Synthetic Biology Journal ›› 2022, Vol. 3 ›› Issue (3): 516-529.DOI: 10.12211/2096-8280.2021-054
• Invited Review • Previous Articles Next Articles
Application of imine reductase in the synthesis of chiral amines
YANG Lu, QU Xudong
- State Key Laboratory of Microbial Metabolism,School of Life Sciences and Biotechnology,Shanghai Jiao Tong University,Shanghai 200240,China
-
Received:2021-05-01Revised:2021-07-27Online:2022-07-13Published:2022-06-30 -
Contact:QU Xudong
亚胺还原酶在手性胺合成中的应用
杨璐, 瞿旭东
- 上海交通大学生命科学技术学院,微生物代谢国家重点实验室,教育部代谢与发育科学国际合作联合实验室,上海 200240
-
通讯作者:瞿旭东 -
作者简介:杨璐 (1993—),女,博士研究生。研究方向为杂环类生物碱的生物合成研究及酶工程改造。E-mail:yl2020@sjtu.edu.cn瞿旭东 (1980—),男,教授,博士生导师。研究方向为生物合成与生物催化。E-mail:quxd19@sjtu.edu.cn -
基金资助:国家自然科学基金(31970054);国家重点研发计划(2018YFC1706200)
CLC Number:
Cite this article
YANG Lu, QU Xudong. Application of imine reductase in the synthesis of chiral amines[J]. Synthetic Biology Journal, 2022, 3(3): 516-529.
杨璐, 瞿旭东. 亚胺还原酶在手性胺合成中的应用[J]. 合成生物学, 2022, 3(3): 516-529.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2021-054
| Entry | Enzyme | Source | Substrate | Products | ee/% | Reference |
|---|---|---|---|---|---|---|
| 1 | IR2 | Actinomaduramadurae | 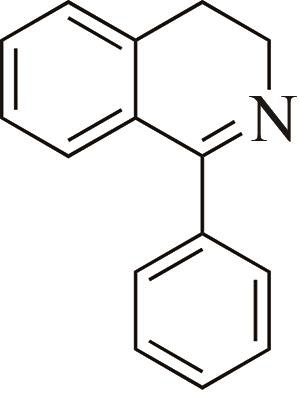 | 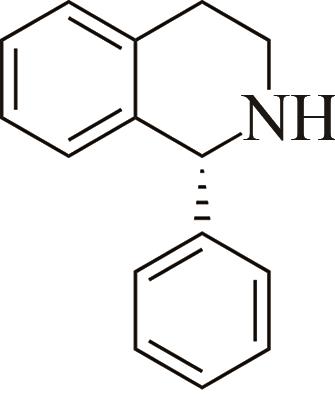 | 99 R | [ |
| 2 | IR45 | Streptomyces aurantiacus | 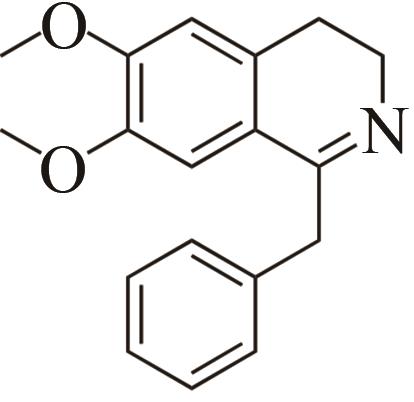 | 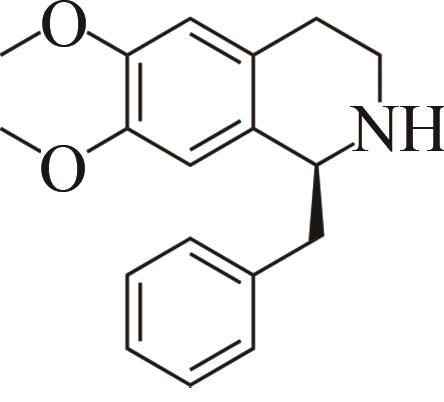 | 99 S | [ |
| 3 | IR46 | Saccharothrixespanaensis | 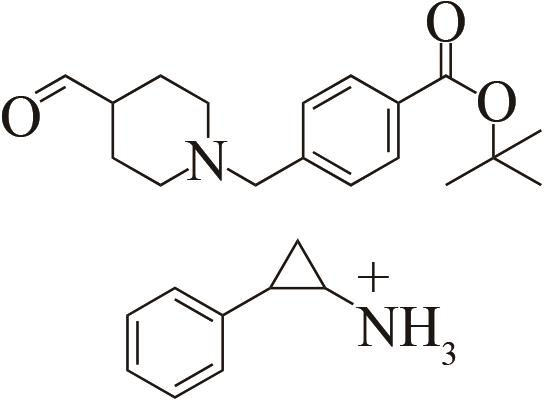 | 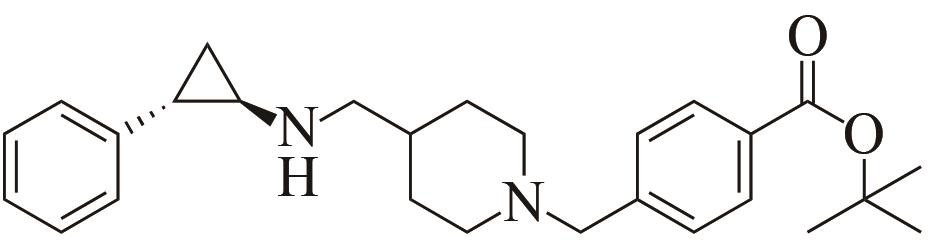 | 99 R | [ |
| 4 | IR1 | Leishmania major | 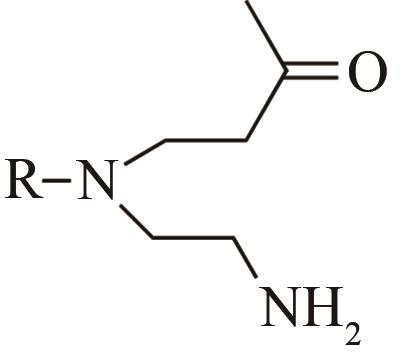 | 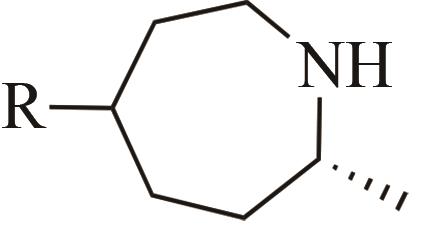 | 99 R | [ |
| 5 | AoIRED | Amycolatopsisorientalis | 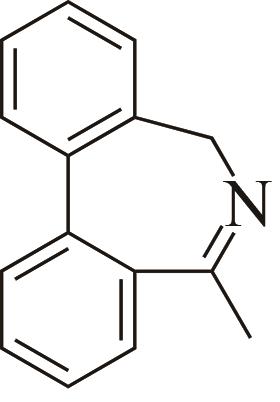 | 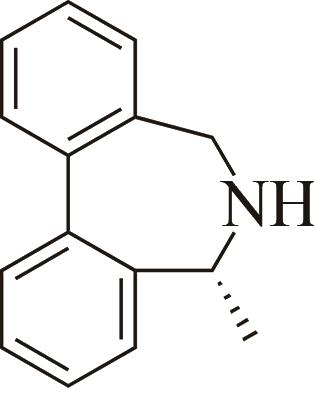 | 40 R | [ |
Tab. 1 Examples of imine reductase modifications
| Entry | Enzyme | Source | Substrate | Products | ee/% | Reference |
|---|---|---|---|---|---|---|
| 1 | IR2 | Actinomaduramadurae |  |  | 99 R | [ |
| 2 | IR45 | Streptomyces aurantiacus |  |  | 99 S | [ |
| 3 | IR46 | Saccharothrixespanaensis |  |  | 99 R | [ |
| 4 | IR1 | Leishmania major |  |  | 99 R | [ |
| 5 | AoIRED | Amycolatopsisorientalis |  |  | 40 R | [ |
| 1 | GHISLIERI D, TURNER N J. Biocatalytic approaches to the synthesis of enantiomerically pure chiral amines[J]. Topics in Catalysis, 2014, 57(5):284-300. |
| 2 | SCHRITTWIESER J H, VELIKOGNE S, KROUTIL W. Biocatalytic imine reduction and reductive amination of ketones[J]. Advanced Synthesis & Catalysis, 2015, 357(8):1655-1685. |
| 3 | GUO F, BERGLUND P. Transaminase biocatalysis: optimization and application[J]. Green Chemistry, 2017, 19(2):333-360. |
| 4 | MATHEW S, YUN H. ω-Transaminases for the production of optically pure amines and unnatural amino acids[J]. ACS Catalysis, 2012, 2(6):993-1001. |
| 5 | KOHLS H, STEFFEN-MUNSBERG F, HÖHNE M. Recent achievements in developing the biocatalytic toolbox for chiral amine synthesis[J]. Current Opinion in Chemical Biology, 2014, 19: 180-192. |
| 6 | JIANG J J, CHEN X, ZHANG D L, et al. Characterization of (R)-selective amine transaminases identified by in silico motif sequence blast[J]. Applied Microbiology and Biotechnology, 2015, 99(6):2613-2621. |
| 7 | JIANG J J, CHEN X, FENG J H, et al. Substrate profile of an ω-transaminase from Burkholderia vietnamiensis and its potential for the production of optically pure amines and unnatural amino acids[J]. Journal of Molecular Catalysis B: Enzymatic, 2014, 100: 32-39. |
| 8 | YANG L, ZHU J M, SUN C H, et al. Biosynthesis of plant tetrahydroisoquinoline alkaloids through an imine reductase route[J]. Chemical Science, 2019, 11(2): 364-371. |
| 9 | KIM Y, PARK J, KIM M J. Dynamic kinetic resolution of amines and amino acids by enzyme-metal cocatalysis[J]. ChemCatChem, 2011, 3(2):271-277. |
| 10 | NUGENT T C, EL-SHAZLY M. Chiral amine synthesis-recent developments and trends for enamide reduction, reductive amination, and imine reduction[J]. Advanced Synthesis & Catalysis, 2010, 352(5):753-819. |
| 11 | CHENG F, ZHU L L, SCHWANEBERG U. Directed evolution 2.0: improving and deciphering enzyme properties[J]. Chemical Communications, 2015, 51(48):9760-9772. |
| 12 | RAMSDEN J I, HEATH R S, DERRINGTON S R, et al. Biocatalytic N-alkylation of amines using either primary alcohols or carboxylic acids via reductive aminase cascades[J]. Journal of the American Chemical Society, 2019, 141(3):1201-1206. |
| 13 | CHENG F, CHEN X L, XIANG C, et al. Fluorescence-based high-throughput screening system for R-ω-transaminase engineering and its substrate scope extension[J]. Applied Microbiology and Biotechnology, 2020, 104(7): 2999-3009. |
| 14 | BALKENHOHL F, DITRICH K, HAUER B, et al. Optically active amines via lipase-catalyzed methoxyacetylation[J]. Journal Fur Praktische Chemie-Chemiker-Zeitung, 1997, 339(4):381-384. |
| 15 | AHN Y, KO S B, KIM M J, et al. Racemization catalysts for the dynamic kinetic resolution of alcohols and amines[J]. Coordination Chemistry Reviews, 2008, 252(5/6/7): 647-658. |
| 16 | SAVILE C K, JANEY J M, MUNDORFF E C, et al. Biocatalytic asymmetric synthesis of chiral amines from ketones applied to sitagliptin manufacture[J]. Science, 2010, 329(5989):305-309. |
| 17 | KELLY S A, POHLE S, WHARRY S, et al. Application of ω-transaminases in the pharmaceutical industry[J]. Chemical Reviews, 2018, 118(1):349-367. |
| 18 | MUTTI F G, FUCHS C S, PRESSNITZ D, et al. Stereoselectivity of four (R)-selective transaminases for the asymmetric amination of ketones[J]. Advanced Synthesis & Catalysis, 2011, 353(17):3227-3233. |
| 19 | GOMM A, O'REILLY E. Transaminases for chiral amine synthesis[J]. Current Opinion in Chemical Biology, 2018, 43: 106-112. |
| 20 | ALEXEEVA M, ENRIGHT A, DAWSON M J, et al. Deracemization of α-methylbenzylamine using an enzyme obtained by in vitro evolution[J]. Angewandte Chemie International Edition, 2002, 41(17):3177-3180. |
| 21 | CARR R, ALEXEEVA M, ENRIGHT A, et al. Directed evolution of an amine oxidase possessing both broad substrate specificity and high enantioselectivity[J]. Angewandte Chemie International Edition, 2003, 42(39):4807-4810. |
| 22 | MUTTI F G, KNAUS T, SCRUTTON N S, et al. Conversion of alcohols to enantiopure amines through dual-enzyme hydrogen-borrowing cascades[J]. Science, 2015, 349(6255):1525-1529. |
| 23 | ALEKU G A, FRANCE S P, MAN H, et al. A reductive aminase from Aspergillus oryzae [J]. Nature Chemistry, 2017, 9(10):961-969. |
| 24 | ZHU J M, TAN H Q, YANG L, et al. Enantioselective synthesis of 1-aryl-substituted tetrahydroisoquinolines employing imine reductase[J]. ACS Catalysis, 2017, 7(10): 7003-7007. |
| 25 | MANGAS-SANCHEZ J, FRANCE S P, MONTGOMERY S L, et al. Imine reductases (IREDs)[J]. Current Opinion in Chemical Biology, 2017, 37:19-25. |
| 26 | HEBERLING M M, WU B, BARTSCH S, et al. Priming ammonia lyases and aminomutases for industrial and therapeutic applications[J]. Current Opinion in Chemical Biology, 2013, 17(2):250-260. |
| 27 | LOVELOCK S L, LLOYD R C, TURNER N J. Phenylalanine ammonia lyase catalyzed synthesis of amino acids by an MIO- cofactor independent pathway[J]. Angewandte Chemie International Edition, 2014, 53(18):4652-4656. |
| 28 | GROGAN G, TURNER N J. InspIRED by nature: NADPH-dependent imine reductases (IREDs) as catalysts for the preparation of chiral amines[J]. Chemistry-A European Journal, 2016, 22(6):1900-1907. |
| 29 | POSNER B A, LI L Y, BETHELL R, et al. Engineering specificity for folate into dihydrofolate reductase from Escherichia coli [J]. Biochemistry, 1996, 35(5):1653-1663. |
| 30 | MENEELY K M, LAMB A L. Two structures of a thiazolinyl imine reductase from Yersinia enterocolitica provide insight into catalysis and binding to the nonribosomal peptide synthetase module of HMWP1 [J]. Biochemistry, 2012, 51(44):9002-9013. |
| 31 | MENEELY K M, RONNEBAUM T A, RILEY A P, et al. Holo structure and steady state kinetics of the thiazolinyl imine reductases for siderophore biosynthesis[J]. Biochemistry, 2016, 55(38):5423-5433. |
| 32 | WINZER T, KERN M, KING A J, et al. Morphinan biosynthesis in opium poppy requires a P450-oxidoreductase fusion protein[J]. Science, 2015, 349(6245):309-312. |
| 33 | FARROW S C, HAGEL J M, BEAUDOIN G A W, et al. Stereochemical inversion of (S)-reticuline by a cytochrome P450 fusion in opium poppy[J]. Nature Chemical Biology, 2015, 11(9): 728-732. |
| 34 | NARDINI M, RICCI G, CACCURI A M, et al. Purification and characterization of a ketimine-reducing enzyme[J]. European Journal of Biochemistry, 1988, 173(3): 689-694. |
| 35 | NARDINI M, RICCI G, VESCI L, et al. Bovine brain ketimine reductase[J]. Biochimica et Biophysica Acta, 1988, 957(2): 286-292. |
| 36 | HALLEN A, COOPER A J L, JAMIE J F, et al. Mammalian forebrain ketimine reductase identified as μ-crystallin; potential regulation by thyroid hormones[J]. Journal of Neurochemistry, 2011, 118(3): 379-387. |
| 37 | LI H, WILLIAMS P, MICKLEFIELD J, et al. A dynamic combinatorial screen for novel imine reductase activity[J]. Tetrahedron, 2004, 60(3): 753-758. |
| 38 | MURAMATSU H, MIHARA H, KAKUTANI R, et al. Enzymatic synthesis of N-methyl-L-phenylalanine by a novel enzyme, N-methyl-L-amino acid dehydrogenase, from Pseudomonas putida [J]. Tetrahedron: Asymmetry, 2004, 15(18): 2841-2843. |
| 39 | MURAMATSU H, MIHARA H, KAKUTANI R, et al. The putative malate/lactate dehydrogenase from Pseudomonas putida is an NADPH-dependent Δ¹-piperideine-2-carboxylate/Δ¹-pyrroline-2-carboxylate reductase involved in the catabolism of D-lysine and D-proline[J]. Journal of Biological Chemistry, 2005, 280(7):5329-5335. |
| 40 | MURAMATSU H, MIHARA H, GOTO M, et al. A new family of NAD(P)H-dependent oxidoreductases distinct from conventional Rossmann-fold proteins[J]. Journal of Bioscience and Bioengineering, 2005, 99(6):541-547. |
| 41 | VAIJAYANTHI T, CHADHA A. Asymmetric reduction of aryl imines using Candida parapsilosis ATCC 7330[J]. Tetrahedron: Asymmetry, 2008, 19(1): 93-96. |
| 42 | ESPINOZA-MORAGA M, PETTA T, VASQUEZ-VASQUEZ M, et al. Bioreduction of β-carboline imines to amines employing Saccharomyces bayanus [J]. Tetrahedron: Asymmetry, 2010, 21(16): 1988-1992. |
| 43 | MITSUKURA K, SUZUKI M, TADA K, et al. Asymmetric synthesis of chiral cyclic amine from cyclic imine by bacterial whole-cell catalyst of enantioselective imine reductase[J]. Organic & Biomolecular Chemistry, 2010, 8(20):4533-4535. |
| 44 | MITSUKURA K, SUZUKI M, SHINODA S, et al. Purification and characterization of a novel (R)-imine reductase from Streptomyces sp. GF3587 [J]. Bioscience, Biotechnology, and Biochemistry, 2011, 75(9): 1778-1782. |
| 45 | SLABU I, GALMAN J L, WEISE N J, et al. Putrescine transaminases for the synthesis of saturated nitrogen heterocycles from polyamines[J]. ChemCatChem, 2016, 8(6):1038-1042. |
| 46 | HUSSAIN S, LEIPOLD F, MAN H, et al. An (R)-imine reductase biocatalyst for the asymmetric reduction of cyclic imines[J]. ChemCatChem, 2015, 7(4):579-583. |
| 47 | LEIPOLD F, HUSSAIN S, GHISLIERI D, et al. Asymmetric reduction of cyclic imines catalyzed by a whole-cell biocatalyst containing an (S)-imine reductase[J]. ChemCatChem, 2013, 5(12): 3505-3508. |
| 48 | SCHELLER P N, FADEMRECHT S, HOFELZER S, et al. Enzyme toolbox: novel enantiocomplementary imine reductases[J]. ChemBioChem, 2014, 15(15):2201-2204. |
| 49 | FADEMRECHT S, SCHELLER P N, NESTL B M, et al. Identification of imine reductase-specific sequence motifs[J]. Proteins: Structure, Function, and Bioinformatics, 2016, 84(5): 600-610. |
| 50 | LENZ M, SCHELLER P N, RICHTER S M, et al. Cultivation and purification of two stereoselective imine reductases from Streptosporangium roseum and Paenibacillus elgii [J]. Protein Expression and Purification, 2017, 133:199-204. |
| 51 | MITSUKURA K, KURAMOTO T, YOSHIDA T, et al. A NADPH-dependent (S)-imine reductase (SIR) from Streptomyces sp. GF3546 for asymmetric synthesis of optically active amines: purification, characterization, gene cloning, and expression[J]. Applied Microbiology and Biotechnology, 2013, 97(18): 8079-8086. |
| 52 | LENZ M, BORLINGHAUS N, WEINMANN L, et al. Recent advances in imine reductase-catalyzed reactions[J]. World Journal of Microbiology & Biotechnology, 2017, 33(11): 199. |
| 53 | MIRABAL-GALLARDO Y, SORIANO M D P C, SANTOS L S. Stereoselective bioreduction of β-carboline imines through cell-free extracts from earthworms (Eisenia foetida)[J]. Tetrahedron: Asymmetry, 2013, 24(8): 440-443. |
| 54 | PENG H D, WEI E M, WANG J L, et al. Deciphering piperidine formation in polyketide-derived indolizidines reveals a thioester reduction, transamination, and unusual imine reduction process[J]. ACS Chemical Biology, 2016, 11(12):3278-3283. |
| 55 | LI H, ZHANG G X, LI L M, et al. A novel (R)-imine reductase from Paenibacillus lactis for asymmetric reduction of 3H-indoles[J]. ChemCatChem, 2016, 8(4): 724-727. |
| 56 | LI H, TIAN P, XU J H, et al. Identification of an imine reductase for asymmetric reduction of bulky dihydroisoquinolines[J]. Organic Letters, 2017, 19(12):3151-3154. |
| 57 | ZHANG Y H, CHEN F F, LI B B, et al. Stereocomplementary synthesis of pharmaceutically relevant chiral 2-aryl-substituted pyrrolidines using imine reductases[J]. Organic Letters, 2020, 22(9):3367-3372. |
| 58 | RODRÍGUEZ-MATA M, FRANK A, WELLS E, et al. Structure and activity of NADPH-dependent reductase Q1EQE0 from Streptomyces kanamyceticus, which catalyses the R-selective reduction of an imine substrate[J]. ChemBioChem, 2013, 14(11): 1372-1379. |
| 59 | LOKANATH N K, OHSHIMA N, TAKIO K, et al. Crystal structure of novel NADP-dependent 3-hydroxyisobutyrate dehydrogenase from Thermus thermophilus HB8[J]. Journal of Molecular Biology, 2005, 352(4): 905-917. |
| 60 | HUBER T, SCHNEIDER L, PRÄG A, et al. Direct reductive amination of ketones: structure and activity of S-selective imine reductases from Streptomyces [J]. ChemCatChem, 2014, 6(8): 2248-2252. |
| 61 | MAN H, WELLS E, HUSSAIN S, et al. Structure, activity and stereoselectivity of NADPH-dependent oxidoreductases catalysing the S-selective reduction of the imine substrate 2-methylpyrroline[J]. ChemBioChem, 2015, 16(7):1052-1059. |
| 62 | ALEKU G A, MAN H, FRANCE S P, et al. Stereoselectivity and structural characterization of an imine reductase (IRED) from Amycolatopsis orientalis [J]. ACS Catalysis, 2016, 6(6): 3880-3889. |
| 63 | MEYER T, ZUMBRÄGEL N, GEERDS C, et al. Structural characterization of an S-enantioselective imine reductase from Mycobacterium smegmatis [J]. Biomolecules, 2020, 10(8):1130-1142. |
| 64 | REETZ M T, KAHAKEAW D, LOHMER R. Addressing the numbers problem in directed evolution[J]. ChemBioChem, 2008, 9(11):1797-1804. |
| 65 | CURRIN A, SWAINSTON N, DAY P J, et al. Synthetic biology for the directed evolution of protein biocatalysts: navigating sequence space intelligently[J]. Chemical Society Reviews, 2015, 44(5):1172-1239. |
| 66 | SCHOBER M, MACDERMAID C, OLLIS A A, et al. Chiral synthesis of LSD1 inhibitor GSK2879552 enabled by directed evolution of an imine reductase[J]. Nature Catalysis, 2019, 2(10):909-915. |
| 67 | XU Z F, YAO P Y, SHENG X, et al. Biocatalytic access to 1,4-diazepanes via imine reductase-catalyzed intramolecular asymmetric reductive amination[J]. ACS Catalysis, 2020, 10(15):8780-8787. |
| 68 | FRANCE S P, ALEKU G A, SHARMA M, et al. Biocatalytic routes to enantiomerically enriched dibenz[c,e]azepines[J]. Angewandte Chemie International Edition, 2017, 56(49): 15589-15593. |
| 69 | RENATA H, WANG Z J, ARNOLD F H. Expanding the enzyme universe: accessing non-natural reactions by mechanism-guided directed evolution[J]. Angewandte Chemie International Edition, 2015, 54(11):3351-3367. |
| 70 | PORTER J L, RUSLI R A, OLLIS D L. Directed evolution of enzymes for industrial biocatalysis[J]. ChemBioChem, 2016, 17(3):197-203. |
| 71 | HESTERICOVÁ M, HEINISCH T, ALONSO-COTCHICO L, et al. Directed evolution of an artificial imine reductase[J]. Angewandte Chemie International Edition, 2018, 57(7): 1863-1868. |
| 72 | NEWTON C R, GRAHAM A, HEPTINSTALL L E, et al. Analysis of any point mutation in DNA. The amplification refractory mutation system (ARMS)[J]. Nucleic Acids Research, 1989, 17(7): 2503-2516. |
| 73 | MORLEY K L, KAZLAUSKAS R J. Improving enzyme properties: when are closer mutations better?[J]. Trends in Biotechnology, 2005, 23(5):231-237. |
| 74 | MINDT M, HANNIBAL S, HEUSER M, et al. Fermentative production of N-alkylated glycine derivatives by recombinant corynebacterium glutamicum using a mutant of imine reductase DpkA from Pseudomonas putida [J]. Frontiers in Bioengineering and Biotechnology, 2019, 7: 232. |
| 75 | PAVLIDIS I V, WEIß M S, GENZ M, et al. Identification of (S)-selective transaminases for the asymmetric synthesis of bulky chiral amines[J]. Nature Chemistry, 2016, 8(11): 1076-1082. |
| 76 | TANIHARA H, INOUE T, YAMAMOTO T, et al. Intra-ocular pressure-lowering effects of a Rho kinase inhibitor, ripasudil (K-115), over 24 hours in primary open-angle glaucoma and ocular hypertension: a randomized, open-label, crossover study[J]. Acta Ophthalmologica, 2015, 93(4): e254-e260. |
| 77 | GARWEG G, REHREN D V, HINTZE U. L-pipecolate formation in the mammalian brain. Regional distribution of Δ1-Pyrroline-2-carboxylate reductase activity[J]. Journal of Neurochemistry, 1980, 35(3): 616-621. |
| 78 | GAND M, THÖLE C, MÜLLER H, et al. A NADH-accepting imine reductase variant: immobilization and cofactor regeneration by oxidative deamination[J]. Journal of Biotechnology, 2016, 230: 11-18. |
| 79 | BORLINGHAUS N, NESTL B M. Switching the cofactor specificity of an imine reductase[J]. ChemCatChem, 2018, 10(1):183-187. |
| 80 | HEATH R S, PONTINI M, HUSSAIN S, et al. Combined imine reductase and amine oxidase catalyzed deracemization of nitrogen heterocycles[J]. ChemCatChem, 2016, 8(1):117-120. |
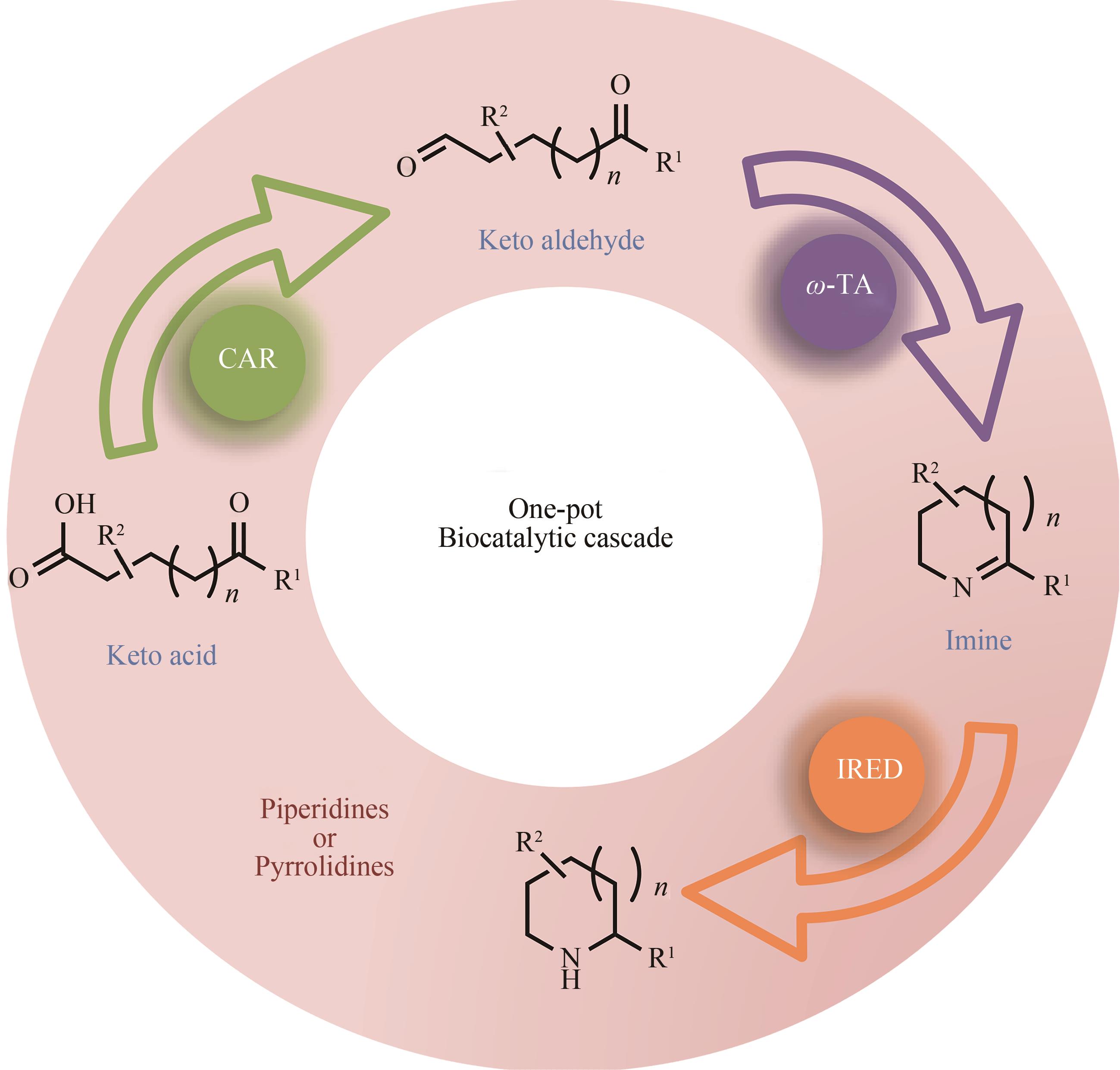
| 81 | FRANCE S P, HUSSAIN S, HILL A M, et al. One-pot cascade synthesis of mono- and disubstituted piperidines and pyrrolidines using carboxylic acid reductase (CAR), ω-transaminase (ω-TA), and imine reductase (IRED) biocatalysts[J]. ACS Catalysis, 2016, 6(6):3753-3759. |
| 82 | HEPWORTH L J, FRANCE S P, HUSSAIN S, et al. Enzyme cascades in whole cells for the synthesis of chiral cyclic amines[J]. ACS Catalysis, 2017, 7(4):2920-2925. |
| 83 | MONTGOMERY S L, MANGAS-SANCHEZ J, THOMPSON M P, et al. Direct alkylation of amines with primary and secondary alcohols through biocatalytic hydrogen borrowing[J]. Angewandte Chemie International Edition, 2017, 56(35): 10491-10494. |
| 84 | FORD G J, KRESS N, MATTEY A P, et al. Synthesis of protected 3-aminopiperidine and 3-aminoazepane derivatives using enzyme cascades[J]. Chemical Communications, 2020, 56(57):7949-7952. |
| 85 | COSGROVE S C, THOMPSON M P, AHMED S T, et al. One-pot synthesis of chiral N-arylamines by combining biocatalytic aminations with Buchwald-Hartwig N-arylation[J]. Angewandte Chemie International Edition, 2020, 59(41):18156-18160. |
| 86 | MARSHALL J R, YAO P Y, MONTGOMERY S L, et al. Screening and characterization of a diverse panel of metagenomic imine reductases for biocatalytic reductive amination[J]. Nature Chemistry, 2021, 13(2): 140-148. |
| 87 | HOPMANN K H, BAYER A. Enantioselective imine hydrogenation with iridium-catalysts: reactions, mechanisms and stereocontrol[J]. Coordination Chemistry Reviews, 2014, 268:59-82. |
| 88 | TANG W J, ZHANG X M. New chiral phosphorus ligands for enantioselective hydrogenation[J]. Chemical Reviews, 2003, 103(8): 3029-3070. |
| 89 | RIANT O, MOSTEFAÏ N, COURMARCEL J. Recent advances in the asymmetric hydrosilylation of ketones, imines and electrophilic double bonds[J]. Synthesis, 2004(18): 2943-2958. |
| 90 | VILAIVAN T, BHANTHUMNAVIN W, SRITANA-ANANT Y. Recent advances in catalytic asymmetric addition to imines and related C=N systems[J]. Current Organic Chemistry, 2005, 9(14):1315-1392. |
| 91 | MUSACCHIO A J, LAINHART B C, ZHANG X, et al. Catalytic intermolecular hydroaminations of unactivated olefins with secondary alkyl amines[J]. Science, 2017, 355(6326):727-730. |
| [1] | DONG Ying, MA Mengdan, HUANG Weiren. Progress in the miniaturization of CRISPR-Cas systems [J]. Synthetic Biology Journal, 2025, 6(1): 105-117. |
| [2] | WEN Yanhua, LIU Hedong, CAO Chunlai, WU Ruibo. Applications of protein engineering in pharmaceutical industry [J]. Synthetic Biology Journal, 2025, 6(1): 65-86. |
| [3] | ZHONG Quanzhou, SHAN Yiyi, PEI Qingyun, JIN Yanyun, WANG Yihan, MENG Luyuan, WANG Xinyun, ZHANG Yuxin, LIU Kunyuan, WANG Huizhong, FENG Shangguo. Research progress in the production of α-arbutin through biosynthesis [J]. Synthetic Biology Journal, 2025, 6(1): 118-135. |
| [4] | WANG Ziyuan, YANG Lirong, WU Jianping, ZHENG Wenlong. A review on enzyme-catalyzed synthesis of chiral amino acids [J]. Synthetic Biology Journal, 2024, 5(6): 1319-1349. |
| [5] | ZHU Fanghuan, CEN Xuecong, CHEN Zhen. Research progress of diols production by microbes [J]. Synthetic Biology Journal, 2024, 5(6): 1367-1385. |
| [6] | LIU Yining, PU Wei, YANG Jinxing, WANG Yu. Recent advances in the biosynthesis of ω-amino acids and lactams [J]. Synthetic Biology Journal, 2024, 5(6): 1350-1366. |
| [7] | CHENG Feng, ZOU Shuping, XU Jianmiao, TANG Heng, XUE Yaping, ZHENG Yuguo. BioHPP®: a benchmark of biomanufacturing for high optically pure L-phosphinothricin [J]. Synthetic Biology Journal, 2024, 5(6): 1404-1418. |
| [8] | FU Yu, ZHONG Fangrui. Recent advances in chemically driven enantioselective photobiocatalysis [J]. Synthetic Biology Journal, 2024, 5(5): 1021-1049. |
| [9] | ZHENG Haotian, LI Chaofeng, LIU Liangxu, WANG Jiawei, LI Hengrun, NI Jun. Design, optimization and application of synthetic carbon-negative phototrophic community [J]. Synthetic Biology Journal, 2024, 5(5): 1189-1210. |
| [10] | CHENG Xiaolei, LIU Tiangang, TAO Hui. Recent research progress in non-canonical biosynthesis of terpenoids [J]. Synthetic Biology Journal, 2024, 5(5): 1050-1071. |
| [11] | LIU Zijian, MU Baiyang, DUAN Zhiqiang, WANG Xuan, LU Xiaojie. Advances in the development of DNA-compatible chemistries [J]. Synthetic Biology Journal, 2024, 5(5): 1102-1124. |
| [12] | ZHANG Shouqi, WANG Tao, KONG Yao, ZOU Jiasheng, LIU Yuanning, XU Zhengren. Chemoenzymatic synthesis of natural products: evolution of synthetic methodology and strategy [J]. Synthetic Biology Journal, 2024, 5(5): 913-940. |
| [13] | XIE Xiangqian, GUO Wen, WANG Huan, LI Jin. Biosynthesis and chemical synthesis of ribosomally synthesized and post-translationally modified peptides containing aminovinyl cysteine [J]. Synthetic Biology Journal, 2024, 5(5): 981-996. |
| [14] | TANG Zhijun, HU Youcai, LIU Wen. Enzymatic (4+2)- and (2+2)-cycloaddition reactions: fundamentals and applications of regio- and stereoselectivity [J]. Synthetic Biology Journal, 2024, 5(3): 401-407. |
| [15] | ZHANG Jun, JIN Shixue, YUN Qian, QU Xudong. Biosynthesis of the unnatural extender units with polyketides and their structural modifications for applications in medicines [J]. Synthetic Biology Journal, 2024, 5(3): 561-570. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||