Synthetic Biology Journal ›› 2022, Vol. 3 ›› Issue (4): 626-637.DOI: 10.12211/2096-8280.2021-087
• Invited Review • Previous Articles Next Articles
Biofilm matrixes-from soft matters to engineered materials
ZHU Runtao, ZHONG Chao, DAI Zhuojun
- Institute of Synthetic Biology,Shenzhen Institude of Advanced Technology,Chinese Acdamey of Science,Shenzhen 518055,Guangdong,China
-
Received:2021-08-27Revised:2021-12-22Online:2022-09-08Published:2022-08-31 -
Contact:ZHONG Chao, DAI Zhuojun
细菌生物被膜的软物质特性及其工程化应用
朱润涛, 钟超, 戴卓君
- 中国科学院深圳先进技术研究院合成生物学研究所,广东 深圳 518055
-
通讯作者:钟超,戴卓君 -
作者简介:朱润涛 (1993—),男,硕士。研究方向为基于工程细菌及工程菌群的活体材料。E-mail:rt.zhu@siat.ac.cn钟超 (1979—),男,博士,研究员。研究方向为利用合成生物学技术发展新材料,包括活体功能材料和蛋白水下黏合材料。E-mail:chao.zhong@siat.ac.cn戴卓君 (1987—),女,博士,副研究员。研究方向是编辑合成功能菌群并结合高分子化学与物理的手段实现生物制剂及活体材料的智能制造。E-mail:zj.dai@siat.ac.cn -
基金资助:国家重点研发计划(2018YFA0903000);国家自然科学基金(32071427);深圳市科技计划(KQTD20180413181837372)
CLC Number:
Cite this article
ZHU Runtao, ZHONG Chao, DAI Zhuojun. Biofilm matrixes-from soft matters to engineered materials[J]. Synthetic Biology Journal, 2022, 3(4): 626-637.
朱润涛, 钟超, 戴卓君. 细菌生物被膜的软物质特性及其工程化应用[J]. 合成生物学, 2022, 3(4): 626-637.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2021-087
| 材料 | 弹性 /GPa | 黏度 /mPa·s | 参考文献 |
|---|---|---|---|
| 钛 | 106~108 | [ | |
| 铝 | 68~70 | [ | |
| 透明质酸基地的组织工程化骨架 | 10-4 | 107 | [ |
| 皮肤 | 0.015~0.15 | [ | |
| 人皮质骨 | 15~30 | [ | |
| 牙釉质 | 80 | [ | |
| 毛发 | 7 | [ | |
| 水 | — | 1 | |
| 唾液 | — | 1.3~2.0 | [ |
| 血液 | 3~4 | [ | |
| 尿液 | 0.8 | [ | |
| Pseudomonas生物外膜(剪切模式) | 10-10 | [ | |
| Pseudomonas全生物膜(剪切模式) | 10-5 | [ | |
| Miscellaneous生物膜(剪切模式) | 10-10~10-4 | 103~1013 | [ |
| 环境与工业的生物膜(拉伸模式) | 10-8 | [ | |
| 口腔生物膜(压缩模式) | 10-8~10-7 | [ |
Tab. 1 Viscoelasticity of different biological and synthetic materials at room temperature
| 材料 | 弹性 /GPa | 黏度 /mPa·s | 参考文献 |
|---|---|---|---|
| 钛 | 106~108 | [ | |
| 铝 | 68~70 | [ | |
| 透明质酸基地的组织工程化骨架 | 10-4 | 107 | [ |
| 皮肤 | 0.015~0.15 | [ | |
| 人皮质骨 | 15~30 | [ | |
| 牙釉质 | 80 | [ | |
| 毛发 | 7 | [ | |
| 水 | — | 1 | |
| 唾液 | — | 1.3~2.0 | [ |
| 血液 | 3~4 | [ | |
| 尿液 | 0.8 | [ | |
| Pseudomonas生物外膜(剪切模式) | 10-10 | [ | |
| Pseudomonas全生物膜(剪切模式) | 10-5 | [ | |
| Miscellaneous生物膜(剪切模式) | 10-10~10-4 | 103~1013 | [ |
| 环境与工业的生物膜(拉伸模式) | 10-8 | [ | |
| 口腔生物膜(压缩模式) | 10-8~10-7 | [ |
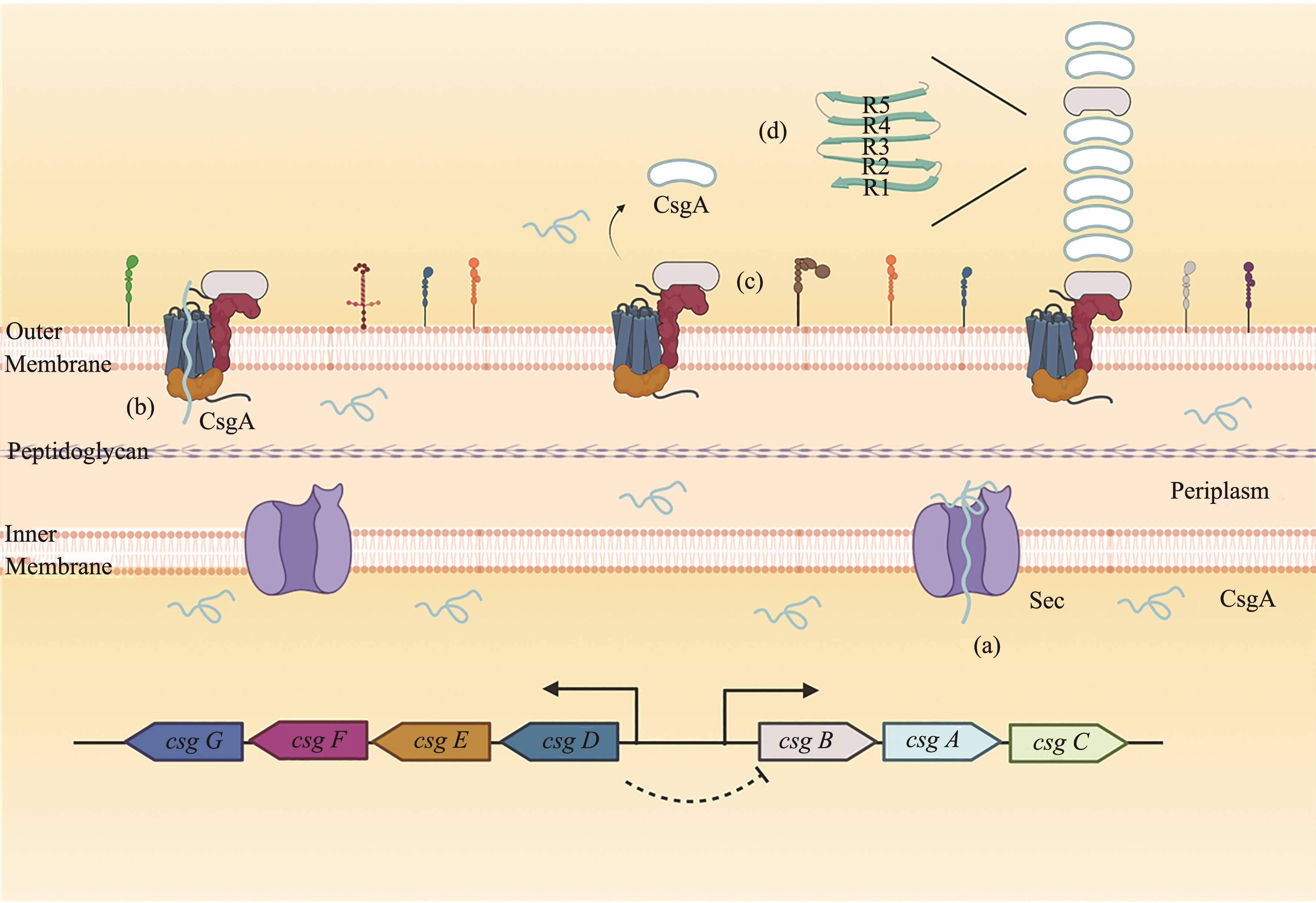
Fig. 1 The molecular mechanism for curli formation[An unfolded CsgA monomer enters the periplasm via the Sec translocon, and CsgB-C and CsgE-F are transported cross the inner membrane(a); A subunit CsgA encapsulated by a chamber of the CsgG: CsgE complex is secreted over outer membrane, which is driven by entropy increase(b); CsgB nucleated polymerization of a soluble subunit CsgA can assemble into a curli system(c); As the major subunit of the curli fiber, the mature CsgA protein is with a β-sheet-turn-β-sheet conformation (d)]
| 功能单位 | 类型 | 参考文献 |
|---|---|---|
| His Tag | 标签 | [ |
| 贻贝足蛋白 | 防水黏合剂 | [ |
| HA | 标签 | [ |
| Flag | 标签 | [ |
| 镧系元素结合标签(LBTs) | 金属结合多肽 | [ |
| A3 | 金属结合多肽 | [ |
| 流感病毒结合肽 | 结合病毒衣壳 | [ |
| 羟基磷灰石结合肽 | 矿化 | [ |
| DNA结合结构域 | 结合DNA | [ |
| 脂酶结合肽 | 结合脂酶 | [ |
| SpyTag | 结合SpyCatcher | [ |
| 金属结合域 | 结合不锈钢 | [ |
| 材料结合多肽 | 合成纳米材料 | [ |
| 几丁质结合域 | 结合几丁质 | [ |
| Mms | 结合磁颗粒 | [ |
| 4-叠氮基-L-苯丙氨酸 | 非天然氨基酸 | [ |
| 人肠三叶因子 | 治疗结肠炎 | [ |
Tab. 2 Domains-fused CsgA functionalizes curli
| 功能单位 | 类型 | 参考文献 |
|---|---|---|
| His Tag | 标签 | [ |
| 贻贝足蛋白 | 防水黏合剂 | [ |
| HA | 标签 | [ |
| Flag | 标签 | [ |
| 镧系元素结合标签(LBTs) | 金属结合多肽 | [ |
| A3 | 金属结合多肽 | [ |
| 流感病毒结合肽 | 结合病毒衣壳 | [ |
| 羟基磷灰石结合肽 | 矿化 | [ |
| DNA结合结构域 | 结合DNA | [ |
| 脂酶结合肽 | 结合脂酶 | [ |
| SpyTag | 结合SpyCatcher | [ |
| 金属结合域 | 结合不锈钢 | [ |
| 材料结合多肽 | 合成纳米材料 | [ |
| 几丁质结合域 | 结合几丁质 | [ |
| Mms | 结合磁颗粒 | [ |
| 4-叠氮基-L-苯丙氨酸 | 非天然氨基酸 | [ |
| 人肠三叶因子 | 治疗结肠炎 | [ |
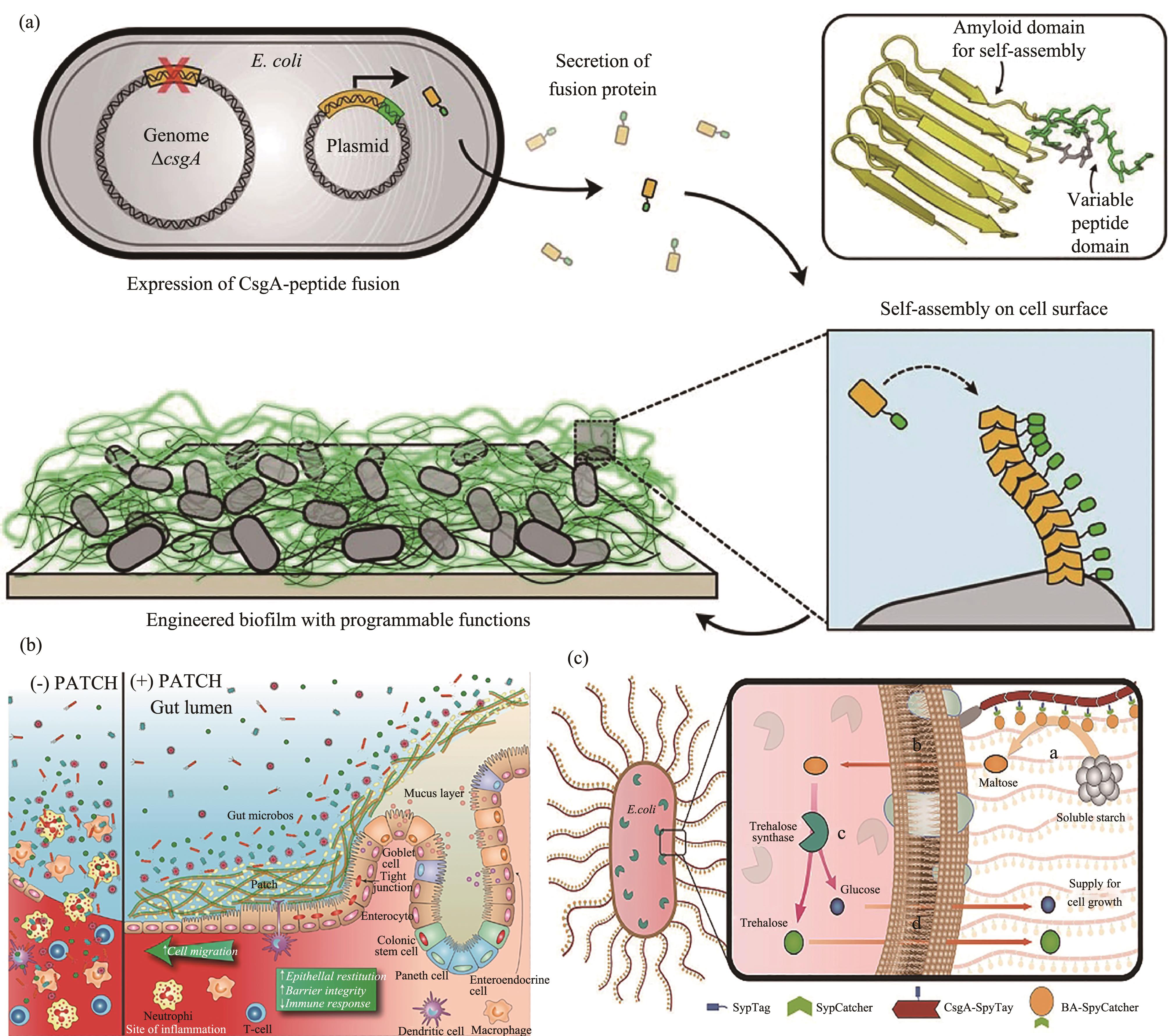
Fig. 2 Functionalization of curli via fusion with CsgA(a) Gene circuit containing inducible expression of CsgA (with subunits engineered to display various peptide tags) was transformed into a host strain with the endogenous csgA gene deleted;(b) Fusing CsgA with trefoil factors (TFFs) led to the formation of curli nanofibers displaying TTFs. The resultant material was proven to promote intestinal barrier function and epithelial restitution;(c) SpyTag displaying curli was fused with SpyCatcher decorated β-amylase. β-amylase converted the starch into maltose. The maltose was then transported intracellularly and further catalyzed into trehalose through the intracellularly expressed trehalase
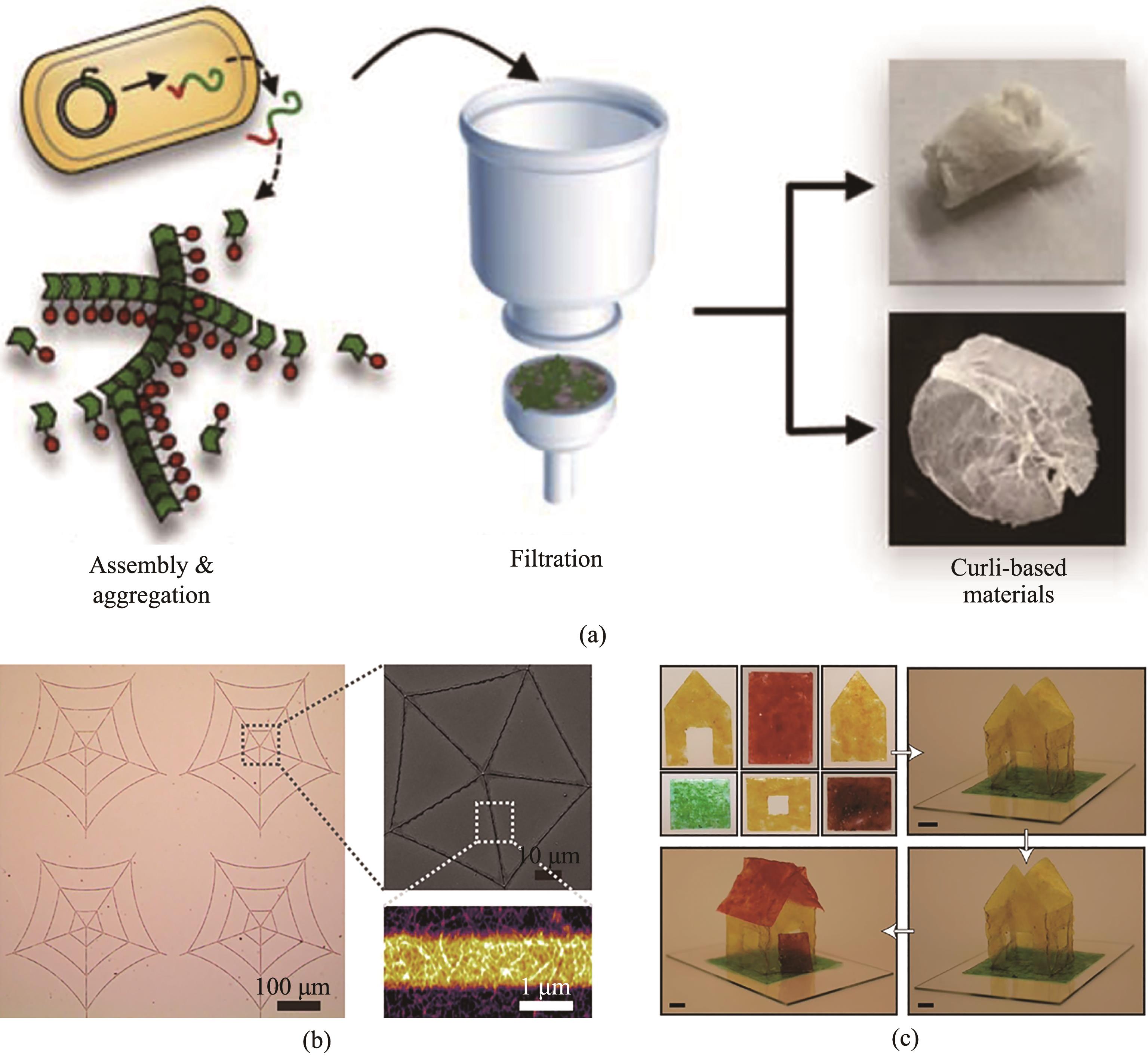
Fig. 3 Purified curli as the materials precursors(a) Curli fiber produced by E. coli were purified using a fast and easily accessible vacuum filtration procedure. The fibers were then disassembled, reassembled into thin films, and recycled for further materials processing[81];(b) Generation of diverse patterns with a generic amyloid monomer inks (consisting of genetically engineered biofilm proteins dissolved in hexafluoroisopropanol), along with methanol-assisted curing[83];(c) Aqua plastic was produced by casting and drying purified curli under ambient conditions[82]. The resultant aqua plastic could withstand strong acid/base and organic solvents. In addition, aqua plastic could be healed and welded to form three-dimensional architectures using water
| 1 | DONLAN R M. Biofilms: microbial life on surfaces[J]. Emerging Infectious Diseases, 2002, 8(9): 881-890. |
| 2 | WATNICK P, KOLTER R. Biofilm, city of microbes[J]. Journal of Bacteriology, 2000, 182(10): 2675-2679. |
| 3 | HALL-STOODLEY L, COSTERTON J W, STOODLEY P. Bacterial biofilms: from the natural environment to infectious diseases[J]. Nature Reviews Microbiology, 2004, 2(2): 95-108. |
| 4 | WILKING J N, ANGELINI T E, SEMINARA A, et al. Biofilms as complex fluids[J]. MRS Bulletin, 2011, 36(5): 385-391. |
| 5 | KLAPPER I, RUPP C J, CARGO R, et al. Viscoelastic fluid description of bacterial biofilm material properties[J]. Biotechnology and Bioengineering, 2002, 80(3): 289-296. |
| 6 | FRANCIUS G, DOMENECH O, MINGEOT-LECLERCQ M P, et al. Direct observation of Staphylococcus aureus cell wall digestion by lysostaphin[J]. Journal of Bacteriology, 2008, 190(24): 7904-7909. |
| 7 | ARNOLDI M, FRITZ M, BÄUERLEIN E, et al. Bacterial turgor pressure can be measured by atomic force microscopy[J]. Physical Review E, Statistical Physics, Plasmas, Fluids, and Related Interdisciplinary Topics, 2000, 62(1 Pt B): 1034-1044. |
| 8 | PETERSON B W, HE Y, REN Y J, et al. Viscoelasticity of biofilms and their recalcitrance to mechanical and chemical challenges[J]. FEMS Microbiology Reviews, 2015, 39(2): 234-245. |
| 9 | BOYER R, WELSCH G, COLLINGS E. Materials properties handbook: Titanium alloys[M]. ASM International, 1994. |
| 10 | COMMITTEE A H. Properties and selection: nonferrous alloys and special-purpose materials[M]. ASM International, 1990. |
| 11 | BORZACCHIELLO A, MAYOL L, RAMIRES P A, et al. Structural and rheological characterization of hyaluronic acid-based scaffolds for adipose tissue engineering[J]. Biomaterials, 2007, 28(30): 4399-4408. |
| 12 | EDWARDS C, MARKS R. Evaluation of biomechanical properties of human skin[J]. Clinics in Dermatology, 1995, 13(4): 375-380. |
| 13 | RATNER B D, HOFFMAN A S, SCHOEN F J, et al. Biomaterials science: an evolving, multidisciplinary endeavor[M]// Biomaterials Science. 3rd . Amsterdam: Elsevier, 2013: xxv-xxxix. |
| 14 | MURCKO A C. Patient-centered interviewing: An evidence-based method. 2nd ed[J]. Clinical Nurse Specialist, 2002, 16(6): 326. |
| 15 | LEE J, KWON H J. Measurement of stress-strain behaviour of human hair fibres using optical techniques[J]. International Journal of Cosmetic Science, 2013, 35(3): 238-243. |
| 16 | RANTONEN P J, MEURMAN J H. Viscosity of whole saliva[J]. Acta Odontologica Scandinavica, 1998, 56(4): 210-214. |
| 17 | ELERT G. Viscosity[EB/OL]// The Physics Hypertextbook, 2014. . |
| 18 | INMAN B A, ETIENNE W, RUBIN R, et al. The impact of temperature and urinary constituents on urine viscosity and its relevance to bladder hyperthermia treatment[J]. International Journal of Hyperthermia, 2013, 29(3): 206-210. |
| 19 | WANG H W, DENG H H, MA L M, et al. Influence of operating conditions on extracellular polymeric substances and surface properties of sludge flocs[J]. Carbohydrate Polymers, 2013, 92(1): 510-515. |
| 20 | KÖRSTGENS V, FLEMMING H C, WINGENDER J, et al. Uniaxial compression measurement device for investigation of the mechanical stability of biofilms[J]. Journal of Microbiological Methods, 2001, 46(1): 9-17. |
| 21 | SHAW T, WINSTON M, RUPP C J, et al. Commonality of elastic relaxation times in biofilms[J]. Physical Review Letters, 2004, 93(9): 098102. |
| 22 | STOODLEY P, LEWANDOWSKI Z, BOYLE J D, et al. Structural deformation of bacterial biofilms caused by short-term fluctuations in fluid shear: an in situ investigation of biofilm rheology[J]. Biotechnology and Bioengineering, 1999, 65(1): 83-92. |
| 23 | PARAMONOVA E, KALMYKOWA O J, VAN DER MEI H C, et al. Impact of hydrodynamics on oral biofilm strength[J]. Journal of Dental Research, 2009, 88(10): 922-926. |
| 24 | CHEN D T N, WEN Q, JANMEY P A, et al. Rheology of soft materials[J]. Annual Review of Condensed Matter Physics, 2010, 1: 301-322. |
| 25 | TERAOKA I. Polymer solution : an introduction to physical properties[M]. New York: John Wiley & Sons, Inc, 2002. |
| 26 | HUNG C, ZHOU Y Z, PINKNER J S, et al. Escherichia coli biofilms have an organized and complex extracellular matrix structure[J]. mBio, 2013, 4(5): e00645-e00613. |
| 27 | GREENBERG E P. Bacterial communication and group behavior[J]. The Journal of Clinical Investigation, 2003, 112(9): 1288-1290. |
| 28 | ZOGAJ X, BOKRANZ W, NIMTZ M, et al. Production of cellulose and curli fimbriae by members of the family Enterobacteriaceae isolated from the human gastrointestinal tract[J]. Infection and Immunity, 2003, 71(7): 4151-4158. |
| 29 | TURSI S A, TÜKEL Ç. Curli-containing enteric biofilms inside and out: matrix composition, immune recognition, and disease implications[J]. Microbiology and Molecular Biology Reviews: MMBR, 2018, 82(4): e00028-e00018. |
| 30 | EVANS M L, CHORELL E, TAYLOR J D, et al. The bacterial curli system possesses a potent and selective inhibitor of amyloid formation[J]. Molecular Cell, 2015, 57(3): 445-455. |
| 31 | CHAPMAN M R, ROBINSON L S, PINKNER J S, et al. Role of Escherichia coli curli operons in directing amyloid fiber formation[J]. Science, 2002, 295(5556): 851-855. |
| 32 | NENNINGER A A, ROBINSON L S, HAMMER N D, et al. CsgE is a curli secretion specificity factor that prevents amyloid fibre aggregation[J]. Molecular Microbiology, 2011, 81(2): 486-499. |
| 33 | HAMMER N D, MCGUFFIE B A, ZHOU Y Z, et al. The C-terminal repeating units of CsgB direct bacterial functional amyloid nucleation[J]. Journal of Molecular Biology, 2012, 422(3): 376-389. |
| 34 | SUNDE M, SERPELL L C, BARTLAM M, et al. Common core structure of amyloid fibrils by synchrotron X-ray diffraction[J]. Journal of Molecular Biology, 1997, 273(3): 729-739. |
| 35 | ZHONG C, GURRY T, CHENG A A, et al. Strong underwater adhesives made by self-assembling multi-protein nanofibres[J]. Nature Nanotechnology, 2014, 9: 858-866. |
| 36 | CHAPMAN M R, ROBINSON L S, PINKNER J S, et al. Role of Escherichia coli curli operons in directing amyloid fiber formation[J]. Science, 2002, 295(5556): 851-855. |
| 37 | HAMMAR M, BIAN Z, NORMARK S. Nucleator-dependent intercellular assembly of adhesive curli organelles in Escherichia coli [J]. Proceedings of the National Academy of Sciences of the United States of America, 1996, 93(13): 6562-6566. |
| 38 | HAMMER N D, SCHMIDT J C, CHAPMAN M R. The curli nucleator protein, CsgB, contains an amyloidogenic domain that directs CsgA polymerization[J]. Proceedings of the National Academy of Sciences of the United States of America, 2007, 104(30): 12494-12499. |
| 39 | EVANS M L, CHAPMAN M R. Curli biogenesis: order out of disorder[J]. Biochimica et Biophysica Acta, 2014, 1843(8): 1551-1558. |
| 40 | ISHIHAMA A. Prokaryotic genome regulation: multifactor promoters, multitarget regulators and hierarchic networks[J]. FEMS Microbiology Reviews, 2010, 34(5): 628-645. |
| 41 | HAMMAR M, ARNQVIST A, BIAN Z, et al. Expression of two csg operons is required for production of fibronectin- and Congo red-binding curli polymers in Escherichia coli K-12[J]. Molecular Microbiology, 1995, 18(4): 661-670. |
| 42 | ZAKIKHANY K, HARRINGTON C R, NIMTZ M, et al. Unphosphorylated CsgD controls biofilm formation in Salmonella enterica serovar Typhimurium[J]. Molecular Microbiology, 2010, 77(3): 771-786. |
| 43 | NENNINGER A A, ROBINSON L S, HULTGREN S J. Localized and efficient curli nucleation requires the chaperone-like amyloid assembly protein CsgF[J]. Proceedings of the National Academy of Sciences of the United States of America, 2009, 106(3): 900-905. |
| 44 | GOYAL P, KRASTEVA P V, VAN GERVEN N, et al. Structural and mechanistic insights into the bacterial amyloid secretion channel CsgG[J]. Nature, 2014, 516(7530): 250-253. |
| 45 | ROBINSON L S, ASHMAN E M, HULTGREN S J, et al. Secretion of curli fibre subunits is mediated by the outer membrane-localized CsgG protein[J]. Molecular Microbiology, 2006, 59(3): 870-881. |
| 46 | LOFERER H, HAMMAR M, NORMARK S. Availability of the fibre subunit CsgA and the nucleator protein CsgB during assembly of fibronectin-binding curli is limited by the intracellular concentration of the novel lipoprotein CsgG[J]. Molecular Microbiology, 1997, 26(1): 11-23. |
| 47 | JIANG J S, PENTELUTE B L, COLLIER R J, et al. Atomic structure of anthrax protective antigen pore elucidates toxin translocation[J]. Nature, 2015, 521(7553): 545-549. |
| 48 | YAN Z F, YIN M, CHEN J N, et al. Assembly and substrate recognition of curli biogenesis system[J]. Nature Communications, 2020, 11: 241. |
| 49 | GARDNER T S, CANTOR C R, COLLINS J J. Construction of a genetic toggle switch in Escherichia coli [J]. Nature, 2000, 403(6767): 339-342. |
| 50 | ELOWITZ M B, LEIBLER S. A synthetic oscillatory network of transcriptional regulators[J]. Nature, 2000, 403(6767): 335-338. |
| 51 | WU F L, BETHKE J H, WANG M D, et al. Quantitative and synthetic biology approaches to combat bacterial pathogens[J]. Current Opinion in Biomedical Engineering, 2017, 4: 116-126. |
| 52 | AUSLÄNDER S, WIELAND M, FUSSENEGGER M. Smart medication through combination of synthetic biology and cell microencapsulation[J]. Metabolic Engineering, 2012, 14(3): 252-260. |
| 53 | YE H F, DAOUD-EL BABA M, PENG R W, et al. A synthetic optogenetic transcription device enhances blood-glucose homeostasis in mice[J]. Science, 2011, 332(6037): 1565-1568. |
| 54 | SEGALL-SHAPIRO T H, SONTAG E D, VOIGT C A. Engineered promoters enable constant gene expression at any copy number in bacteria[J]. Nature Biotechnology, 2018, 36(4): 352-358. |
| 55 | KYLILIS N, TUZA Z A, STAN G B, et al. Tools for engineering coordinated system behaviour in synthetic microbial consortia[J]. Nature Communications, 2018, 9(1): 2677. |
| 56 | VILABOA N, FENNA M, MUNSON J, et al. Novel gene switches for targeted and timed expression of proteins of interest[J]. Molecular Therapy, 2005, 12(2): 290-298. |
| 57 | GAO C, HOU J S, XU P, et al. Programmable biomolecular switches for rewiring flux in Escherichia coli [J]. Nature Communications, 2019, 10(1): 3751. |
| 58 | NOVÁK B, TYSON J J. Design principles of biochemical oscillators[J]. Nature Reviews Molecular Cell Biology, 2008, 9(12): 981-991. |
| 59 | POTVIN-TROTTIER L, LORD N D, VINNICOMBE G, et al. Synchronous long-term oscillations in a synthetic gene circuit[J]. Nature, 2016, 538(7626): 514-517. |
| 60 | CHEN A Y, DENG Z T, BILLINGS A N, et al. Synthesis and patterning of tunable multiscale materials with engineered cells[J]. Nature Materials, 2014, 13(5): 515-523. |
| 61 | MOSER F, THAM E, GONZÁLEZ L M, et al. Light-controlled, high-resolution patterning of living engineered bacteria onto textiles, ceramics, and plastic[J]. Advanced Functional Materials, 2019, 29(30): 1901788. |
| 62 | WANG X Y, PU J H, LIU Y, et al. Immobilization of functional nano-objects in living engineered bacterial biofilms for catalytic applications[J]. National Science Review, 2019, 6(5): 929-943. |
| 63 | CAO Y, FENG Y Y, RYSER M D, et al. Programmable assembly of pressure sensors using pattern-forming bacteria[J]. Nature Biotechnology, 2017, 35(11): 1087-1093. |
| 64 | LI Y F, LI K, WANG X Y, et al. Conformable self-assembling amyloid protein coatings with genetically programmable functionality[J]. Science Advances, 2020, 6(21): eaba1425. |
| 65 | CUI M K, QI Q, GURRY T, et al. Modular genetic design of multi-domain functional amyloids: insights into self-assembly and functional properties[J]. Chemical Science, 2019, 10(14): 4004-4014. |
| 66 | WANG Y Y, AN B L, XUE B, et al. Living materials fabricated via gradient mineralization of light-inducible biofilms[J]. Nature Chemical Biology, 2021, 17(3): 351-359. |
| 67 | AN B L, WANG Y Y, JIANG X Y, et al. Programming living glue systems to perform autonomous mechanical repairs[J]. Matter, 2020, 3(6): 2080-2092. |
| 68 | NGUYEN P Q, BOTYANSZKI Z, TAY P K R, et al. Programmable biofilm-based materials from engineered curli nanofibres[J]. Nature Communications, 2014, 5: 4945. |
| 69 | TAY P K R, MANJULA-BASAVANNA A, JOSHI N S. Repurposing bacterial extracellular matrix for selective and differential abstraction of rare earth elements[J]. Green Chemistry, 2018, 20(15): 3512-3520. |
| 70 | PU J H, LIU Y, ZHANG J C, et al. Virus disinfection: virus disinfection from environmental water sources using living engineered biofilm materials[J]. Advanced Science, 2020, 7(14): 1903558. |
| 71 | ABDALI Z, AMINZARE M, ZHU X D, et al. Curli-mediated self-assembly of a fibrous protein scaffold for hydroxyapatite mineralization[J]. ACS Synthetic Biology, 2020, 9(12): 3334-3343. |
| 72 | DONG H, ZHANG W X, XUAN Q Z, et al. Binding peptide-guided immobilization of lipases with significantly improved catalytic performance using Escherichia coli BL21(DE3) biofilms as a platform[J]. ACS Applied Materials & Interfaces, 2021, 13(5): 6168-6179. |
| 73 | BOTYANSZKI Z, TAY P K R, NGUYEN P Q, et al. Engineered catalytic biofilms: site-specific enzyme immobilization onto E. coli curli nanofibers[J]. Biotechnology and Bioengineering, 2015, 112(10): 2016-2024. |
| 74 | BAO J J, LIU N, ZHU L Y, et al. Programming a biofilm-mediated multienzyme-assembly-cascade system for the biocatalytic production of glucosamine from chitin[J]. Journal of Agricultural and Food Chemistry, 2018, 66(30): 8061-8068. |
| 75 | OLMEZ T T, SAHIN KEHRIBAR E, ISILAK M E, et al. Synthetic genetic circuits for self-actuated cellular nanomaterial fabrication devices[J]. ACS Synthetic Biology, 2019, 8(9): 2152-2162. |
| 76 | PRAVESCHOTINUNT P, DORVAL COURCHESNE N M, DEN HARTOG I, et al. Tracking of engineered bacteria in vivo using nonstandard amino acid incorporation[J]. ACS Synthetic Biology, 2018, 7(6): 1640-1650. |
| 77 | PRAVESCHOTINUNT P, DURAJ-THATTE A M, GELFAT I, et al. Engineered E. coli Nissle 1917 for the delivery of matrix-tethered therapeutic domains to the gut[J]. Nature Communications, 2019, 10: 5580. |
| 78 | DURAJ-THATTE A M, PRAVESCHOTINUNT P, NASH T R, et al. Modulating bacterial and gut mucosal interactions with engineered biofilm matrix proteins[J]. Scientific Reports, 2018, 8: 3475. |
| 79 | AXPE E, DURAJ-THATTE A, CHANG Y, et al. Fabrication of amyloid curli fibers-alginate nanocomposite hydrogels with enhanced stiffness[J]. ACS Biomaterials Science & Engineering, 2018, 4(6): 2100-2105. |
| 80 | JIANG L, SONG X G, LI Y F, et al. Programming integrative extracellular and intracellular biocatalysis for rapid, robust, and recyclable synthesis of trehalose[J]. ACS Catalysis, 2018, 8(3): 1837-1842. |
| 81 | DORVAL COURCHESNE N M, DURAJ-THATTE A, TAY P K R, et al. Scalable production of genetically engineered nanofibrous macroscopic materials via filtration[J]. ACS Biomaterials Science & Engineering, 2017, 3(5): 733-741. |
| 82 | DURAJ-THATTE A M, MANJULA-BASAVANNA A, COURCHESNE N M D, et al. Water-processable, biodegradable and coatable aquaplastic from engineered biofilms[J]. Nature Chemical Biology, 2021, 17(6): 732-738. |
| 83 | LI Y F, LI K, WANG X Y, et al. Patterned amyloid materials integrating robustness and genetically programmable functionality[J]. Nano Letters, 2019, 19(12): 8399-8408. |
| 84 | GALATSIS K, WANG K L, OZKAN M, et al. Patterning and templating for nanoelectronics[J]. Advanced Materials, 2010, 22(6): 769-778. |
| 85 | CAO Y, RYSER M D, PAYNE S, et al. Collective space-sensing coordinates pattern scaling in engineered bacteria[J]. Cell, 2016, 165(3): 620-630. |
| 86 | FERNANDEZ-RODRIGUEZ J, MOSER F, SONG M, et al. Engineering RGB color vision into Escherichia coli [J]. Nature Chemical Biology, 2017, 13(7): 706-708. |
| 87 | NGUYEN P Q, COURCHESNE N M D, DURAJ-THATTE A, et al. Engineered living materials: prospects and challenges for using biological systems to direct the assembly of smart materials[J]. Advanced Materials, 2018, 30(19): e1704847. |
| 88 | GILBERT C, TANG T C, OTT W, et al. Living materials with programmable functionalities grown from engineered microbial co-cultures[J]. Nature Materials, 2021, 20(5): 691-700. |
| 89 | LIU X Y, YUK H, LIN S T, et al. 3D printing of living responsive materials and devices[J]. Advanced Materials, 2018, 30(4): 1704821. |
| 90 | GILBERT C, ELLIS T. Biological engineered living materials: Growing functional materials with genetically programmable properties[J]. ACS Synthetic Biology, 2019, 8(1): 1-15. |
| 91 | CHEN A Y, ZHONG C, LU T K. Engineering living functional materials[J]. ACS Synthetic Biology, 2015, 4(1): 8-11. |
| 92 | YANG P D. Liquid sunlight: the evolution of photosynthetic biohybrids[J]. Nano Letters, 2021, 21(13): 5453-5456. |
| 93 | LE FEUVRE R A, SCRUTTON N S. A living foundry for synthetic biological materials: a synthetic biology roadmap to new advanced materials[J]. Synthetic and Systems Biotechnology, 2018, 3(2): 105-112. |
| 94 | TANG T C, AN B L, HUANG Y Y, et al. Materials design by synthetic biology[J]. Nature Reviews Materials, 2021, 6(4): 332-350. |
| 95 | DAI Z J, LEE A J, ROBERTS S, et al. Versatile biomanufacturing through stimulus-responsive cell-material feedback[J]. Nature Chemical Biology, 2019, 15(10): 1017-1024. |
| 96 | DAI Z J, YANG X Y, WU F L, et al. Living fabrication of functional semi-interpenetrating polymeric materials[J]. Nature Communications, 2021, 12: 3422. |
| 97 | MOLINARI S, TESORIERO R F JR, AJO-FRANKLIN C M. Bottom-up approaches to engineered living materials: Challenges and future directions[J]. Matter, 2021, 4(10): 3095-3120. |
| [1] | GAO Ge, BIAN Qi, WANG Baojun. Synthetic genetic circuit engineering: principles, advances and prospects [J]. Synthetic Biology Journal, 2025, 6(1): 45-64. |
| [2] | LI Jiyuan, WU Guosheng. Two hypothesises for the origins of organisms from the synthetic biology perspective [J]. Synthetic Biology Journal, 2025, 6(1): 190-202. |
| [3] | JIAO Hongtao, QI Meng, SHAO Bin, JIANG Jinsong. Legal issues for the storage of DNA data [J]. Synthetic Biology Journal, 2025, 6(1): 177-189. |
| [4] | TANG Xinghua, LU Qianneng, HU Yilin. Philosophical reflections on synthetic biology in the Anthropocene [J]. Synthetic Biology Journal, 2025, 6(1): 203-212. |
| [5] | XU Huaisheng, SHI Xiaolong, LIU Xiaoguang, XU Miaomiao. Key technologies for DNA storage: encoding, error correction, random access, and security [J]. Synthetic Biology Journal, 2025, 6(1): 157-176. |
| [6] | SHI Ting, SONG Zhan, SONG Shiyi, ZHANG Yi-Heng P. Job. In vitro BioTransformation (ivBT): a new frontier of industrial biomanufacturing [J]. Synthetic Biology Journal, 2024, 5(6): 1437-1460. |
| [7] | CHAI Meng, WANG Fengqing, WEI Dongzhi. Synthesis of organic acids from lignocellulose by biotransformation [J]. Synthetic Biology Journal, 2024, 5(6): 1242-1263. |
| [8] | SHAO Mingwei, SUN Simian, YANG Shimao, CHEN Guoqiang. Bioproduction based on extremophiles [J]. Synthetic Biology Journal, 2024, 5(6): 1419-1436. |
| [9] | CHEN Yu, ZHANG Kang, QIU Yijing, CHENG Caiyun, YIN Jingjing, SONG Tianshun, XIE Jingjing. Progress of microbial electrosynthesis for conversion of CO2 [J]. Synthetic Biology Journal, 2024, 5(5): 1142-1168. |
| [10] | ZHENG Haotian, LI Chaofeng, LIU Liangxu, WANG Jiawei, LI Hengrun, NI Jun. Design, optimization and application of synthetic carbon-negative phototrophic community [J]. Synthetic Biology Journal, 2024, 5(5): 1189-1210. |
| [11] | CHEN Ziling, XIANG Yangfei. Integrated development of organoid technology and synthetic biology [J]. Synthetic Biology Journal, 2024, 5(4): 795-812. |
| [12] | CAI Bingyu, TAN Xiangtian, LI Wei. Advances in synthetic biology for engineering stem cell [J]. Synthetic Biology Journal, 2024, 5(4): 782-794. |
| [13] | XIE Huang, ZHENG Yilei, SU Yiting, RUAN Jingyi, LI Yongquan. An overview on reconstructing the biosynthetic system of actinomycetes for polyketides production [J]. Synthetic Biology Journal, 2024, 5(3): 612-630. |
| [14] | ZHA Wenlong, BU Lan, ZI Jiachen. Advances in synthetic biology for producing potent pharmaceutical ingredients of traditional Chinese medicine [J]. Synthetic Biology Journal, 2024, 5(3): 631-657. |
| [15] | HUI Zhen, TANG Xiaoyu. Applications of the CRISPR/Cas9 editing system in the study of microbial natural products [J]. Synthetic Biology Journal, 2024, 5(3): 658-671. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||