Synthetic Biology Journal ›› 2024, Vol. 5 ›› Issue (5): 1169-1188.DOI: 10.12211/2096-8280.2024-023
• Invited Review • Previous Articles Next Articles
Bioconversion of one carbon feedstocks for producing organic acids
YU Wei1,2, GAO Jiaoqi1,2, ZHOU Yongjin1,2
- 1.Division of Biotechnology,Dalian Institute of Chemical Physics,Chinese Academy of Sciences,Dalian 116023,Liaoning,China
2.Dalian Key Laboratory of Energy Biotechnology,Dalian 116023,Liaoning,China
-
Received:2024-03-19Revised:2024-06-04Online:2024-11-20Published:2024-10-31 -
Contact:ZHOU Yongjin
一碳生物转化合成有机酸的研究进展
禹伟1,2, 高教琪1,2, 周雍进1,2
- 1.中国科学院大连化学物理研究所,生物技术研究部,辽宁 大连 116023
2.大连市能源生物技术重点实验室,辽宁 大连 116023
-
通讯作者:周雍进 -
作者简介:禹伟 (1992—),男,博士。研究方向为多形汉逊酵母甲醇生物转化与代谢工程。 E-mail:yuweibio@dicp.ac.cn周雍进 (1984—),男,博士,研究员。研究方向为甲醇生物转化与天然产物生物合成。 E-mail:zhouyongjin@dicp.ac.cn -
基金资助:国家重点研发计划(2022YFC2105900);国家自然科学基金(22308351);国家资助博士后研究人员计划(GZB20230727)
CLC Number:
Cite this article
YU Wei, GAO Jiaoqi, ZHOU Yongjin. Bioconversion of one carbon feedstocks for producing organic acids[J]. Synthetic Biology Journal, 2024, 5(5): 1169-1188.
禹伟, 高教琪, 周雍进. 一碳生物转化合成有机酸的研究进展[J]. 合成生物学, 2024, 5(5): 1169-1188.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2024-023
| 产物 | 宿主 | 底物 | 培养条件 | 产量/(g/L) | 得率/% | 生产强度/[g/(L·d)] | 参考文献 |
|---|---|---|---|---|---|---|---|
| 3-HP | 蓝细菌 | CO2 | MM,摇瓶, 50 mL发酵体积 | 0.67 | 2.6 | 0.07 | [ |
| 蓝细菌 | CO2 | MM,摇瓶, 20 mL发酵体积 | 0.84 | 8.0 | 0.14 | [ | |
| 甲基弯菌 | 甲烷 | MM,发酵罐, 50 mL发酵体积 | 0.06 | 2.4 | 0.03 | [ | |
| 扭脱甲基杆菌 | 甲醇 | MM,摇瓶, 50 mL发酵体积 | 0.07 | 2.0 | 0.04 | [ | |
| 扭脱甲基杆菌 | 甲醇 | MM,发酵罐, 1.8 L发酵体积 | 0.86 | 3.1 | 0.21 | [ | |
| 多形汉逊酵母 | 甲醇 | MM,摇瓶, 50 mL发酵体积 | 7.10 | 14.2 | 1.20 | [ | |
| 巴斯德毕赤酵母 | 甲醇 | MM,发酵罐, 300 mL发酵体积 | 48.20 | 23.0 | 3.70 | [ | |
| L-乳酸 | 蓝细菌 | CO2 | MM,摇瓶, 80 mL发酵体积 | 1.00 | N.A. | 0.03 | [ |
| 甲烷氧化菌 | 甲烷 | MM,摇瓶, 2 mL发酵体积 | 0.60 | N.A. | 0.15 | [ | |
| 多形汉逊酵母 | 甲醇 | MM,摇瓶, 50 mL发酵体积 | 3.80 | 8.0 | 0.69 | [ | |
| D-乳酸 | 蓝细菌 | CO2 | MM,发酵罐, 100 mL发酵体积 | 1.30 | N.A. | 0.13 | [ |
| 甲基单胞菌 | 甲烷 | MM,摇瓶, 12.5 mL发酵体积 | 1.20 | 24.5 | 0.20 | [ | |
| 巴斯德毕赤酵母 | 甲醇 | CM,摇瓶, 5 mL发酵体积 | 3.50 | 22.0 | 0.87 | [ | |
| 琥珀酸 | 蓝细菌 | CO2 | MM,发酵罐, 发酵体积未知 | 0.93 | N.A. | 0.19 | [ |
| 蓝细菌 | CO2 | MM,摇瓶, 40 mL发酵体积 | 0.63 | N.A. | 0.32 | [ | |
| 蓝细菌 | CO2 | MM,摇瓶, 发酵体积未知 | 1.80 | N.A. | 0.60 | [ | |
| 蓝细菌 | CO2 | MM,发酵罐, 1 L发酵体积 | 2.50 | N.A. | 0.23 | [ | |
| 甲基单胞菌 | 甲烷 | MM,发酵罐, 3.2 L发酵体积 | 0.20 | 7.9 | 0.04 | [ |
Table 1 Bio-production of organic acids from C1 feedstocks
| 产物 | 宿主 | 底物 | 培养条件 | 产量/(g/L) | 得率/% | 生产强度/[g/(L·d)] | 参考文献 |
|---|---|---|---|---|---|---|---|
| 3-HP | 蓝细菌 | CO2 | MM,摇瓶, 50 mL发酵体积 | 0.67 | 2.6 | 0.07 | [ |
| 蓝细菌 | CO2 | MM,摇瓶, 20 mL发酵体积 | 0.84 | 8.0 | 0.14 | [ | |
| 甲基弯菌 | 甲烷 | MM,发酵罐, 50 mL发酵体积 | 0.06 | 2.4 | 0.03 | [ | |
| 扭脱甲基杆菌 | 甲醇 | MM,摇瓶, 50 mL发酵体积 | 0.07 | 2.0 | 0.04 | [ | |
| 扭脱甲基杆菌 | 甲醇 | MM,发酵罐, 1.8 L发酵体积 | 0.86 | 3.1 | 0.21 | [ | |
| 多形汉逊酵母 | 甲醇 | MM,摇瓶, 50 mL发酵体积 | 7.10 | 14.2 | 1.20 | [ | |
| 巴斯德毕赤酵母 | 甲醇 | MM,发酵罐, 300 mL发酵体积 | 48.20 | 23.0 | 3.70 | [ | |
| L-乳酸 | 蓝细菌 | CO2 | MM,摇瓶, 80 mL发酵体积 | 1.00 | N.A. | 0.03 | [ |
| 甲烷氧化菌 | 甲烷 | MM,摇瓶, 2 mL发酵体积 | 0.60 | N.A. | 0.15 | [ | |
| 多形汉逊酵母 | 甲醇 | MM,摇瓶, 50 mL发酵体积 | 3.80 | 8.0 | 0.69 | [ | |
| D-乳酸 | 蓝细菌 | CO2 | MM,发酵罐, 100 mL发酵体积 | 1.30 | N.A. | 0.13 | [ |
| 甲基单胞菌 | 甲烷 | MM,摇瓶, 12.5 mL发酵体积 | 1.20 | 24.5 | 0.20 | [ | |
| 巴斯德毕赤酵母 | 甲醇 | CM,摇瓶, 5 mL发酵体积 | 3.50 | 22.0 | 0.87 | [ | |
| 琥珀酸 | 蓝细菌 | CO2 | MM,发酵罐, 发酵体积未知 | 0.93 | N.A. | 0.19 | [ |
| 蓝细菌 | CO2 | MM,摇瓶, 40 mL发酵体积 | 0.63 | N.A. | 0.32 | [ | |
| 蓝细菌 | CO2 | MM,摇瓶, 发酵体积未知 | 1.80 | N.A. | 0.60 | [ | |
| 蓝细菌 | CO2 | MM,发酵罐, 1 L发酵体积 | 2.50 | N.A. | 0.23 | [ | |
| 甲基单胞菌 | 甲烷 | MM,发酵罐, 3.2 L发酵体积 | 0.20 | 7.9 | 0.04 | [ |
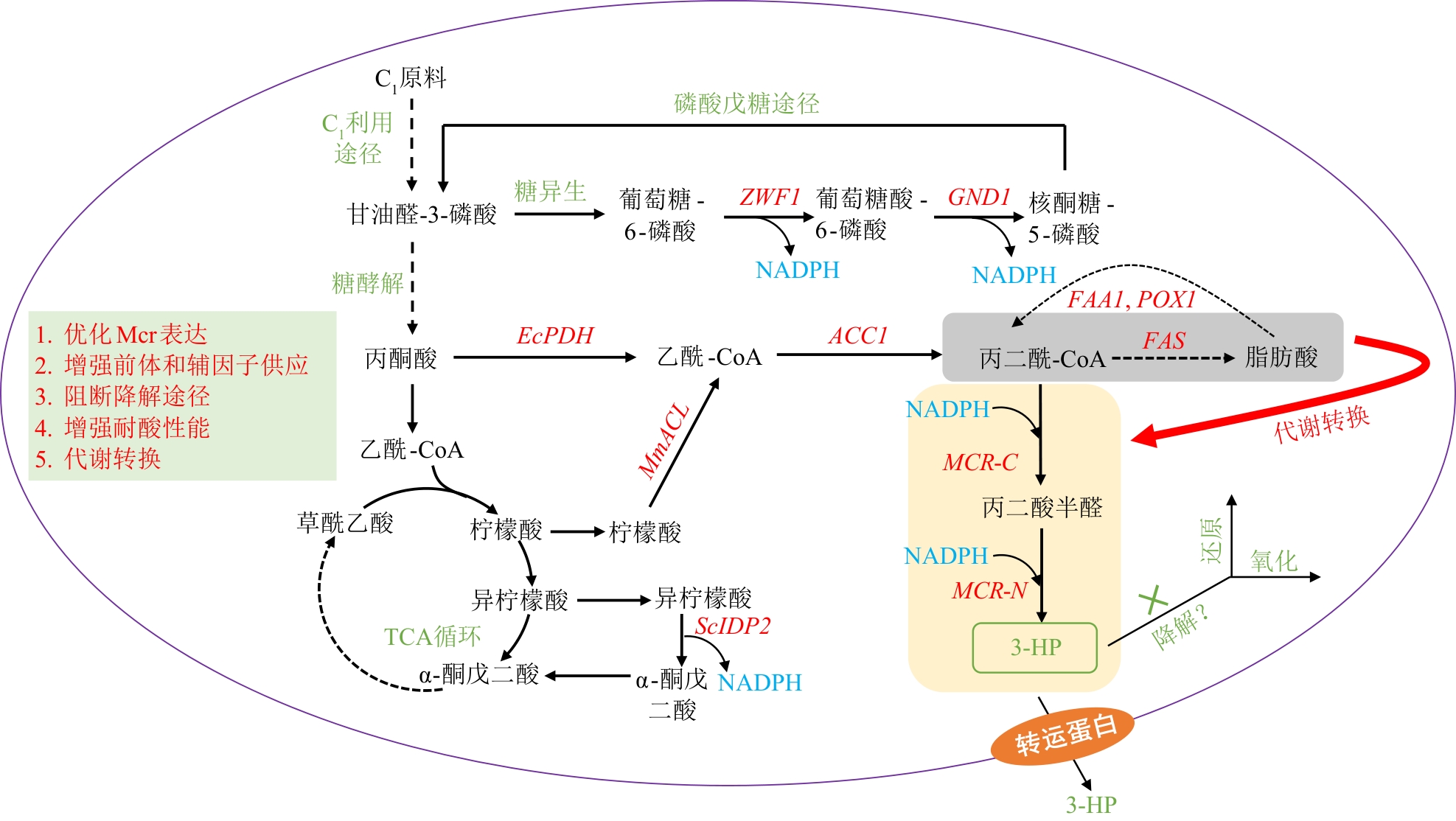
Fig. 2 Biosynthetic pathway and engineering strategies for 3-HP production from C1 feedstocksZWF1—Glucose-6-phosphate dehydrogenase gene; GND1—6-Phosphogluconate dehydrogenase gene; EcPDH—Escherichia coli pyruvate dehydrogenase complex gene; ACC1—Acetyl-CoA carboxylase gene; FAA1—Fatty acyl-CoA synthetase gene; POX1—Fatty acyl-CoA oxidase gene; FAS—Fatty acid synthase gene; MmACL—Mouse ATP-citrate lyase gene; ScIDP2—Saccharomyces cerevisiae isocitrate dehydrogenase gene; MCR-C—Malonyl-CoA reductase C-terminal gene; MCR-N—Malonyl-CoA reductase N-terminal gene
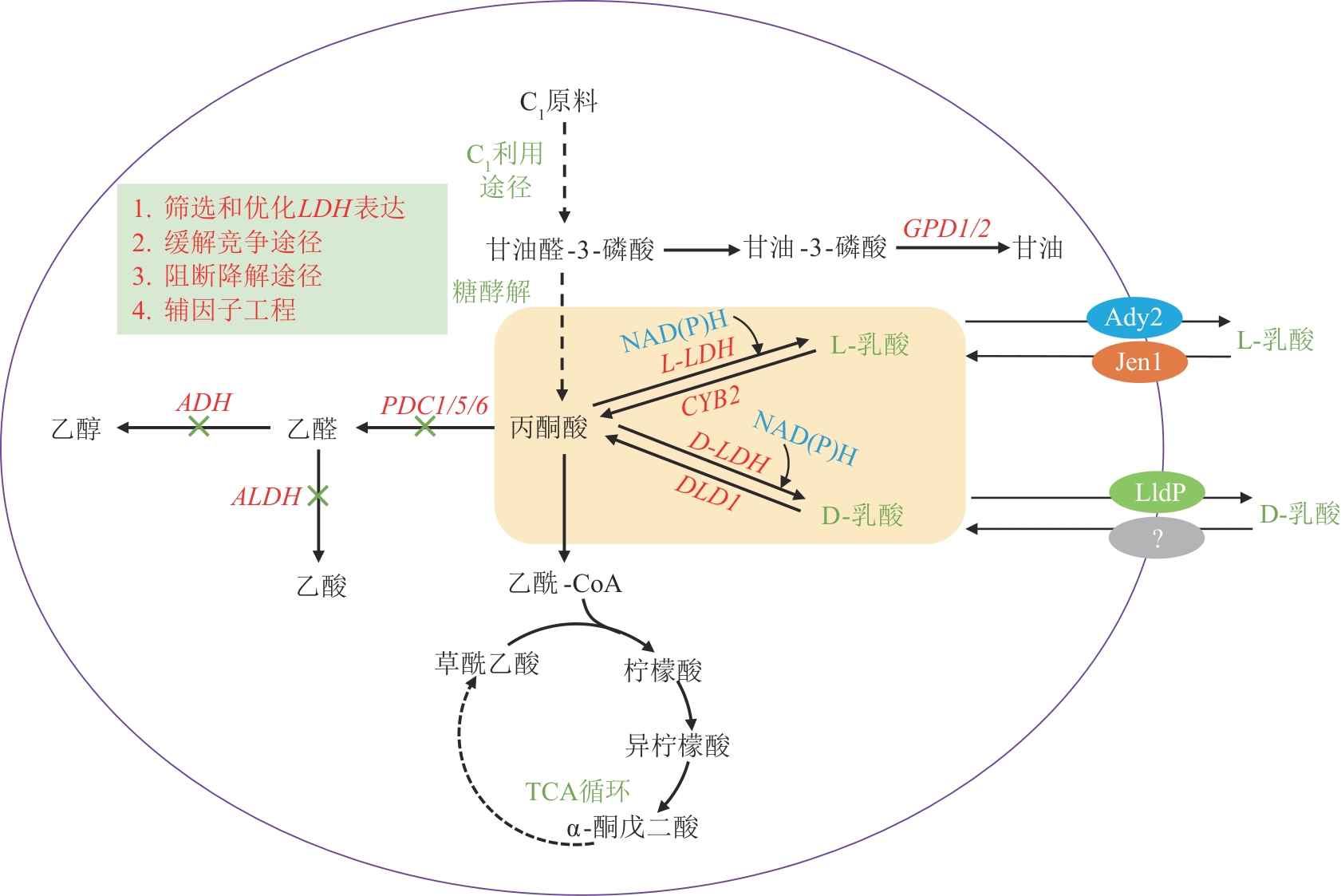
Fig. 3 Biosynthetic pathway and engineering strategies for lactic acid production from C1 feedstocksGPD1/2—Glycerol-3-phosphate dehydrogenase 1/2 genes; ADH—Alcohol dehydrogenase gene; PDC1/5/6—Pyruvate decarboxylase 1/5/6 genes; ALDH—Aldehyde dehydrogenase gene; L-LDH—l-lactate dehydrogenase gene; D-LDH—D-lactate dehydrogenase gene; CYB2—L-lactate dehydrogenase (cytochrome) gene; DLD1—D-lactate dehydrogenase gene; Ady2—Acetate transporter; Jen1—Monocarboxylate/H+ symporter; LldP—D-lactate transporter
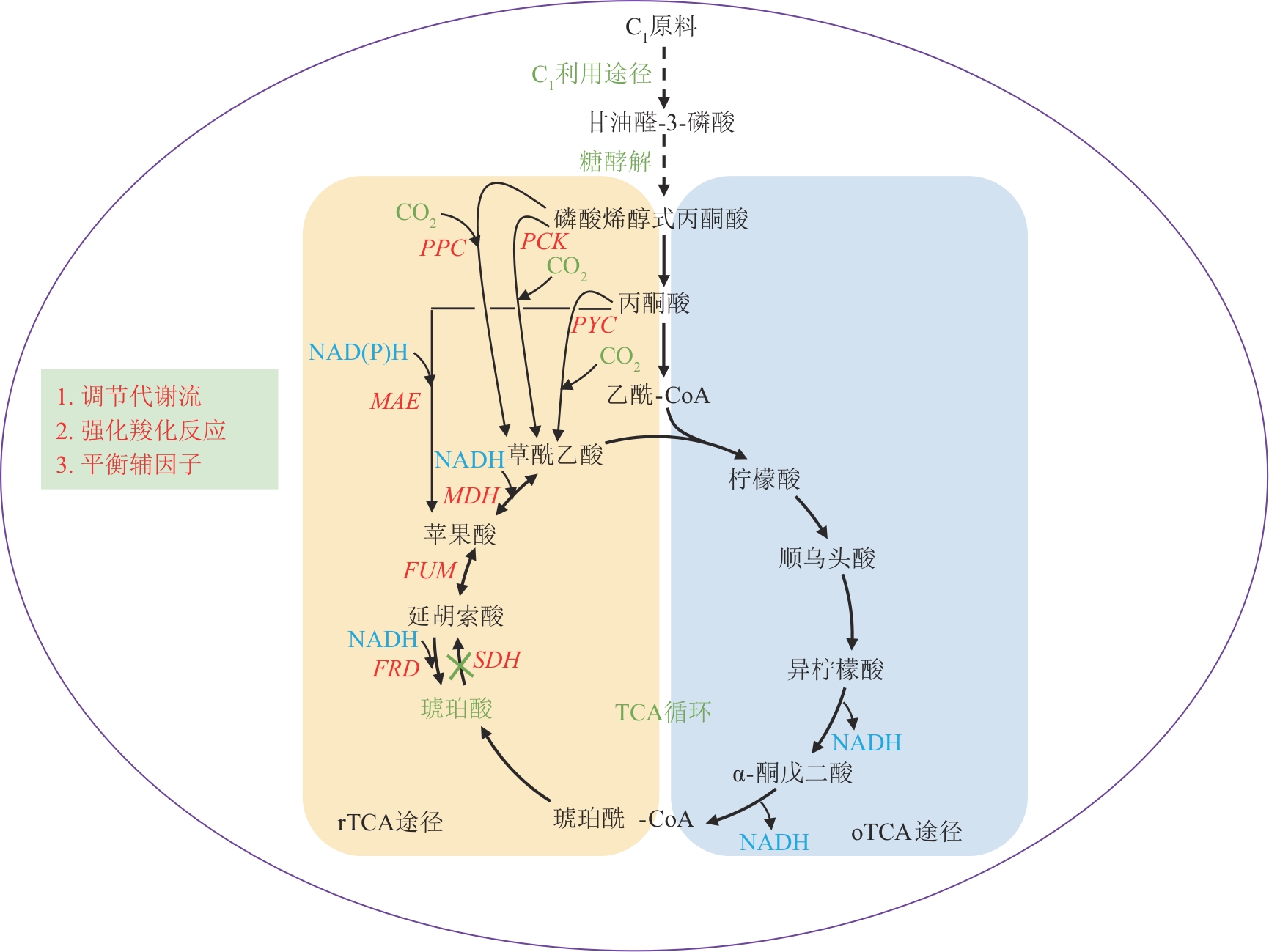
Fig. 4 Biosynthetic pathway and engineering strategies for succinic acid production from C1 feedstocksPYC—Pyruvate carboxylase gene; PPC—Phosphoenolpyruvate carboxylase gene; PCK—Phosphoenolpyruvate carboxykinase gene; MAE—Malic enzyme gene; MDH—Malate dehydrogenase gene; FUM—Fumarase gene; FRD—Fumarate reductase gene; SDH—Succinate dehydrogenase gene
| 1 | WERPY T, PETERSEN G. Top value added Chemicals from Biomass—VolumeⅠ: results of screening for potential candidates from sugars and synthesis Gas[R/OL]. Pacific Northwest National Laboratory National Renewable Energy Laboratory and Department of Energy. (2004-08-01)[2024-02-01]. . |
| 2 | BOZELL J J, PETERSEN G R. Technology development for the production of biobased products from biorefinery carbohydrates—the US Department of Energy’s “Top 10” revisited[J]. Green Chemistry, 2010, 12(4): 539-554. |
| 3 | 张瑞元, 朱翊凡, 曾杜文, 等. 利用酵母菌生产有机酸的研究进展[J]. 生物工程学报, 2023, 39(6): 2231-2247. |
| ZHANG R Y, ZHU Y F, ZENG D W, et al. Advances on the production of organic acids by yeast[J]. Chinese Journal of Biotechnology, 2023, 39(6): 2231-2247. | |
| 4 | LOMWONGSOPON P, VARRONE C. Contribution of fermentation technology to building blocks for renewable plastics[J]. Fermentation, 2022, 8(2): 47. |
| 5 | 张媛媛, 曾艳, 王钦宏. 合成生物制造进展[J]. 合成生物学, 2021, 2(2): 145-160. |
| ZHANG Y Y, ZENG Y, WANG Q H. Advances in synthetic biomanufacturing[J]. Synthetic Biology Journal, 2021, 2(2): 145-160. | |
| 6 | ZOU L H, OUYANG S P, HU Y L, et al. Efficient lactic acid production from dilute acid-pretreated lignocellulosic biomass by a synthetic consortium of engineered Pseudomonas putida and Bacillus coagulans [J]. Biotechnology for Biofuels, 2021, 14(1): 227. |
| 7 | LI Y, HUGENHOLTZ J, CHEN J, et al. Enhancement of pyruvate production by Torulopsis glabrata using a two-stage oxygen supply control strategy[J]. Applied Microbiology and Biotechnology, 2002, 60(1-2): 101-106. |
| 8 | YU W, CAO X, GAO J Q, et al. Overproduction of 3-hydroxypropionate in a super yeast chassis[J]. Bioresource Technology, 2022, 361: 127690. |
| 9 | ZHOU Y J, KERKHOVEN E J, NIELSEN J. Barriers and opportunities in bio-based production of hydrocarbons[J]. Nature Energy, 2018, 3(11): 925-935. |
| 10 | SANTOS CORREA S, SCHULTZ J, LAUERSEN K J, et al. Natural carbon fixation and advances in synthetic engineering for redesigning and creating new fixation pathways[J]. Journal of Advanced Research, 2023, 47: 75-92. |
| 11 | SARMA S, SHARMA S, RUDAKIYA D, et al. Valorization of microalgae biomass into bioproducts promoting circular bioeconomy: a holistic approach of bioremediation and biorefinery[J]. 3 Biotech, 2021, 11(8): 378. |
| 12 | VEAUDOR T, BLANC-GARIN V, CHENEBAULT C, et al. Recent advances in the photoautotrophic metabolism of cyanobacteria: biotechnological implications[J]. Life, 2020, 10(5): 71. |
| 13 | STEPHENS S, MAHADEVAN R, ALLEN D G. Engineering photosynthetic bioprocesses for sustainable chemical production: a review[J]. Frontiers in Bioengineering and Biotechnology, 2021, 8: 610723. |
| 14 | BAR-EVEN A, NOOR E, LEWIS N E, et al. Design and analysis of synthetic carbon fixation pathways[J]. Proceedings of the National Academy of Sciences of the United States of America, 2010, 107(19): 8889-8894. |
| 15 | SCHWANDER T, VON BORZYSKOWSKI L S, BURGENER S, et al. A synthetic pathway for the fixation of carbon dioxide in vitro [J]. Science, 2016, 354(6314): 900-904. |
| 16 | BOUZON M, PERRET A, LOREAU O, et al. A synthetic alternative to canonical one-carbon metabolism[J]. ACS Synthetic Biology, 2017, 6(8): 1520-1533. |
| 17 | LUO S S, DIEHL C, HE H, et al. Construction and modular implementation of the THETA cycle for synthetic CO2 fixation[J]. Nature Catalysis, 2023, 6(12): 1228-1240. |
| 18 | SARWAR A, LEE E Y. Methanol-based biomanufacturing of fuels and chemicals using native and synthetic methylotrophs[J]. Synthetic and Systems Biotechnology, 2023, 8(3): 396-415. |
| 19 | ZHAN C J, LI X W, YANG Y K, et al. Strategies and challenges with the microbial conversion of methanol to high-value chemicals[J]. Biotechnology and Bioengineering, 2021, 118(10): 3655-3668. |
| 20 | ZHAI X X, GAO J Q, LI Y X, et al. Peroxisomal metabolic coupling improves fatty alcohol production from sole methanol in yeast[J]. Proceedings of the National Academy of Sciences of the United States of America, 2023, 120(12): e2220816120. |
| 21 | SEMRAU J D, DISPIRITO A A, YOON S. Methanotrophs and copper[J]. FEMS Microbiology Reviews, 2010, 34(4): 496-531. |
| 22 | KALYUZHNAYA M G, PURI A W, LIDSTROM M E. Metabolic engineering in methanotrophic bacteria[J]. Metabolic Engineering, 2015, 29: 142-152. |
| 23 | BARIK S, PRIETO S, HARRISON S B, et al. Biological production of alcohols from coal through indirect liquefaction[J]. Applied Biochemistry and Biotechnology, 1988, 18(1): 363-378. |
| 24 | KÖPKE M, MIHALCEA C, LIEW F, et al. 2,3-Butanediol production by acetogenic bacteria, an alternative route to chemical synthesis, using industrial waste gas[J]. Applied and Environmental Microbiology, 2011, 77(15): 5467-5475. |
| 25 | FERNÁNDEZ-NAVEIRA Á, ABUBACKAR H N, VEIGA M C, et al. Efficient butanol-ethanol (B-E) production from carbon monoxide fermentation by Clostridium carboxidivorans [J]. Applied Microbiology and Biotechnology, 2016, 100(7): 3361-3370. |
| 26 | LITTY D, KREMP F, MÜLLER V. One substrate, many fates: different ways of methanol utilization in the acetogen Acetobacterium woodii [J]. Environmental Microbiology, 2022, 24(7): 3124-3133. |
| 27 | HEISKANEN H, VIRKAJÄRVI I, VIIKARI L. The effect of syngas composition on the growth and product formation of Butyribacterium methylotrophicum [J]. Enzyme and Microbial Technology, 2007, 41(3): 362-367. |
| 28 | SCHUCHMANN K, MÜLLER V. Autotrophy at the thermodynamic limit of life: a model for energy conservation in acetogenic bacteria[J]. Nature Reviews Microbiology, 2014, 12(12): 809-821. |
| 29 | GOYAL N, ZHOU Z, KARIMI I A. Metabolic processes of Methanococcus maripaludis and potential applications[J]. Microbial Cell Factories, 2016, 15(1): 107. |
| 30 | HEIDELBERG J F, SESHADRI R, HAVEMAN S A, et al. The genome sequence of the anaerobic, sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough[J]. Nature Biotechnology, 2004, 22(5): 554-559. |
| 31 | CROWTHER G J, KOSÁLY G, LIDSTROM M E. Formate as the main branch point for methylotrophic metabolism in Methylobacterium extorquens AM1[J]. Journal of Bacteriology, 2008, 190(14): 5057-5062. |
| 32 | CRAMM R. Genomic view of energy metabolism in Ralstonia eutropha H16[J]. Journal of Molecular Microbiology and Biotechnology, 2009, 16(1-2): 38-52. |
| 33 | GLEIZER S, BEN-NISSAN R, BAR-ON Y M, et al. Conversion of Escherichia coli to generate all biomass carbon from CO2 [J]. Cell, 2019, 179(6): 1255-1263.e12. |
| 34 | GASSLER T, SAUER M, GASSER B, et al. The industrial yeast Pichia pastoris is converted from a heterotroph into an autotroph capable of growth on CO2 [J]. Nature Biotechnology, 2020, 38(2): 210-216. |
| 35 | TIAN J Z, DENG W, ZHANG Z W, et al. Discovery and remodeling of Vibrio natriegens as a microbial platform for efficient formic acid biorefinery[J]. Nature Communications, 2023, 14(1): 7758. |
| 36 | HENRY C S, BROADBELT L J, HATZIMANIKATIS V. Discovery and analysis of novel metabolic pathways for the biosynthesis of industrial chemicals: 3-hydroxypropanoate[J]. Biotechnology and Bioengineering, 2010, 106(3): 462-473. |
| 37 | KILDEGAARD K R, HALLSTRÖM B M, BLICHER T H, et al. Evolution reveals a glutathione-dependent mechanism of 3-hydroxypropionic acid tolerance[J]. Metabolic Engineering, 2014, 26: 57-66. |
| 38 | SCHWARZ M, KÖPCKE B, WEBER R W, et al. 3-Hydroxypropionic acid as a nematicidal principle in endophytic fungi[J]. Phytochemistry, 2004, 65(15): 2239-2245. |
| 39 | CHEN Y, BAO J C, KIM I K, et al. Coupled incremental precursor and co-factor supply improves 3-hydroxypropionic acid production in Saccharomyces cerevisiae [J]. Metabolic Engineering, 2014, 22: 104-109. |
| 40 | LI Y, WANG X, GE X Z, et al. High production of 3-hydroxypropionic acid in Klebsiella pneumoniae by systematic optimization of glycerol metabolism[J]. Scientific Reports, 2016, 6: 26932. |
| 41 | BORODINA I, KILDEGAARD K R, JENSEN N B, et al. Establishing a synthetic pathway for high-level production of 3-hydroxypropionic acid in Saccharomyces cerevisiae via β-alanine[J]. Metabolic Engineering, 2015, 27: 57-64. |
| 42 | JIANG X R, YAN X, YU L P, et al. Hyperproduction of 3-hydroxypropionate by Halomonas bluephagenesis [J]. Nature Communications, 2021, 12(1): 1513. |
| 43 | TONG T, TAO Z Y, CHEN X L, et al. A biosynthesis pathway for 3-hydroxypropionic acid production in genetically engineered Saccharomyces cerevisiae [J]. Green Chemistry, 2021, 23(12): 4502-4509. |
| 44 | ZHAO P, MA C L, XU L D, et al. Exploiting tandem repetitive promoters for high-level production of 3-hydroxypropionic acid[J]. Applied Microbiology and Biotechnology, 2019, 103(10): 4017-4031. |
| 45 | WANG C, REN J, ZHOU L B, et al. An aldolase-catalyzed new metabolic pathway for the assimilation of formaldehyde and methanol to synthesize 2-keto-4-hydroxybutyrate and 1,3-propanediol in Escherichia coli [J]. ACS Synthetic Biology, 2019, 8(11): 2483-2493. |
| 46 | GAO J Q, YU W, LI Y X, et al. Engineering co-utilization of glucose and xylose for chemical overproduction from lignocellulose[J]. Nature Chemical Biology, 2023, 19(12): 1524-1531. |
| 47 | WANG Y P, SUN T, GAO X Y, et al. Biosynthesis of platform chemical 3-hydroxypropionic acid (3-HP) directly from CO2 in cyanobacterium Synechocystis sp. PCC 6803[J]. Metabolic Engineering, 2016, 34: 60-70. |
| 48 | NGUYEN D T N, LEE O K, LIM C, et al. Metabolic engineering of type Ⅱ methanotroph, Methylosinus trichosporium OB3b, for production of 3-hydroxypropionic acid from methane via a malonyl-CoA reductase-dependent pathway[J]. Metabolic Engineering, 2020, 59: 142-150. |
| 49 | WU X Y, CAI P, GAO L H, et al. Efficient bioproduction of 3-hydroxypropionic acid from methanol by a synthetic yeast cell factory[J]. ACS Sustainable Chemistry & Engineering, 2023, 11(16): 6445-6453. |
| 50 | LAN E I, CHUANG D S, SHEN C R, et al. Metabolic engineering of cyanobacteria for photosynthetic 3-hydroxypropionic acid production from CO2 using Synechococcus elongatus PCC 7942[J]. Metabolic Engineering, 2015, 31: 163-170. |
| 51 | YANG Y M, CHEN W J, YANG J, et al. Production of 3-hydroxypropionic acid in engineered Methylobacterium extorquens AM1 and its reassimilation through a reductive route[J]. Microbial Cell Factories, 2017, 16(1): 179. |
| 52 | YUAN X J, CHEN W J, MA Z X, et al. Rewiring the native methanol assimilation metabolism by incorporating the heterologous ribulose monophosphate cycle into Methylorubrum extorquens [J]. Metabolic Engineering, 2021, 64: 95-110. |
| 53 | YU W, GAO J Q, YAO L, et al. Bioconversion of methanol to 3-hydroxypropionate by engineering Ogataea polymorpha [J]. Chinese Journal of Catalysis, 2023, 46: 84-90. |
| 54 | SHABESTARY K, HERNÁNDEZ H P, MIAO R, et al. Cycling between growth and production phases increases cyanobacteria bioproduction of lactate[J]. Metabolic Engineering, 2021, 68: 131-141. |
| 55 | GARG S, CLOMBURG J M, GONZALEZ R. A modular approach for high-flux lactic acid production from methane in an industrial medium using engineered Methylomicrobium buryatense 5GB1[J]. Journal of Industrial Microbiology & Biotechnology, 2018, 45(6): 379-391. |
| 56 | WEFELMEIER K, SCHMITZ S, HAUT A M, et al. Engineering the methylotrophic yeast Ogataea polymorpha for lactate production from methanol[J]. Frontiers in Bioengineering and Biotechnology, 2023, 11: 1223726. |
| 57 | LI C, TAO F, NI J, et al. Enhancing the light-driven production of D-lactate by engineering cyanobacterium using a combinational strategy[J]. Scientific Reports, 2015, 5: 9777. |
| 58 | LEE J K, KIM S J, KIM W S, et al. Efficient production of D-lactate from methane in a lactate-tolerant strain of Methylomonas sp. DH-1 generated by adaptive laboratory evolution[J]. Biotechnology for Biofuels, 2019, 12: 234. |
| 59 | YAMADA R, OGURA K, KIMOTO Y, et al. Toward the construction of a technology platform for chemicals production from methanol: D-lactic acid production from methanol by an engineered yeast Pichia pastoris [J]. World Journal of Microbiology & Biotechnology, 2019, 35(2): 37. |
| 60 | SENGUPTA S, JAISWAL D, SENGUPTA A, et al. Metabolic engineering of a fast-growing Cyanobacterium Synechococcus elongatus PCC 11801 for photoautotrophic production of succinic acid[J]. Biotechnology for Biofuels, 2020, 13: 89. |
| 61 | HUANG C H, SHEN C R, LI H, et al. CRISPR interference (CRISPRi) for gene regulation and succinate production in cyanobacterium S. elongatus PCC 7942[J]. Microbial Cell Factories, 2016, 15(1): 196. |
| 62 | HASUNUMA T, MATSUDA M, KATO Y, et al. Temperature enhanced succinate production concurrent with increased central metabolism turnover in the cyanobacterium Synechocystis sp. PCC 6803[J]. Metabolic Engineering, 2018, 48: 109-120. |
| 63 | IIJIMA H, WATANABE A, SUKIGARA H, et al. Four-carbon dicarboxylic acid production through the reductive branch of the open cyanobacterial tricarboxylic acid cycle in Synechocystis sp. PCC 6803[J]. Metabolic Engineering, 2021, 65: 88-98. |
| 64 | NGUYEN D T N, LEE O K, HADIYATI S, et al. Metabolic engineering of the typeⅠmethanotroph Methylomonas sp. DH-1 for production of succinate from methane[J]. Metabolic Engineering, 2019, 54: 170-179. |
| 65 | LIU C S, WANG Q, XIAN M, et al. Dissection of malonyl-coenzyme A reductase of Chloroflexus aurantiacus results in enzyme activity improvement[J]. PLoS One, 2013, 8(9): e75554. |
| 66 | LIU C S, DING Y M, ZHANG R B, et al. Functional balance between enzymes in malonyl-CoA pathway for 3-hydroxypropionate biosynthesis[J]. Metabolic Engineering, 2016, 34: 104-111. |
| 67 | ZHOU Y J, BUIJS N A, ZHU Z W, et al. Production of fatty acid-derived oleochemicals and biofuels by synthetic yeast cell factories[J]. Nature Communications, 2016, 7: 11709. |
| 68 | YU T, ZHOU Y J, HUANG M T, et al. Reprogramming yeast metabolism from alcoholic fermentation to lipogenesis[J]. Cell, 2018, 174(6): 1549-1558.e14. |
| 69 | SCHNEIDER K, ASAO M, CARTER M S, et al. Rhodobacter sphaeroides uses a reductive route via propionyl coenzyme A to assimilate 3-hydroxypropionate[J]. Journal of Bacteriology, 2012, 194(2): 225-232. |
| 70 | ZHOU S F, ASHOK S, KO Y, et al. Development of a deletion mutant of Pseudomonas denitrificans that does not degrade 3-hydroxypropionic acid[J]. Applied Microbiology and Biotechnology, 2014, 98(10): 4389-4398. |
| 71 | NGUYEN-VO T P, RYU H, SAUER M, et al. Improvement of 3-hydroxypropionic acid tolerance in Klebsiella pneumoniae by novel transporter YohJK[J]. Bioresource Technology, 2022, 346: 126613. |
| 72 | NGUYEN-VO T P, LIANG Y X, SANKARANARAYANAN M, et al. Development of 3-hydroxypropionic-acid-tolerant strain of Escherichia coli W and role of minor global regulator yieP [J]. Metabolic Engineering, 2019, 53: 48-58. |
| 73 | CHUN A Y, YUNXIAO L, ASHOK S, et al. Elucidation of toxicity of organic acids inhibiting growth of Escherichia coli W[J]. Biotechnology and Bioprocess Engineering, 2014, 19(5): 858-865. |
| 74 | LI J, ZHU K, MIAO L, et al. Simultaneous improvement of limonene production and tolerance in Yarrowia lipolytica through tolerance engineering and evolutionary engineering[J]. ACS Synthetic Biology, 2021, 10(4): 884-896. |
| 75 | YANG M M, AN Y F, ZABED H M, et al. Random mutagenesis of Clostridium butyricum strain and optimization of biosynthesis process for enhanced production of 1,3-propanediol[J]. Bioresource Technology, 2019, 284: 188-196. |
| 76 | ZHANG W, GENG A L. Improved ethanol production by a xylose-fermenting recombinant yeast strain constructed through a modified genome shuffling method[J]. Biotechnology for Biofuels, 2012, 5(1): 46. |
| 77 | ZHU Y, ZHOU C, WANG Y, et al. Transporter engineering for microbial manufacturing[J]. Biotechnology Journal, 2020, 15(9): e1900494. |
| 78 | LIN Z L, ZHANG Y, WANG J Q. Engineering of transcriptional regulators enhances microbial stress tolerance[J]. Biotechnology Advances, 2013, 31(6): 986-991. |
| 79 | CHO J S, KIM G B, EUN H M, et al. Designing microbial cell factories for the production of chemicals[J]. JACS Au, 2022, 2(8): 1781-1799. |
| 80 | NGUYEN-VO T P, KO S, RYU H, et al. Systems evaluation reveals novel transporter YohJK renders 3-hydroxypropionate tolerance in Escherichia coli [J]. Scientific Reports, 2020, 10(1): 19064. |
| 81 | LIU D, HWANG H J, OTOUPAL P B, et al. Engineering Rhodosporidium toruloides for production of 3-hydroxypropionic acid from lignocellulosic hydrolysate[J]. Metabolic Engineering, 2023, 78: 72-83. |
| 82 | VINK E T H, RÁBAGO K R, GLASSNER D A, et al. Applications of life cycle assessment to NatureWorks™ polylactide (PLA) production[J]. Polymer Degradation and Stability, 2003, 80(3): 403-419. |
| 83 | DING X W, RONG J, PAN Z P, et al. De novo multienzyme synthetic pathways for lactic acid production[J]. ACS Catalysis, 2024, 14(7): 4665-4674. |
| 84 | UPADHYAYA B P, DEVEAUX L C, CHRISTOPHER L P. Metabolic engineering as a tool for enhanced lactic acid production[J]. Trends in Biotechnology, 2014, 32(12): 637-644. |
| 85 | HENARD C A, SMITH H, DOWE N, et al. Bioconversion of methane to lactate by an obligate methanotrophic bacterium[J]. Scientific Reports, 2016, 6: 21585. |
| 86 | YU W, GAO J Q, ZHAI X X, et al. Screening neutral sites for metabolic engineering of methylotrophic yeast Ogataea polymorpha [J]. Synthetic and Systems Biotechnology, 2021, 6(2): 63-68. |
| 87 | CAI P, DUAN X P, WU X Y, et al. Recombination machinery engineering facilitates metabolic engineering of the industrial yeast Pichia pastoris [J]. Nucleic Acids Research, 2021, 49(13): 7791-7805. |
| 88 | WANG M, LUAN G D, LU X F. Systematic identification of a neutral site on chromosome of Synechococcus sp. PCC7002, a promising photosynthetic chassis strain[J]. Journal of Biotechnology, 2019, 295: 37-40. |
| 89 | PORRO D, BIANCHI M M, BRAMBILLA L, et al. Replacement of a metabolic pathway for large-scale production of lactic acid from engineered yeasts[J]. Applied and Environmental Microbiology, 1999, 65(9): 4211-4215. |
| 90 | HIDESE R, MATSUDA M, OSANAI T, et al. Malic enzyme facilitates D-lactate production through increased pyruvate supply during anoxic dark fermentation in Synechocystis sp. PCC 6803[J]. ACS Synthetic Biology, 2020, 9(2): 260-268. |
| 91 | BIANCHI M M, BRAMBILLA L, PROTANI F, et al. Efficient homolactic fermentation by Kluyveromyces lactis strains defective in pyruvate utilization and transformed with the heterologous LDH gene[J]. Applied and Environmental Microbiology, 2001, 67(12): 5621-5625. |
| 92 | ANGERMAYR S A, VAN DER WOUDE A D, CORREDDU D, et al. Exploring metabolic engineering design principles for the photosynthetic production of lactic acid by Synechocystis sp. PCC6803[J]. Biotechnology for Biofuels, 2014, 7: 99. |
| 93 | BAEK S H, KWON E Y, KIM Y H, et al. Metabolic engineering and adaptive evolution for efficient production of D-lactic acid in Saccharomyces cerevisiae [J]. Applied Microbiology and Biotechnology, 2016, 100(6): 2737-2748. |
| 94 | PACHECO A, TALAIA G, SÁ-PESSOA J, et al. Lactic acid production in Saccharomyces cerevisiae is modulated by expression of the monocarboxylate transporters Jen1 and Ady2[J]. FEMS Yeast Research, 2012, 12(3): 375-381. |
| 95 | WAKAMATSU M, TOMITAKA M, TANI T, et al. Improvement of ethanol production from D-lactic acid by constitutive expression of lactate transporter Jen1p in Saccharomyces cerevisiae [J]. Bioscience, Biotechnology, and Biochemistry, 2013, 77(5): 1114-1116. |
| 96 | GUIARD B. Structure, expression and regulation of a nuclear gene encoding a mitochondrial protein: the yeast L(+)-lactate cytochrome c oxidoreductase (cytochrome b2)[J]. The EMBO Journal, 1985, 4(12): 3265-3272. |
| 97 | MOURIER A, VALLORTIGARA J, YOBOUE E D, et al. Kinetic activation of yeast mitochondrial D-lactate dehydrogenase by carboxylic acids[J]. Biochimica et Biophysica Acta (BBA) - Bioenergetics, 2008, 1777(10): 1283-1288. |
| 98 | BAUMSCHABL M, ATA Ö, MITIC B M, et al. Conversion of CO2 into organic acids by engineered autotrophic yeast[J]. Proceedings of the National Academy of Sciences of the United States of America, 2022, 119(47): e2211827119. |
| 99 | TONG T, CHEN X L, HU G P, et al. Engineering microbial metabolic energy homeostasis for improved bioproduction[J]. Biotechnology Advances, 2021, 53: 107841. |
| 100 | LEE J Y, KANG C D, LEE S H, et al. Engineering cellular redox balance in Saccharomyces cerevisiae for improved production of L-lactic acid[J]. Biotechnology and Bioengineering, 2015, 112(4): 751-758. |
| 101 | KOMATI REDDY G, LINDNER S N, WENDISCH V F. Metabolic engineering of an ATP-neutral Embden-Meyerhof-Parnas pathway in Corynebacterium glutamicum: growth restoration by an adaptive point mutation in NADH dehydrogenase[J]. Applied and Environmental Microbiology, 2015, 81(6): 1996-2005. |
| 102 | QI H S, LI S S, ZHAO S M, et al. Model-driven redox pathway manipulation for improved isobutanol production in Bacillus subtilis complemented with experimental validation and metabolic profiling analysis[J]. PLoS One, 2014, 9(4): e93815. |
| 103 | MULLINEAUX C W. Electron transport and light-harvesting switches in cyanobacteria[J]. Frontiers in Plant Science, 2014, 5: 7. |
| 104 | LIU X T, ZHAO G, SUN S J, et al. Biosynthetic pathway and metabolic engineering of succinic acid[J]. Frontiers in Bioengineering and Biotechnology, 2022, 10: 843887. |
| 105 | LIU Y P, ZHENG P, SUN Z H, et al. Economical succinic acid production from cane molasses by Actinobacillus succinogenes [J]. Bioresource Technology, 2008, 99(6): 1736-1742. |
| 106 | LEE P C, LEE S Y, HONG S H, et al. Batch and continuous cultures of Mannheimia succiniciproducens MBEL55E for the production of succinic acid from whey and corn steep liquor[J]. Bioprocess and Biosystems Engineering, 2003, 26(1): 63-67. |
| 107 | MEYNIAL-SALLES I, DOROTYN S, SOUCAILLE P. A new process for the continuous production of succinic acid from glucose at high yield, titer, and productivity[J]. Biotechnology and Bioengineering, 2008, 99(1): 129-135. |
| 108 | KUHNERT P, SCHOLTEN E, HAEFNER S, et al. Basfia succiniciproducens gen. nov., sp. nov., a new member of the family Pasteurellaceae isolated from bovine rumen[J]. International Journal of Systematic and Evolutionary Microbiology, 2010, 60(Pt 1): 44-50. |
| 109 | LEE S J, LEE D Y, KIM T Y, et al. Metabolic engineering of Escherichia coli for enhanced production of succinic acid, based on genome comparison and in silico gene knockout simulation[J]. Applied and Environmental Microbiology, 2005, 71(12): 7880-7887. |
| 110 | LITSANOV B, BROCKER M, BOTT M. Toward homosuccinate fermentation: metabolic engineering of Corynebacterium glutamicum for anaerobic production of succinate from glucose and formate[J]. Applied and Environmental Microbiology, 2012, 78(9): 3325-3337. |
| 111 | CUI Z Y, GAO C J, LI J J, et al. Engineering of unconventional yeast Yarrowia lipolytica for efficient succinic acid production from glycerol at low pH[J]. Metabolic Engineering, 2017, 42: 126-133. |
| 112 | LAI M J, TSAI J C, LAN E I. CRISPRi-enhanced direct photosynthetic conversion of carbon dioxide to succinic acid by metabolically engineered cyanobacteria[J]. Bioresource Technology, 2022, 366: 128131. |
| 113 | HASUNUMA T, MATSUDA M, KONDO A. Improved sugar-free succinate production by Synechocystis sp. PCC 6803 following identification of the limiting steps in glycogen catabolism[J]. Metabolic Engineering Communications, 2016, 3: 130-141. |
| 114 | LAN E I, WEI C T. Metabolic engineering of cyanobacteria for the photosynthetic production of succinate[J]. Metabolic Engineering, 2016, 38: 483-493. |
| 115 | LU S Y, EITEMAN M A, ALTMAN E. Effect of CO2 on succinate production in dual-phase Escherichia coli fermentations[J]. Journal of Biotechnology, 2009, 143(3): 213-223. |
| 116 | COTELESAGE J J H, PUTTICK J, GOLDIE H, et al. How does an enzyme recognize CO2?[J]. The International Journal of Biochemistry & Cell Biology, 2007, 39(6): 1204-1210. |
| 117 | XIAO M Y, ZHU X N, BI C H, et al. Improving succinate productivity by engineering a cyanobacterial CO2 concentrating system (CCM) in Escherichia coli [J]. Biotechnology Journal, 2017, 12(9): 1700199. |
| 118 | PRICE G D, WOODGER F J, BADGER M R, et al. Identification of a SulP-type bicarbonate transporter in marine cyanobacteria[J]. Proceedings of the National Academy of Sciences of the United States of America, 2004, 101(52): 18228-18233. |
| 119 | SHIBATA M, KATOH H, SONODA M, et al. Genes essential to sodium-dependent bicarbonate transport in cyanobacteria: function and phylogenetic analysis[J]. Journal of Biological Chemistry, 2002, 277(21): 18658-18664. |
| 120 | ZHU L W, ZHANG L, WEI L N, et al. Collaborative regulation of CO2 transport and fixation during succinate production in Escherichia coli [J]. Scientific Reports, 2015, 5: 17321. |
| 121 | DURALL C, KUKIL K, HAWKES J A, et al. Production of succinate by engineered strains of Synechocystis PCC 6803 overexpressing phosphoenolpyruvate carboxylase and a glyoxylate shunt[J]. Microbial Cell Factories, 2021, 20(1): 39. |
| 122 | TAPSCOTT T, GUARNIERI M T, HENARD C A. Development of a CRISPR/Cas9 system for Methylococcus capsulatus in vivo gene editing[J]. Applied and Environmental Microbiology, 2019, 85(11): e00340-19. |
| 123 | MO X H, ZHANG H, WANG T M, et al. Establishment of CRISPR interference in Methylorubrum extorquens and application of rapidly mining a new phytoene desaturase involved in carotenoid biosynthesis[J]. Applied Microbiology and Biotechnology, 2020, 104(10): 4515-4532. |
| 124 | SCHULTENKÄMPER K, BRITO L F, LÓPEZ M G, et al. Establishment and application of CRISPR interference to affect sporulation, hydrogen peroxide detoxification, and mannitol catabolism in the methylotrophic thermophile Bacillus methanolicus [J]. Applied Microbiology and Biotechnology, 2019, 103(14): 5879-5889. |
| 125 | GAO J Q, GAO N, ZHAI X X, et al. Recombination machinery engineering for precise genome editing in methylotrophic yeast Ogataea polymorpha [J]. iScience, 2021, 24(3): 102168. |
| 126 | ZHAI X X, JI L L, GAO J Q, et al. Characterizing methanol metabolism-related promoters for metabolic engineering of Ogataea polymorpha [J]. Applied Microbiology and Biotechnology, 2021, 105(23): 8761-8769. |
| 127 | NIELSEN J, KEASLING J D. Engineering cellular metabolism[J]. Cell, 2016, 164(6): 1185-1197. |
| 128 | LIEBAL U W, FABRY B A, RAVIKRISHNAN A, et al. Genome-scale model reconstruction of the methylotrophic yeast Ogataea polymorpha [J]. BMC Biotechnology, 2021, 21(1): 23. |
| 129 | KING Z A, LU J, DRÄGER A, et al. BiGG Models: a platform for integrating, standardizing and sharing genome-scale models[J]. Nucleic Acids Research, 2016, 44(D1): D515-D522. |
| 130 | SHIH C F, ZHANG T, LI J H, et al. Powering the future with liquid sunshine[J]. Joule, 2018, 2(10): 1925-1949. |
| [1] | GUO Shuyuan, ZHANG Qiannan, Gulikezi· MAIMAITIREXIATI, YANG Yiqun, YU Tao. Advances in microbial production of liquid biofuels [J]. Synthetic Biology Journal, 2025, 6(1): 18-44. |
| [2] | SHAO Mingwei, SUN Simian, YANG Shimao, CHEN Guoqiang. Bioproduction based on extremophiles [J]. Synthetic Biology Journal, 2024, 5(6): 1419-1436. |
| [3] | ZHAO Liang, LI Zhenshuai, FU Liping, LYU Ming, WANG Shi’an, ZHANG Quan, LIU Licheng, LI Fuli, LIU Ziyong. Progress in biomanufacturing of lipids and single cell protein from one-carbon compounds [J]. Synthetic Biology Journal, 2024, 5(6): 1300-1318. |
| [4] | ZHU Fanghuan, CEN Xuecong, CHEN Zhen. Research progress of diols production by microbes [J]. Synthetic Biology Journal, 2024, 5(6): 1367-1385. |
| [5] | CHEN Xiwei, ZHANG Huaran, ZOU Yi. Biosynthesis and metabolic engineering of fungal non-ribosomal peptides [J]. Synthetic Biology Journal, 2024, 5(3): 571-592. |
| [6] | XIE Huang, ZHENG Yilei, SU Yiting, RUAN Jingyi, LI Yongquan. An overview on reconstructing the biosynthetic system of actinomycetes for polyketides production [J]. Synthetic Biology Journal, 2024, 5(3): 612-630. |
| [7] | ZHAO Jingyu, ZHANG Jian, QI Qingsheng, WANG Qian. Research progress in biosensors based on bacterial two-component systems [J]. Synthetic Biology Journal, 2024, 5(1): 38-52. |
| [8] | SUN Huili, CUI Jinyu, LUAN Guodong, LYU Xuefeng. Progress of cyanobacterial synthetic biotechnology for efficient light-driven carbon fixation and ethanol production [J]. Synthetic Biology Journal, 2023, 4(6): 1161-1177. |
| [9] | YAN Xiongying, WANG Zhen, LOU Jiyun, ZHANG Haoyu, HUANG Xingyu, WANG Xia, YANG Shihui. Progress in the construction of microbial cell factories for efficient biofuel production [J]. Synthetic Biology Journal, 2023, 4(6): 1082-1121. |
| [10] | CHENG Zhenzhen, ZHANG Jian, GAO Cong, LIU Liming, CHEN Xiulai. Progress in metabolic engineering of microorganisms for the utilization of formate [J]. Synthetic Biology Journal, 2023, 4(4): 756-778. |
| [11] | Jiayu LIU, Zhihan YANG, Lei YANG, Liying ZHU, Zhengming ZHU, Ling JIANG. Advances in the development of Clostridium tyrobutyricum cell factories driven by synthetic biotechnology [J]. Synthetic Biology Journal, 2022, 3(6): 1174-1200. |
| [12] | Shuyuan GUO, Lianghuan WU, Xiangjian LIU, Bo WANG, Tao YU. Developing C1-based metabolic network in methylotrophy for biotransformation [J]. Synthetic Biology Journal, 2022, 3(1): 116-137. |
| [13] | Jiuzhou CHEN, Yu WANG, Wei PU, Ping ZHENG, Jibin SUN. Advances and perspective on bioproduction of 5-aminolevulinic acid [J]. Synthetic Biology Journal, 2021, 2(6): 1000-1016. |
| [14] | Qingzhuo WANG, Ping SONG, He HUANG. Synthetic biotechnology drives the development of natural eukaryotic lipid cell factories [J]. Synthetic Biology Journal, 2021, 2(6): 920-941. |
| [15] | Wei YAN, Hao GAO, Yujia JIANG, Xiujuan QIAN, Jie ZHOU, Weiliang DONG, Wenming ZHANG, Fengxue XIN, Min JIANG. Research progress in 2-phenylethanol production through biological processes [J]. Synthetic Biology Journal, 2021, 2(6): 1030-1045. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||