12 | MAO Xuming, LUO Shuai, LI Yongquan. Negative regulation of daptomycin production by DepR2, an ArsR-family transcriptional factor [J]. Journal of Industrial Microbiology & Biotechnology, 2017, 44(6): 1653-1658. |
| 13 | RAMOS J L, MARTÍNEZ-BUENO M, MOLINA-HENARES A J, et al. The TetR family of transcriptional repressors [J]. Microbiology and Molecular Biology Reviews, 2005, 69(2): 326-356. |
| 14 | XU Delin, SEGHEZZI N, ESNAULT C, et al. Repression of antibiotic production and sporulation in Streptomyces coelicolor by overexpression of a TetR family transcriptional regulator [J]. Applied and Environmental Microbiology, 2010, 76(23): 7741-7753. |
| 15 | LIU Wenshuai, ZHANG Qinling, GUO Jia, et al. Increasing avermectin production in Streptomyces avermitilis by manipulating the expression of a novel TetR-family regulator and its target gene product [J]. Applied and Environmental Microbiology, 2015, 81(15): 5157-5173. |
| 16 | WEI Junhong, TIAN Yuqing, NIU Guoqing, et al. GouR, a TetR family transcriptional regulator, coordinates the biosynthesis and export of gougerotin in Streptomyces graminearus [J]. Applied and Environmental Microbiology, 2014, 80(2): 714-722. |
| 17 | BUSENLEHNER L S, PENNELLA M A, GIEDROC D P. The SmtB/ArsR family of metalloregulatory transcriptional repressors: structural insights into prokaryotic metal resistance [J]. FEMS Microbiology Reviews, 2003, 27(23): 131-143. |
| 18 | KIM Hae Mi, Bo-Eun AHN, Ju-Hyung LEE, et al. Regulation of a nickel-cobalt efflux system and nickel homeostasis in a soil actinobacterium Streptomyces coelicolor [J]. Metallomics, 2015, 7(4): 702-709. |
| 19 | ZMIJEWSKI M J, BRIGGS B, OCCOLOWITZ J. Role of branched chain fatty acid precursors in regulating factor profile in the biosynthesis of A21978C complex [J]. The Journal of Antibiotics, 1986, 39(10): 1483-1485. |
| 20 | TARDU M, BULUT S, KAVAKLI I H. MerR and ChrR mediate blue light induced photo-oxidative stress response at the transcriptional level in Vibrio cholerae [J]. Scientific Reports, 2017, 7: 40817. |
| 21 | BROWN N L, STOYANOV J V, KIDD S P, et al. The MerR family of transcriptional regulators [J]. FEMS Microbiology Reviews, 2003, 27(2): 145-163. |
| 22 | SCHUMACHER M A, HENGST C D DEN, BUSH M J, et al. The MerR-like protein BldC binds DNA direct repeats as cooperative multimers to regulate Streptomyces development [J]. Nature Communications, 2018, 9(1): 1139. |
| 23 | MIAO V, COËFFET-LE GAL M F, BRIAN P, et al. Daptomycin biosynthesis in Streptomyces roseosporus: cloning and analysis of the gene cluster and revision of peptide stereochemistry [J]. Microbiology, 2005, 151(5): 1507-1523. |
| 24 | 靳旭, 魏维, 饶敏, 等. 链霉菌HCCB10043中调控基因dptR1、dptR2及dptR3对A21978C合成的影响[J]. 中国抗生素杂志, 2014, 39(7): 490-493. |
| JIN Xu, WEI Wei, RAO Min, et al. Influence of regulatory genes dptR1, dptR2 and dptR3 on A21978C production in Streptomyces sp. HCCB10043 [J]. Chinese Journal of Antibiotics, 2014, 39(7): 490-493. | |
| 25 | ULANOVA D, KITANI S, FUKUSAKI E, et al. SdrA, a new DeoR family regulator involved in Streptomyces avermitilis morphological development and antibiotic production [J]. Applied and Environmental Microbiology, 2013, 79(24): 7916-7921. |
| 26 | GE Beibei, LIU Yan, LIU Binghua, et al. Characterization of novel DeoR-family member from the Streptomyces ahygroscopicus strain CK-15 that acts as a repressor of morphological development [J]. Applied Microbiology and Biotechnology, 2016, 100(20): 8819-8828. |
| 27 | JEON Jong-Min, CHOI Tae-Rim, Bo-Rahm LEE, et al. Decreased growth and antibiotic production in Streptomyces coelicolor A3(2) by deletion of a highly conserved DeoR family regulator, SCO1463 [J]. Biotechnology and Bioprocess Engineering 2019, 24(4): 613-621. |
| 28 | WANG Feng, REN Nini, LUO Shuai, et al. DptR2, a DeoR-type auto-regulator, is required for daptomycin production in Streptomyces roseosporus [J]. Gene, 2014, 544(2): 208-215. |
| 29 | MARTIN R G, ROSNER J L. Binding of purified multiple antibiotic-resistance repressor protein (MarR) to mar operator sequences [J]. Proceedings of the National Academy of Sciences of the United States of America, 1995, 92(12): 5456-5460. |
| 30 | INOKA C P, GROVE A. Molecular mechanisms of ligand-mediated attenuation of DNA binding by MarR family transcriptional regulators [J]. Journal of Molecular Cell Biology, 2010, 2(5): 243-254 |
| 31 | GROVE A. Regulation of metabolic pathways by MarR family transcription factors [J]. Computational and Structural Biotechnology Journal, 2017, 15: 366-371. |
| 32 | So-Young OH, SHIN Jung-Ho, Jung-Hye ROE. Dual role of OhrR as a repressor and an activator in response to organic hydroperoxides in Streptomyces coelicolor [J]. Journal of Bacteriology, 2007, 189(17): 6284-6292. |
| 33 | HUANG Hao, GROVE A. The transcriptional regulator TamR from Streptomyces coelicolor controls a key step in central metabolism during oxidative stress [J]. Mol. Microbiol., 2013, 87(6): 1151-1166. |
| 34 | ZHANG Qinling, CHEN Qiong, ZHUANG Shuai, et al. A MarR family transcriptional regulator, DptR3, activates daptomycin biosynthesis and morphological differentiation in Streptomyces roseosporus [J]. Applied and Environmental Microbiology, 2015, 81(11): 3753-3765. |
| 35 | BUSH M J, BIBB M J, CHANDRA G, et al. Genes required for aerial growth, cell division, and chromosome segregation are targets of WhiA before sporulation in Streptomyces venezuelae [J] mBio, 2013, 4(5): e00684-00613. |
| 36 | HUANG Xingwei, MA Tingmei, TIAN Jun, et al. wblA, a pleiotropic regulatory gene modulating morphogenesis and daptomycin production in Streptomyces roseosporus [J]. Journal of Applied Microbiology, 2017, 123(3): 669-677. |
| 37 | NISHIDA H, OHNISHI Y, BEPPU T, et al. Evolution of γ-butyrolactone synthases and receptors in Streptomyces [J]. Environmental Microbiology, 2007, 9(8): 1986-1994. |
| 38 | OHNISHI Y, HORINOUCHI S. The A-factor regulatory cascade that leads to morphological development and secondary metabolism in Streptomyces [J]. Biofilms, 2004, 1(4): 319-328. |
| 39 | KIM Hyun Soo, NIHIRA T, TADA H, et al. Identification of binding protein of virginiae butanolide C, an autoregulator in virginiamycin production, from Streptomyces virginiae [J]. The Journal of Antibiotics, 1989, 42(5): 769-778. |
| 40 | HASHIMOTO K, NIHIRA T, SAKUDA S, et al. IM-2, a butyrolactone autoregulator, induces production of several nucleoside antibiotics in Streptomyces sp. FRI-5 [J]. Journal of Fermentation and Bioengineering, 1992, 73(6): 449-455. |
| 41 | OHNISHI Y, YAMAZAKI H, KATO J, et al. AdpA, a central transcriptional regulator in the A-factor regulatory cascade that leads to morphological development and secondary metabolism in Streptomyces griseus [J]. Bioscience, Biotechnology, and Biochemistry, 2005, 69(3): 431-439. |
| 42 | RODRÍGUEZ H, RICO S, DÍAZ M, et al. Two-component systems in Streptomyces: key regulators of antibiotic complex pathways. [J] Microbial Cell Factories, 2013, 12(1): 127. |
| 43 | SOLALANDA A, MOURA R S, MARTIN J F. The two-component PhoR-PhoP system controls both primary metabolism and secondary metabolite biosynthesis in Streptomyces lividans [J]. Proceedings of the National Academy of Sciences of the United States of America 2003, 100(10): 6133-6138. |
| 44 | BRIAN P, RIGGLE P, SANTOS R A, et al. Global negative regulation of Streptomyces coelicolor antibiotic synthesis mediated by an absA-encoded putative signal transduction system [J]. Journal of Bacteriology, 1996, 178(11): 3221-3231. |
| 45 | WANG Rui, MAST Y, WANG Jin, et al. Identification of two-component system AfsQ1/Q2 regulon and its cross-regulation with GlnR in Streptomyces coelicolor [J]. Molecular Microbiology, 2013, 87(1): 30-48. |
| 46 | UMEYAMA T, Ping-Chin LEE, UEDA K, et al. An AfsK/AfsR system involved in the response of aerial mycelium formation to glucose in Streptomyces griseus [J]. Microbiology, 1999, 145(9): 2281-2292. |
| 47 | MARTIN J F. Phosphate control of the biosynthesis of antibiotics and other secondary metabolites is mediated by the PhoR-PhoP system: an unfinished story [J]. Journal of Bacteriology, 2004, 186(16): 5197-5201. |
| 48 | ZHENG Yang, SUN Chenfan, FU Yu,et al. Dual regulation between the two-component system PhoRP and AdpA regulates antibiotic production in Streptomyces [J]. Journal of Industrial Microbiology & Biotechnology, 2019, 46(5): 725-737. |
| 49 | DEBONO M, BARNHART M, CARRELL C B, et al. A21978C, a complex of new acidic peptide antibiotics: isolation, chemistry, and mass spectral structure elucidation [J]. The Journal of Antibiotics, 1987, 40(6): 761-777. |
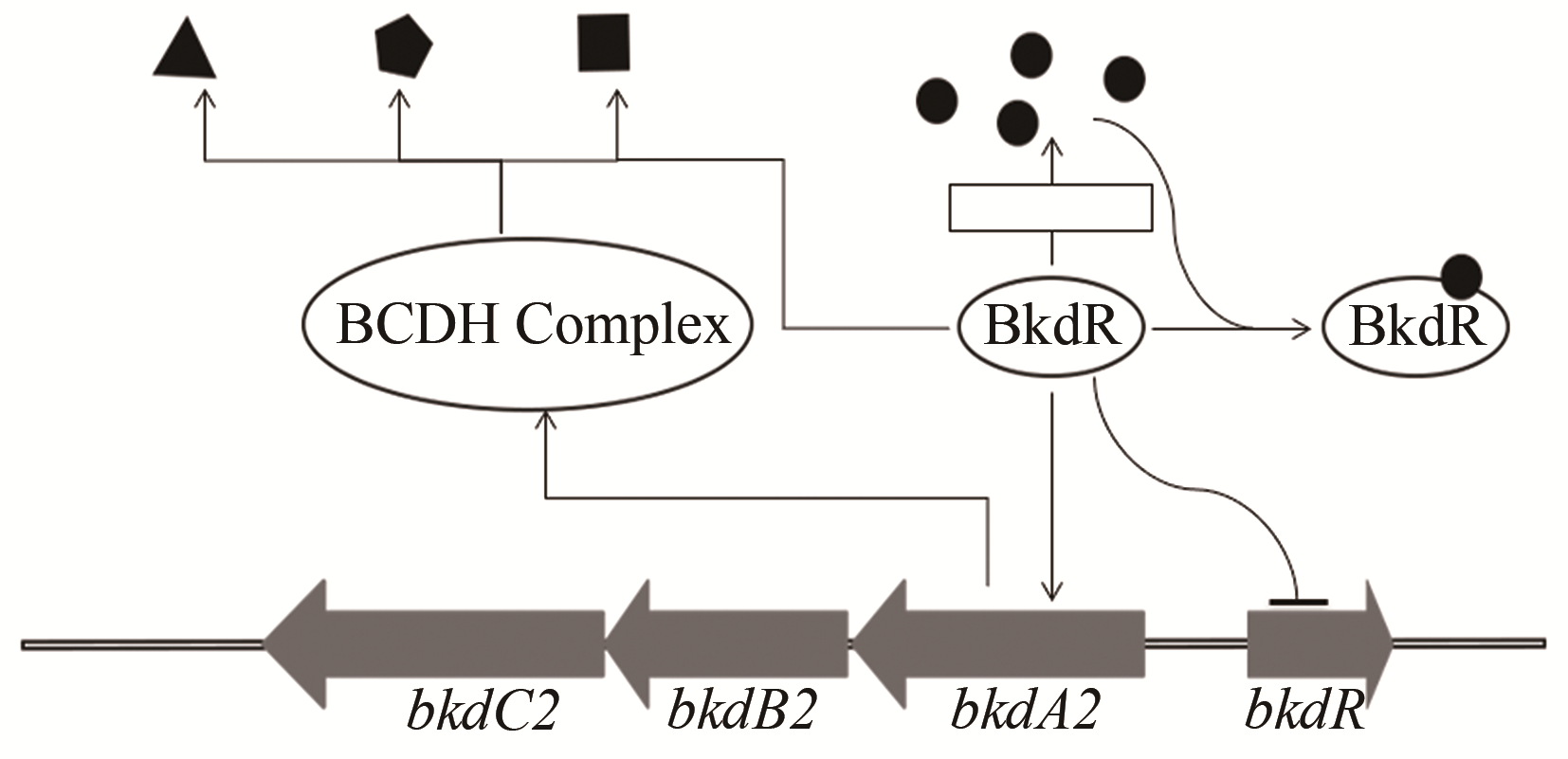
| 50 | LUO Shuai, CHEN Xin'ai, MAO Xuming, et al. Regulatory and biosynthetic effects of the bkd gene clusters on the production of daptomycin and its analogs A21978C1-3 [J]. Journal of Industrial Microbiology & Biotechnology, 2018, 45(4): 271-279. |
| 51 | SPRUSANSKY O, STIRRETT K, SKINNER D D, et al. The bkdR gene of Streptomyces coelicolor is required for morphogenesis and antibiotic production and encodes a transcriptional regulator of a branched-chain amino acid dehydrogenase complex [J]. Journal of Bacteriology, 2005, 187(2): 664-671. |
| [1] | ZHONG Quanzhou, SHAN Yiyi, PEI Qingyun, JIN Yanyun, WANG Yihan, MENG Luyuan, WANG Xinyun, ZHANG Yuxin, LIU Kunyuan, WANG Huizhong, FENG Shangguo. Research progress in the production of α-arbutin through biosynthesis [J]. Synthetic Biology Journal, 2025, 6(1): 118-135. |
| [2] | ZHU Fanghuan, CEN Xuecong, CHEN Zhen. Research progress of diols production by microbes [J]. Synthetic Biology Journal, 2024, 5(6): 1367-1385. |
| [3] | LIU Yining, PU Wei, YANG Jinxing, WANG Yu. Recent advances in the biosynthesis of ω-amino acids and lactams [J]. Synthetic Biology Journal, 2024, 5(6): 1350-1366. |
| [4] | ZHENG Haotian, LI Chaofeng, LIU Liangxu, WANG Jiawei, LI Hengrun, NI Jun. Design, optimization and application of synthetic carbon-negative phototrophic community [J]. Synthetic Biology Journal, 2024, 5(5): 1189-1210. |
| [5] | CHENG Xiaolei, LIU Tiangang, TAO Hui. Recent research progress in non-canonical biosynthesis of terpenoids [J]. Synthetic Biology Journal, 2024, 5(5): 1050-1071. |
| [6] | LIU Zijian, MU Baiyang, DUAN Zhiqiang, WANG Xuan, LU Xiaojie. Advances in the development of DNA-compatible chemistries [J]. Synthetic Biology Journal, 2024, 5(5): 1102-1124. |
| [7] | ZHANG Shouqi, WANG Tao, KONG Yao, ZOU Jiasheng, LIU Yuanning, XU Zhengren. Chemoenzymatic synthesis of natural products: evolution of synthetic methodology and strategy [J]. Synthetic Biology Journal, 2024, 5(5): 913-940. |
| [8] | XIE Xiangqian, GUO Wen, WANG Huan, LI Jin. Biosynthesis and chemical synthesis of ribosomally synthesized and post-translationally modified peptides containing aminovinyl cysteine [J]. Synthetic Biology Journal, 2024, 5(5): 981-996. |
| [9] | TANG Zhijun, HU Youcai, LIU Wen. Enzymatic (4+2)- and (2+2)-cycloaddition reactions: fundamentals and applications of regio- and stereoselectivity [J]. Synthetic Biology Journal, 2024, 5(3): 401-407. |
| [10] | ZHANG Jun, JIN Shixue, YUN Qian, QU Xudong. Biosynthesis of the unnatural extender units with polyketides and their structural modifications for applications in medicines [J]. Synthetic Biology Journal, 2024, 5(3): 561-570. |
| [11] | CHEN Xiwei, ZHANG Huaran, ZOU Yi. Biosynthesis and metabolic engineering of fungal non-ribosomal peptides [J]. Synthetic Biology Journal, 2024, 5(3): 571-592. |
| [12] | FENG Jin, PAN Haixue, TANG Gongli. Research advances in biosynthesis of natural product drugs within the past decade [J]. Synthetic Biology Journal, 2024, 5(3): 408-446. |
| [13] | XI Mengyu, HU Yiling, GU Yucheng, GE Huiming. Genome mining-directed discovery for natural medicinal products [J]. Synthetic Biology Journal, 2024, 5(3): 447-473. |
| [14] | SHI Xinjie, DU Yiling. Research advances in the biosynthesis of nonribosomal peptides within the bisintercalator family as anticancer drugs [J]. Synthetic Biology Journal, 2024, 5(3): 593-611. |
| [15] | SONG Yongxiang, ZHANG Xiufeng, LI Yanqin, XIAO Hua, YAN Yan. Resistance-gene directed discovery of bioactive natural products [J]. Synthetic Biology Journal, 2024, 5(3): 474-491. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||