Synthetic Biology Journal ›› 2024, Vol. 5 ›› Issue (3): 561-570.DOI: 10.12211/2096-8280.2023-093
• Invited Review • Previous Articles Next Articles
Biosynthesis of the unnatural extender units with polyketides and their structural modifications for applications in medicines
ZHANG Jun1, JIN Shixue2, YUN Qian2, QU Xudong1
- 1.Zhangjiang Institute for Advanced Study,Shanghai Jiao Tong University,Shanghai 201204,China
2.School of Pharmacy,Fudan University,Shanghai 201203,China
-
Received:2023-11-30Revised:2024-01-08Online:2024-07-12Published:2024-06-30 -
Contact:QU Xudong
聚酮化合物非天然延伸单元的生物合成与结构改造应用
张俊1, 金诗雪2, 云倩2, 瞿旭东1
- 1.上海交通大学张江高等研究院,上海 201204
2.复旦大学药学院,上海 201203
-
通讯作者:瞿旭东 -
作者简介:张俊 (1992—),女,博士后。研究方向为聚酮化合物的生物合成与结构改造。E-mail:zhangjun2021@sjtu.edu.cn瞿旭东 (1980—),男,教授,博士生导师。研究方向为天然产物生物合成与生物催化,通过生物合成与化学合成结合的方式,发展针对各类天然产物骨架的普适、高效的合成与编辑策略,用于拓展结构多样性和提高药物的生产效率,实现对天然产物资源的深度开发及天然药物的高效创制。E-mail:quxd19@sjtu.edu.cn -
基金资助:国家自然科学基金青年科学基金(32200033)
CLC Number:
Cite this article
ZHANG Jun, JIN Shixue, YUN Qian, QU Xudong. Biosynthesis of the unnatural extender units with polyketides and their structural modifications for applications in medicines[J]. Synthetic Biology Journal, 2024, 5(3): 561-570.
张俊, 金诗雪, 云倩, 瞿旭东. 聚酮化合物非天然延伸单元的生物合成与结构改造应用[J]. 合成生物学, 2024, 5(3): 561-570.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2023-093

Fig. 1 Selected polyketide drugs (a) and the classical polyketide synthase assembly line for the biosynthesis of erythromycin A (b)AT—acyltransferase; ACP—acyl carrier protein; DEBS—6-deoxyerythronolide B synthase; DH—dehydratase; ER—enoylreductase; KR—ketoreductase; KS—ketosynthase; TE—thioesterase
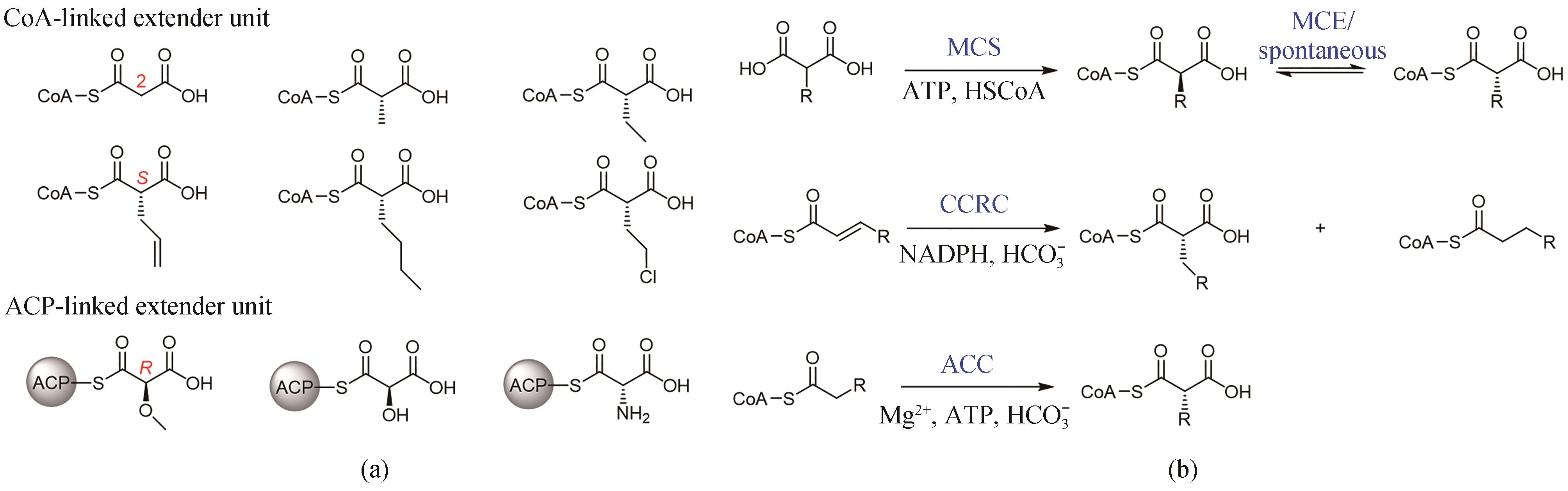
Fig. 2 Two classes of natural extender units (a) and the biosynthesis of malonyl-CoA extender units (b)MCS—malonyl-CoA synthetase; CCRC—crotonyl-CoA reductase/carboxylase; ACC—acyl-CoA carboxylase; MCE—methyl malonyl-CoA epimerase; CoA—coenzyme A; ACP—acyl carrier protein
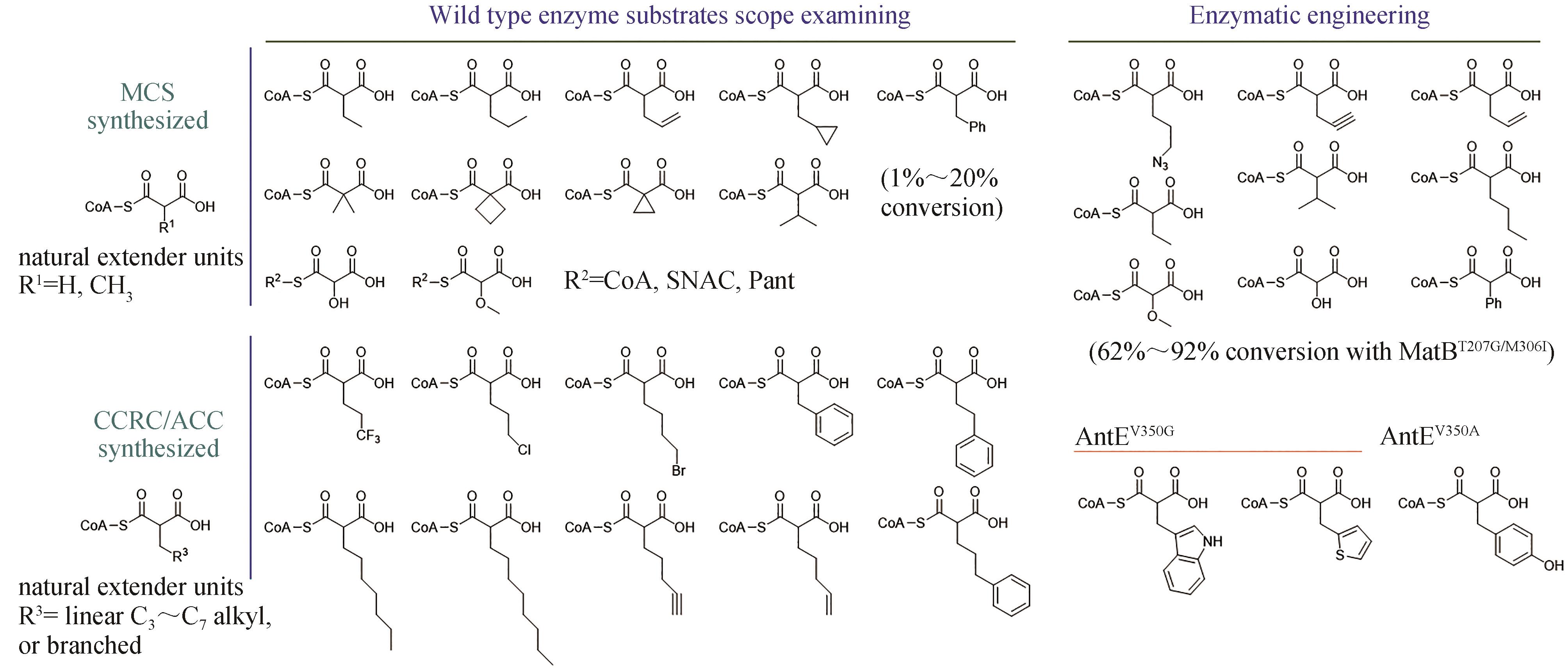
Fig. 3 Biosynthesis of unnatural extender units through engineering substrate spectrum or the enzymesMCS—malonyl-CoA synthetase; CCRC—crotonyl-CoA reductase/carboxylase; ACC—acyl-CoA carboxylase; CoA—Coenzyme A; SNAC—N-acetylcysteamine; Pant—pantetheine

Fig. 4 Modifications of polyketide sidechains through the biosynthesis of unnatural extender units with a natural promiscuous AT(Blue represents unnatural sidechains introduced by unnatural extender units)
| 1 | ROBERTSEN H L, MUSIOL-KROLL E M. Actinomycete-derived polyketides as a source of antibiotics and lead structures for the development of new antimicrobial drugs[J]. Antibiotics, 2019, 8(4): 157. |
| 2 | DAVISON E K, BRIMBLE M A. Natural product derived privileged scaffolds in drug discovery[J]. Current Opinion in Chemical Biology, 2019, 52: 1-8. |
| 3 | NEWMAN D J, CRAGG G M. Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019[J]. Journal of Natural Products, 2020, 83(3): 770-803. |
| 4 | LI G, LOU H X. Strategies to diversify natural products for drug discovery[J]. Medicinal Research Reviews, 2018, 38(4): 1255-1294. |
| 5 | KENNEDY J. Mutasynthesis, chemobiosynthesis, and back to semi-synthesis: combining synthetic chemistry and biosynthetic engineering for diversifying natural products[J]. Natural Product Reports, 2008, 25(1): 25-34. |
| 6 | LIN Z, QU X D. Emerging diversity in polyketide synthase[J]. Tetrahedron Letters, 2022, 110: 154183. |
| 7 | YAN X L, ZHANG J, TAN H Q, et al. A pair of atypical KAS Ⅲ homologues with initiation and elongation functions program the polyketide biosynthesis in Asukamycin[J]. Angewandte Chemie International Edition, 2022, 61(19): e202200879. |
| 8 | KEATINGE-CLAY A T. The structures of type Ⅰ polyketide synthases[J]. Natural Product Reports, 2012, 29(10): 1050-1073. |
| 9 | ROBBINS T, LIU Y C, CANE D E, et al. Structure and mechanism of assembly line polyketide synthases[J]. Current Opinion in Structural Biology, 2016, 41: 10-18. |
| 10 | HERTWECK C. The biosynthetic logic of polyketide diversity[J]. Angewandte Chemie International Edition, 2009, 48(26): 4688-4716. |
| 11 | FISCHBACH M A, WALSH C T. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: logic, machinery, and mechanisms[J]. Chemical Reviews, 2006, 106(8): 3468-3496. |
| 12 | WALKER P D, WEIR A N M, WILLIS C L, et al. Polyketide β-branching: diversity, mechanism and selectivity[J]. Natural Product Reports, 2021, 38(4): 723-756. |
| 13 | CHAN Y A, PODEVELS A M, KEVANY B M, et al. Biosynthesis of polyketide synthase extender units[J]. Natural Product Reports, 2009, 26(1): 90-114. |
| 14 | LITTLE R F, HERTWECK C. Chain release mechanisms in polyketide and non-ribosomal peptide biosynthesis[J]. Natural Product Reports, 2022, 39(1): 163-205. |
| 15 | MOORE B S, HERTWECK C. Biosynthesis and attachment of novel bacterial polyketide synthase starter units[J]. Natural Product Reports, 2002, 19(1): 70-99. |
| 16 | WILSON M C, MOORE B S. Beyond ethylmalonyl-CoA: the functional role of crotonyl-CoA carboxylase/reductase homologs in expanding polyketide diversity[J]. Natural Product Reports, 2012, 29(1): 72-86. |
| 17 | RAY L, MOORE B S. Recent advances in the biosynthesis of unusual polyketide synthase substrates[J]. Natural Product Reports, 2016, 33(2): 150-161. |
| 18 | BISSELL A U, RAUTSCHEK J, HOEFGEN S, et al. Biosynthesis of the sphingolipid inhibitors sphingofungins in filamentous fungi requires aminomalonate as a metabolic precursor[J]. ACS Chemical Biology, 2022, 17(2): 386-394. |
| 19 | ZHAO C H, COUGHLIN J M, JU J H, et al. Oxazolomycin biosynthesis in Streptomyces albus JA3453 featuring an “acyltransferase-less” type Ⅰ polyketide synthase that incorporates two distinct extender units[J]. Journal of Biological Chemistry, 2010, 285(26): 20097-20108. |
| 20 | CHEN D D, ZHANG Q, ZHANG Q L, et al. Improvement of FK506 production in Streptomyces tsukubaensis by genetic enhancement of the supply of unusual polyketide extender units via utilization of two distinct site-specific recombination systems[J]. Applied and Environmental Microbiology, 2012, 78(15): 5093-5103. |
| 21 | KATO Y, BAI L Q, XUE Q, et al. Functional expression of genes involved in the biosynthesis of the novel polyketide chain extension unit, methoxymalonyl-acyl carrier protein, and engineered biosynthesis of 2-desmethyl-2-methoxy-6-deoxyerythronolide B[J]. Journal of the American Chemical Society, 2002, 124(19): 5268-5269. |
| 22 | CARPENTER S M, WILLIAMS G J. Extender unit promiscuity and orthogonal protein interactions of an aminomalonyl-ACP utilizing trans-acyltransferase from zwittermicin biosynthesis[J]. ACS Chemical Biology, 2018, 13(12): 3361-3373. |
| 23 | DODGE G J, MALONEY F P, SMITH J L. Protein-protein interactions in “cis-AT” polyketide synthases[J]. Natural Product Reports, 2018, 35(10): 1082-1096. |
| 24 | KOSOL S, JENNER M, LEWANDOWSKI J R, et al. Protein-protein interactions in trans-AT polyketide synthases[J]. Natural Product Reports, 2018, 35(10): 1097-1109. |
| 25 | ZHANG F, JI H N, ALI I, et al. Structural and biochemical insight into the recruitment of acyl carrier protein-linked extender units in ansamitocin biosynthesis[J]. Chembiochem, 2020, 21(9): 1309-1314. |
| 26 | ZHENG M M, ZHANG J, ZHANG W, et al. An atypical acyl-CoA synthetase enables efficient biosynthesis of extender units for engineering a polyketide carbon scaffold[J]. Angewandte Chemie International Edition, 2022, 61(43): e202208734. |
| 27 | AN J H, KIM Y S. A gene cluster encoding malonyl-CoA decarboxylase (MatA), malonyl-CoA synthetase (MatB) and a putative dicarboxylate carrier protein (MatC) in Rhizobium trifolii: cloning, sequencing, and expression of the enzymes in Escherichia coli [J]. European Journal of Biochemistry, 1998, 257(2): 395-402. |
| 28 | POHL N L, HANS M, LEE H Y, et al. Remarkably broad substrate tolerance of malonyl-CoA synthetase, an enzyme capable of intracellular synthesis of polyketide precursors[J]. Journal of the American Chemical Society, 2001, 123(24): 5822-5823. |
| 29 | HUGHES A J, KEATINGE-CLAY A. Enzymatic extender unit generation for in vitro polyketide synthase reactions: structural and functional showcasing of Streptomyces coelicolor MatB[J]. Chemistry & Biology, 2011, 18(2): 165-176. |
| 30 | ERB T J, BERG I A, BRECHT V, et al. Synthesis of C5-dicarboxylic acids from C2-units involving crotonyl-CoA carboxylase/reductase: the ethylmalonyl-CoA pathway[J]. Proceedings of the National Academy of Sciences of the United States of America, 2007, 104(25): 10631-10636. |
| 31 | YAN Y, CHEN J, ZHANG L H, et al. Multiplexing of combinatorial chemistry in antimycin biosynthesis: expansion of molecular diversity and utility[J]. Angewandte Chemie International Edition, 2013, 52(47): 12308-12312. |
| 32 | VÖGELI B, GEYER K, GERLINGER P D, et al. Combining promiscuous acyl-CoA oxidase and enoyl-CoA carboxylase/reductases for atypical polyketide extender unit biosynthesis[J]. Cell Chemical Biology, 2018, 25(7): 833-839.e4. |
| 33 | RAY L, VALENTIC T R, MIYAZAWA T, et al. A crotonyl-CoA reductase-carboxylase independent pathway for assembly of unusual alkylmalonyl-CoA polyketide synthase extender units[J]. Nature Communications, 2016, 7: 13609. |
| 34 | ZHANG J, ZHENG M M, YAN J Y, et al. A permissive medium chain acyl-CoA carboxylase enables the efficient biosynthesis of extender units for engineering polyketide carbon scaffolds[J]. ACS Catalysis, 2021, 11(19): 12179-12185. |
| 35 | TRAN T H, HSIAO Y S, JO J, et al. Structure and function of a single-chain, multi-domain long-chain acyl-CoA carboxylase[J]. Nature, 2015, 518(7537): 120-124. |
| 36 | FARINAS E T, BULTER T, ARNOLD F H. Directed enzyme evolution[J]. Current Opinion in Biotechnology, 2001, 12(6): 545-551. |
| 37 | NODA-GARCIA L, TAWFIK D S. Enzyme evolution in natural products biosynthesis: target- or diversity-oriented?[J]. Current Opinion in Chemical Biology, 2020, 59: 147-154. |
| 38 | KORYAKINA I, WILLIAMS G J. Mutant malonyl-CoA synthetases with altered specificity for polyketide synthase extender unit generation[J]. Chembiochem, 2011, 12(15): 2289-2293. |
| 39 | KORYAKINA I, MCARTHUR J, RANDALL S, et al. Poly specific trans-acyltransferase machinery revealed via engineered acyl-CoA synthetases[J]. ACS Chemical Biology, 2013, 8(1): 200-208. |
| 40 | ZHANG L H, MORI T, ZHENG Q F, et al. Rational control of polyketide extender units by structure-based engineering of a crotonyl-CoA carboxylase/reductase in antimycin biosynthesis[J]. Angewandte Chemie International Edition, 2015, 54(45): 13462-13465. |
| 41 | QUADE N, HUO L J, RACHID S, et al. Unusual carbon fixation gives rise to diverse polyketide extender units[J]. Nature Chemical Biology, 2012, 8(1): 117-124. |
| 42 | TIAN W Y, CHEN X R, ZHANG J, et al. Biosynthesis of tetronates by a nonribosomal peptide synthetase-polyketide synthase system[J]. Organic Letters, 2023, 25(10): 1628-1632. |
| 43 | AWAKAWA T, FUJIOKA T, ZHANG L H, et al. Reprogramming of the antimycin NRPS-PKS assembly lines inspired by gene evolution[J]. Nature Communications, 2018, 9(1): 3534. |
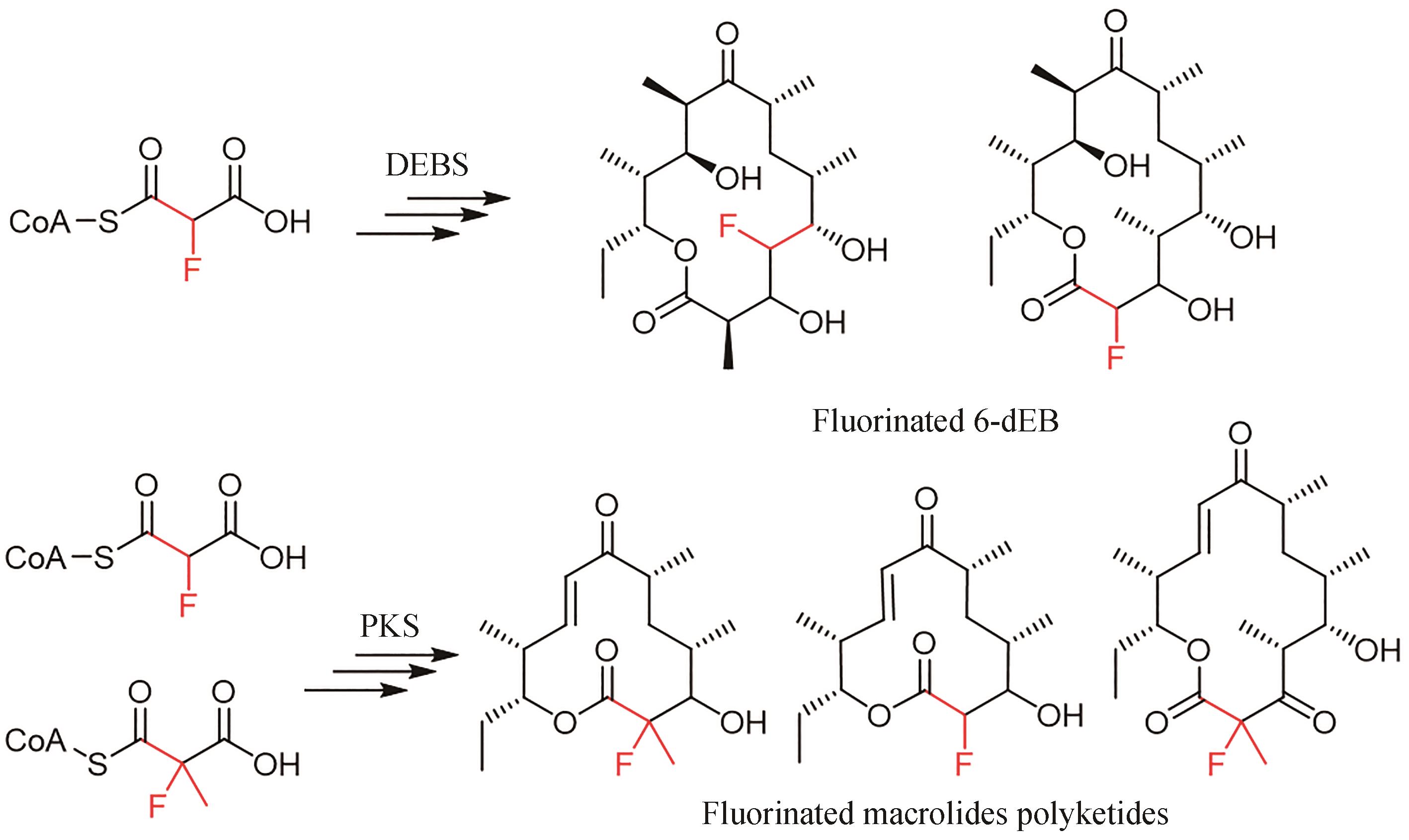
| 44 | WALKER M C, THURONYI B W, CHARKOUDIAN L K, et al. Expanding the fluorine chemistry of living systems using engineered polyketide synthase pathways[J]. Science, 2013, 341(6150): 1089-1094. |
| 45 | SIRIRUNGRUANG S, AD O, PRIVALSKY T M, et al. Engineering site-selective incorporation of fluorine into polyketides[J]. Nature Chemical Biology, 2022, 18(8): 886-893. |
| 46 | RITTNER A, JOPPE M, SCHMIDT J J, et al. Chemoenzymatic synthesis of fluorinated polyketides[J]. Nature Chemistry, 2022, 14(9): 1000-1006. |
| 47 | LI Y, ZHANG W, ZHANG H, et al. Structural basis of a broadly selective acyltransferase from the polyketide synthase of splenocin[J]. Angewandte Chemie International Edition, 2018, 57(20): 5823-5827. |
| 48 | KALKREUTER E, CROWETIPTON J M, LOWELL A N, et al. Engineering the substrate specificity of a modular polyketide synthase for installation of consecutive non-natural extender units[J]. Journal of the American Chemical Society, 2019, 141(5): 1961-1969. |
| 49 | ENGLUND E, SCHMIDT M, NAVA A A, et al. Expanding extender substrate selection for unnatural polyketide biosynthesis by acyltransferase domain exchange within a modular polyketide synthase[J]. Journal of the American Chemical Society, 2023, 145(16): 8822-8832. |
| 50 | MUSIOL-KROLL E M, WOHLLEBEN W. Acyltransferases as tools for polyketide synthase engineering[J]. Antibiotics, 2018, 7(3): 62. |
| 51 | CHANG C C, HUANG R, YAN Y, et al. Uncovering the formation and selection of benzylmalonyl-CoA from the biosynthesis of splenocin and enterocin reveals a versatile way to introduce amino acids into polyketide carbon scaffolds[J]. Journal of the American Chemical Society, 2015, 137(12): 4183-4190. |
| 52 | ZHU X J, LIU J, ZHANG W J. De novo biosynthesis of terminal alkyne-labeled natural products[J]. Nature Chemical Biology, 2015, 11(2): 115-120. |
| [1] | ZHONG Quanzhou, SHAN Yiyi, PEI Qingyun, JIN Yanyun, WANG Yihan, MENG Luyuan, WANG Xinyun, ZHANG Yuxin, LIU Kunyuan, WANG Huizhong, FENG Shangguo. Research progress in the production of α-arbutin through biosynthesis [J]. Synthetic Biology Journal, 2025, 6(1): 118-135. |
| [2] | ZHU Fanghuan, CEN Xuecong, CHEN Zhen. Research progress of diols production by microbes [J]. Synthetic Biology Journal, 2024, 5(6): 1367-1385. |
| [3] | LIU Yining, PU Wei, YANG Jinxing, WANG Yu. Recent advances in the biosynthesis of ω-amino acids and lactams [J]. Synthetic Biology Journal, 2024, 5(6): 1350-1366. |
| [4] | ZHENG Haotian, LI Chaofeng, LIU Liangxu, WANG Jiawei, LI Hengrun, NI Jun. Design, optimization and application of synthetic carbon-negative phototrophic community [J]. Synthetic Biology Journal, 2024, 5(5): 1189-1210. |
| [5] | CHENG Xiaolei, LIU Tiangang, TAO Hui. Recent research progress in non-canonical biosynthesis of terpenoids [J]. Synthetic Biology Journal, 2024, 5(5): 1050-1071. |
| [6] | LIU Zijian, MU Baiyang, DUAN Zhiqiang, WANG Xuan, LU Xiaojie. Advances in the development of DNA-compatible chemistries [J]. Synthetic Biology Journal, 2024, 5(5): 1102-1124. |
| [7] | ZHANG Shouqi, WANG Tao, KONG Yao, ZOU Jiasheng, LIU Yuanning, XU Zhengren. Chemoenzymatic synthesis of natural products: evolution of synthetic methodology and strategy [J]. Synthetic Biology Journal, 2024, 5(5): 913-940. |
| [8] | XIE Xiangqian, GUO Wen, WANG Huan, LI Jin. Biosynthesis and chemical synthesis of ribosomally synthesized and post-translationally modified peptides containing aminovinyl cysteine [J]. Synthetic Biology Journal, 2024, 5(5): 981-996. |
| [9] | TANG Zhijun, HU Youcai, LIU Wen. Enzymatic (4+2)- and (2+2)-cycloaddition reactions: fundamentals and applications of regio- and stereoselectivity [J]. Synthetic Biology Journal, 2024, 5(3): 401-407. |
| [10] | Xiwei CHEN, Huaran ZHANG, Yi ZOU. Biosynthesis and metabolic engineering of fungal non-ribosomal peptides [J]. Synthetic Biology Journal, 2024, 5(3): 571-592. |
| [11] | YU Xuchang, WU Hui, LI Lei. Library construction and targeted BGC screening for more efficient discovery of microbial natural products [J]. Synthetic Biology Journal, 2024, 5(3): 492-506. |
| [12] | FENG Jin, PAN Haixue, TANG Gongli. Research advances in biosynthesis of natural product drugs within the past decade [J]. Synthetic Biology Journal, 2024, 5(3): 408-446. |
| [13] | XI Mengyu, HU Yiling, GU Yucheng, GE Huiming. Genome mining-directed discovery for natural medicinal products [J]. Synthetic Biology Journal, 2024, 5(3): 447-473. |
| [14] | LEI Ru, TAO Hui, LIU Tiangang. Deep genome mining boosts the discovery of microbial terpenoids [J]. Synthetic Biology Journal, 2024, 5(3): 507-526. |
| [15] | Xinjie SHI, Yiling DU. Research advances in the biosynthesis of nonribosomal peptides within the bisintercalator family as anticancer drugs [J]. Synthetic Biology Journal, 2024, 5(3): 593-611. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||