Synthetic biology empowers breakthroughs in addressing immunotherapy limitations
WANG Ziheng1,2, LIU Ziyi2, MA Yuqian1,2, QIN Hongyan2, ZHAO Junlong2, ZHANG Xiangqian1
- 1.School of Life Sciences,Yan’an University,Yan’an 716000,Shannxi,China
2.Department of Medical Genetics and Developmental Biology,School of Basic Medical Sciences,Fourth Military Medical University,Xi’an 710000,Shannxi,China
-
Received:2025-07-21Revised:2025-10-28Published:2025-11-07 -
Contact:ZHANG Xiangqian
合成生物学助力突破免疫治疗局限性
王子恒1,2, 刘子怡2, 马毓谦1,2, 秦鸿雁2, 赵俊龙2, 张向前1
- 1.延安大学生命科学学院,陕西,延安 716000
2.第四军医大学基础医学院医学遗传与发育生物学教研室,陕西,西安 710000
-
通讯作者:张向前 -
作者简介:王子恒 (1998—),男,硕士研究生。研究方向为合成生物学和免疫细胞治疗。 E-mail:1355985130@qq.com张向前 (1973—),男,教授,硕士生导师。研究方向为植物天然产物及miRNA调控机制方面的研究和学科教学(生物)领域的教学研究。 E-mail: zxq@yau.edu.cn -
基金资助:国家自然科学基金(82373270);陕西省自然科学基金(2024SF-ZDCYL-03-28)
CLC Number:
Cite this article
WANG Ziheng, LIU Ziyi, MA Yuqian, QIN Hongyan, ZHAO Junlong, ZHANG Xiangqian. Synthetic biology empowers breakthroughs in addressing immunotherapy limitations[J]. Synthetic Biology Journal, DOI: 10.12211/2096-8280.2025-074.
王子恒, 刘子怡, 马毓谦, 秦鸿雁, 赵俊龙, 张向前. 合成生物学助力突破免疫治疗局限性[J]. 合成生物学, DOI: 10.12211/2096-8280.2025-074.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2025-074
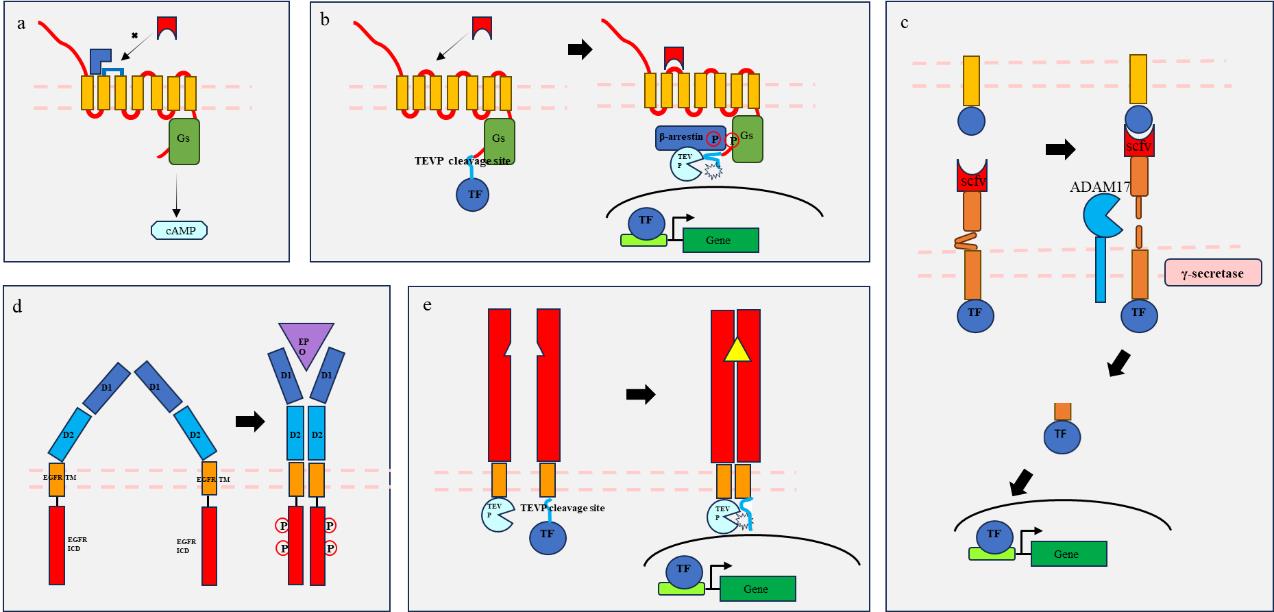
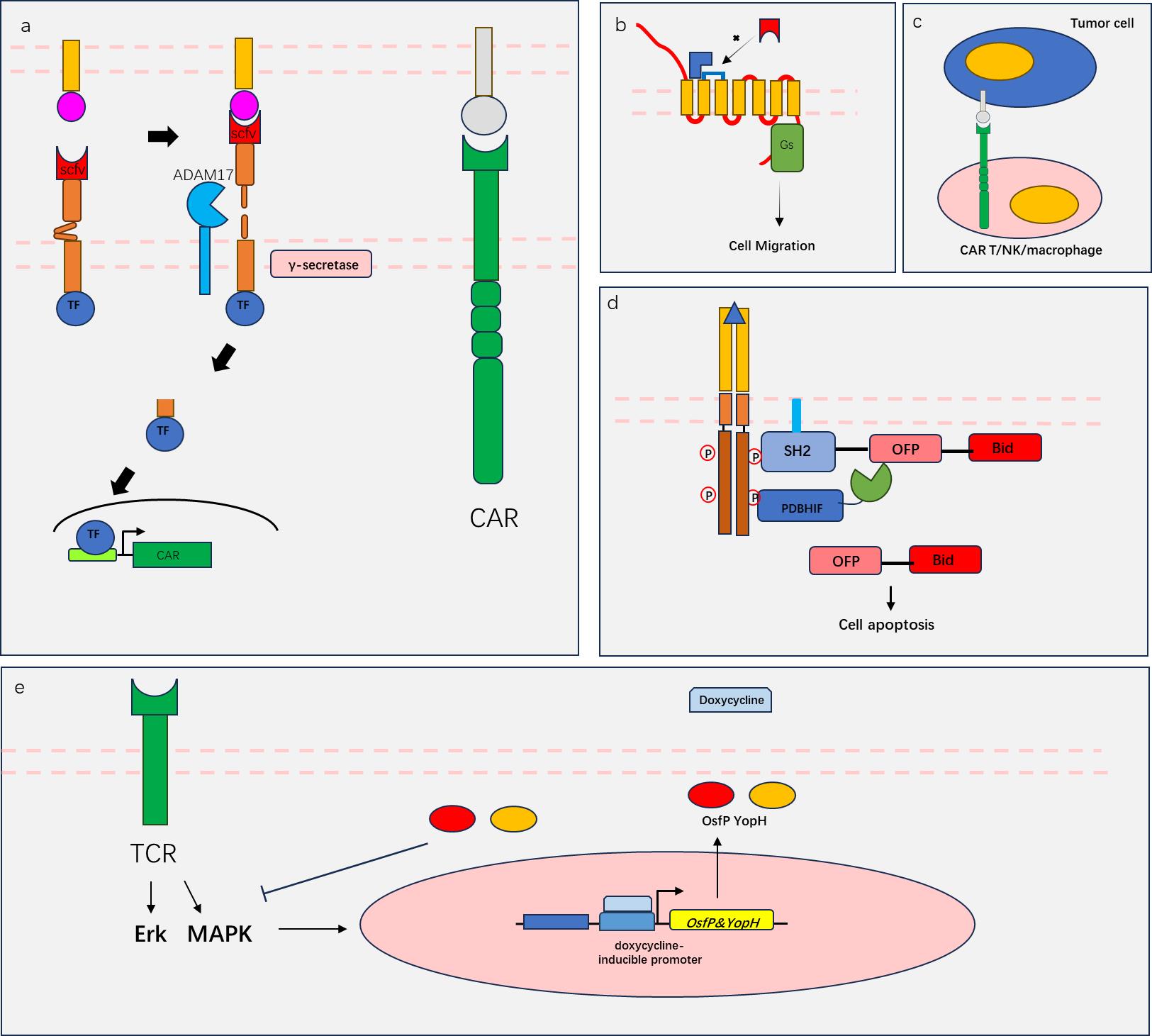
Fig. 1 Schematic diagram of engineering receptor principle(a) The RASSL system utilizes GPCR mutations to recognize only specific signals (blue) such as small molecule drugs; (b) The "Tango" system utilizes the proximity of β-arrestin and phosphorylated GPCR spatial positions to initiate protease cleavage and release of TF; (c) the "Syn-Notch" system utilizes two cleavage processes activated by notch receptors (extracellular ADAM17 cleavage and proximal γ-secretase cleavage) to achieve TF release; (d) The GEMS system utilizes the signaling characteristics of EPOR receptors to transmit editable signals (extracellular and intracellular signal remodeling); (e) The "MESA" system utilizes receptor dimerization spatial proximity to initiate protease cleavage and release of TF.
| 工程化受体 | 优点 | 缺点 |
|---|---|---|
| RASSL[ | 1.GPCR种类繁多且可接受多种配体激活 2.改造GPCR结合配体特异性高 | 1. 依赖于突变 2. 受限于GPCR的配体种类 3. 局限于GPCR自身激活的通路 |
| Tango[ | 1.GPCR种类繁多且可接受多种配体激活 2.下游基因激活可编辑 | 受限于GPCR的配体种类 |
| Syn-Notch[ | 1. 受体与配体结合特异性高 2.配体可选择范围广 3.可实现广泛性的目的基因表达 | 仅用于与细胞之间的配受体结合 |
| GEMS[ | 1. 配体可选择范围广 2.配体可选择为游离性配体 3. 天然信号通路,信号传导成功率高 4. 同源二聚体,工作效率与改造成功率高 5. 自激活概率低 | 信号传导局限于可选择的天然信号通路 |
| MESA[ | 1配体可选择范围广 2.配体可选择为游离性配体 3. 能够选择性表达目的产物 | 1. 异二聚体工作效率低 2. 多种质粒转染,细胞负担大 3. 有概率自激活 |
Table 1 Advantages and disadvantages of classical engineered receptors
| 工程化受体 | 优点 | 缺点 |
|---|---|---|
| RASSL[ | 1.GPCR种类繁多且可接受多种配体激活 2.改造GPCR结合配体特异性高 | 1. 依赖于突变 2. 受限于GPCR的配体种类 3. 局限于GPCR自身激活的通路 |
| Tango[ | 1.GPCR种类繁多且可接受多种配体激活 2.下游基因激活可编辑 | 受限于GPCR的配体种类 |
| Syn-Notch[ | 1. 受体与配体结合特异性高 2.配体可选择范围广 3.可实现广泛性的目的基因表达 | 仅用于与细胞之间的配受体结合 |
| GEMS[ | 1. 配体可选择范围广 2.配体可选择为游离性配体 3. 天然信号通路,信号传导成功率高 4. 同源二聚体,工作效率与改造成功率高 5. 自激活概率低 | 信号传导局限于可选择的天然信号通路 |
| MESA[ | 1配体可选择范围广 2.配体可选择为游离性配体 3. 能够选择性表达目的产物 | 1. 异二聚体工作效率低 2. 多种质粒转染,细胞负担大 3. 有概率自激活 |
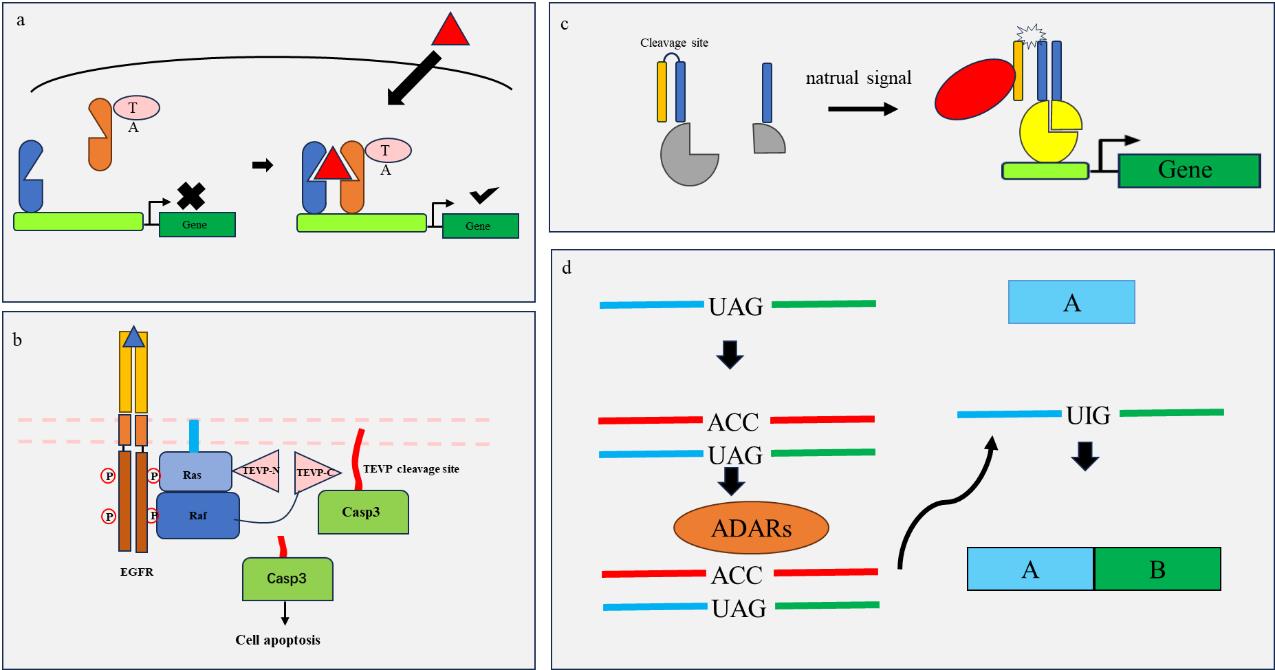
Fig. 2 Schematic diagram of engineering components principle(a) After dimerization of the "HTH" transcription factor dependent signal (red), it becomes a complete transcription factor with transcription activator (TA), initiating downstream gene expression; (b) The "CHOMP" system uses the Ras signaling pathway to activate the binding of Ras and Raf, achieving the assembly and cleavage of caspase 3 by protease TEV, causing cell apoptosis; (c) The "SPOC" logic utilizes endogenous signals to activate intracellular proteases (red) to cleave enzyme cleavage sites between useless coiled coils (orange yellow) and effector coiled coils (blue), carrying effector coiled coils (blue) to bind with each other, causing effector activation (yellow) and initiating downstream gene transcription; (d) The "DART VADAR" system uses engineered RNA strands (red) to target the target RNA (blue, green), and mismatches trigger ADARs to repair stop codons, allowing transcription to continue.
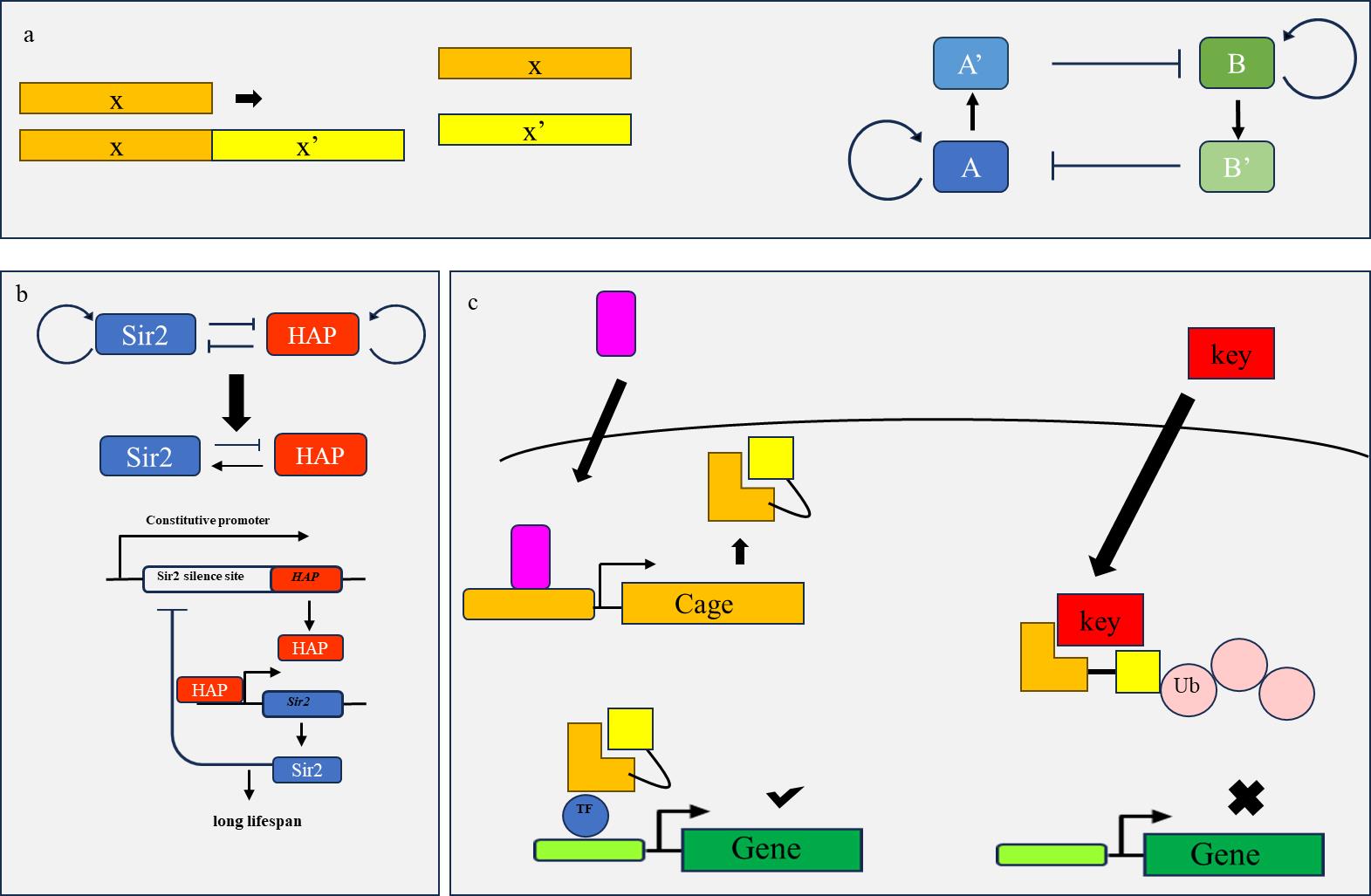
Fig. 3 Schematic diagram of engineering loop principle(a) In a bistable system, the two nucleic acid products A and B self-activate and mutually inhibit each other, achieving the transition between the two states of A and B; (b) Two mutually inhibitory genes in yeast achieve overall negative feedback regulation by downstream modification of their promoters and silencing sites, allowing their respective gene products to be expressed separately and achieving sustained cell cycle elongation; (c) The "LOCKR" system utilizes KEY competitive binding with Cage to control the degradation of Cage gene products, achieving tolerance to the signal (purple) that regulates Cage.
| CAR免疫细胞 | 优点 | 缺点 |
|---|---|---|
| CAR-T | 1.肿瘤杀伤强: T细胞具有多种肿瘤杀伤手段,能够分泌穿孔素、颗粒酶引发肿瘤细胞死亡,通过死亡受体途径(Fas/FasL)引发肿瘤细胞凋亡,通过大量细胞因子分泌增强自身,抑制及杀伤肿瘤[ 2.持久性强:T细胞 一旦激活,能够大规模扩增,并在体内形成长期免疫记忆,提供持续的监视和杀伤效果[ 3.杀伤效率高:T细胞杀伤肿瘤后并不会发生凋亡,可立即脱离寻找下一个肿瘤细胞[ 4.技术相对成熟: 是第一个成功商业化的细胞疗法,拥有大量的临床数据、生产工艺和经验[ | 1.毒副作用强:T细胞激活后细胞因子释放能力强,易造成细胞因子释放综合征[ 2.治疗实体瘤效果差: 肿瘤免疫微环境使得T细胞迁移进入肿瘤的信号不足,难以浸润实体瘤[ 3.自体来源为主: 目前多为自体疗法,导致制备成本高、时间长[ 4.可能脱靶:靶抗原在正常组织上也有低水平表达会导致T细胞攻击正常细胞[ |
| CAR-NK | 1.毒性低: NK细胞通过不同的机制杀伤靶细胞,不会引起严重的细胞因子释放[ 2.通用性高:不同于T细胞这类适应性免疫细胞,NK细胞作为固有免疫细胞 可以从健康供体的外周血、脐带血或NK细胞系扩增而来,有望成为通用型产品,成本更低[ 3.对实体瘤潜力更大: 具有更好的肿瘤组织浸润能力,且在肿瘤微环境中仍能保持一定的活性[ | 1.持久作用短: NK细胞在体内的存活时间较短,无法形成长期的免疫记忆[ 2.增殖能力有限: 在体内的增殖能力不如T细胞[ 3.技术尚不成熟: 实验数据与临床数据远少于CAR-T[ |
| CAR-M | 1.实体瘤浸润能力强: 巨噬细胞天生具有浸润到实体瘤深部的能力[ 2.具有重塑肿瘤微环境的能力: 活化的CAR-M不仅能直接杀伤肿瘤,还能分泌细胞因子(如IFN-γ)将抑制性的M2型巨噬细胞转化为杀伤性的“M1型”,并招募其他的免疫细胞(如T细胞)到肿瘤部位[ 3.吞噬作用: 能够通过强大的吞噬作用直接吞噬肿瘤细胞[ 4.通用性强:作为固有免疫细胞,可在体外大量扩增改造[ | 1.存在促瘤风险: 巨噬细胞如果功能失调,反而可能被肿瘤微环境刺激成为M2型巨噬细胞,加重肿瘤进展[ 2.肿瘤杀伤效率低:巨噬细胞随着吞噬肿瘤细胞数量增多,自身会发生凋亡,无法做到T细胞的持续杀伤[ 3.体内增殖能力差:相较于T细胞,体内活化后的巨噬细胞增殖能力差[ |
Table 2 Comparison of different CAR immune cells
| CAR免疫细胞 | 优点 | 缺点 |
|---|---|---|
| CAR-T | 1.肿瘤杀伤强: T细胞具有多种肿瘤杀伤手段,能够分泌穿孔素、颗粒酶引发肿瘤细胞死亡,通过死亡受体途径(Fas/FasL)引发肿瘤细胞凋亡,通过大量细胞因子分泌增强自身,抑制及杀伤肿瘤[ 2.持久性强:T细胞 一旦激活,能够大规模扩增,并在体内形成长期免疫记忆,提供持续的监视和杀伤效果[ 3.杀伤效率高:T细胞杀伤肿瘤后并不会发生凋亡,可立即脱离寻找下一个肿瘤细胞[ 4.技术相对成熟: 是第一个成功商业化的细胞疗法,拥有大量的临床数据、生产工艺和经验[ | 1.毒副作用强:T细胞激活后细胞因子释放能力强,易造成细胞因子释放综合征[ 2.治疗实体瘤效果差: 肿瘤免疫微环境使得T细胞迁移进入肿瘤的信号不足,难以浸润实体瘤[ 3.自体来源为主: 目前多为自体疗法,导致制备成本高、时间长[ 4.可能脱靶:靶抗原在正常组织上也有低水平表达会导致T细胞攻击正常细胞[ |
| CAR-NK | 1.毒性低: NK细胞通过不同的机制杀伤靶细胞,不会引起严重的细胞因子释放[ 2.通用性高:不同于T细胞这类适应性免疫细胞,NK细胞作为固有免疫细胞 可以从健康供体的外周血、脐带血或NK细胞系扩增而来,有望成为通用型产品,成本更低[ 3.对实体瘤潜力更大: 具有更好的肿瘤组织浸润能力,且在肿瘤微环境中仍能保持一定的活性[ | 1.持久作用短: NK细胞在体内的存活时间较短,无法形成长期的免疫记忆[ 2.增殖能力有限: 在体内的增殖能力不如T细胞[ 3.技术尚不成熟: 实验数据与临床数据远少于CAR-T[ |
| CAR-M | 1.实体瘤浸润能力强: 巨噬细胞天生具有浸润到实体瘤深部的能力[ 2.具有重塑肿瘤微环境的能力: 活化的CAR-M不仅能直接杀伤肿瘤,还能分泌细胞因子(如IFN-γ)将抑制性的M2型巨噬细胞转化为杀伤性的“M1型”,并招募其他的免疫细胞(如T细胞)到肿瘤部位[ 3.吞噬作用: 能够通过强大的吞噬作用直接吞噬肿瘤细胞[ 4.通用性强:作为固有免疫细胞,可在体外大量扩增改造[ | 1.存在促瘤风险: 巨噬细胞如果功能失调,反而可能被肿瘤微环境刺激成为M2型巨噬细胞,加重肿瘤进展[ 2.肿瘤杀伤效率低:巨噬细胞随着吞噬肿瘤细胞数量增多,自身会发生凋亡,无法做到T细胞的持续杀伤[ 3.体内增殖能力差:相较于T细胞,体内活化后的巨噬细胞增殖能力差[ |
| [1] | COUZIN-FRANKEL J. Cancer immunotherapy[J]. Science, 2013, 342(6165): 1432-1433. |
| [2] | RIEDEL S. Edward Jenner and the history of smallpox and vaccination[J]. Proceedings, 2005, 18(1): 21-25. |
| [3] | MCCARTHY E F. The toxins of William B. Coley and the treatment of bone and soft-tissue sarcomas[J]. The Iowa Orthopaedic Journal, 2006, 26: 154-158. |
| [4] | MANOHAR S, JHAVERI K D, PERAZELLA M A. Immunotherapy-related acute kidney injury[J]. Advances in Chronic Kidney Disease, 2021, 28(5): 429-437.e1. |
| [5] | KENNEDY L B, SALAMA A K S. A review of cancer immunotherapy toxicity[J]. CA: A Cancer Journal for Clinicians, 2020, 70(2): 86-104. |
| [6] | YANG H X, YAO Z R, ZHOU X X, et al. Immune-related adverse events of checkpoint inhibitors: Insights into immunological dysregulation[J]. Clinical Immunology, 2020, 213: 108377. |
| [7] | MITRA A, BARUA A, HUANG L P, et al. From bench to bedside: the history and progress of CAR T cell therapy[J]. Frontiers in Immunology, 2023, 14: 1188049. |
| [8] | TURTLE C J, HAY K A, HANAFI L A, et al. Durable molecular remissions in chronic lymphocytic leukemia treated with CD19-specific chimeric antigen receptor-modified T cells after failure of ibrutinib[J]. Journal of Clinical Oncology, 2017, 35(26): 3010-3020. |
| [9] | CAMERON D E, BASHOR C J, COLLINS J J. A brief history of synthetic biology[J]. Nature Reviews Microbiology, 2014, 12(5): 381-390. |
| [10] | YANG S, SLEIGHT S C, SAURO H M. Rationally designed bidirectional promoter improves the evolutionary stability of synthetic genetic circuits[J]. Nucleic Acids Research, 2013, 41(1): e33. |
| [11] | CHAPPELL J, WATTERS K E, TAKAHASHI M K, et al. A renaissance in RNA synthetic biology: new mechanisms, applications and tools for the future[J]. Current Opinion in Chemical Biology, 2015, 28: 47-56. |
| [12] | KHALIL A S, LU T K, BASHOR C J, et al. A synthetic biology framework for programming eukaryotic transcription functions[J]. Cell, 2012, 150(3): 647-658. |
| [13] | QUINN J Y, COX R S, ADLER A, et al. SBOL visual: a graphical language for genetic designs[J]. PLoS Biology, 2015, 13(12): e1002310. |
| [14] | MISHRA D, RIVERA P M, LIN A, et al. A load driver device for engineering modularity in biological networks[J]. Nature Biotechnology, 2014, 32(12): 1268-1275. |
| [15] | GARDNER T S, CANTOR C R, COLLINS J J. Construction of a genetic toggle switch in Escherichia coli [J]. Nature, 2000, 403(6767): 339-342. |
| [16] | LIU Y, HUANG Y X, LU R, et al. Synthetic biology applications of the yeast mating signal pathway[J]. Trends in Biotechnology, 2022, 40(5): 620-631. |
| [17] | RYU J, PARK S H. Simple synthetic protein scaffolds can create adjustable artificial MAPK circuits in yeast and mammalian cells[J]. Science Signaling, 2015, 8(383): ra66. |
| [18] | LI S Y, TANG H, LI C, et al. Synthetic biology technologies and genetically engineering strategies for enhanced cell therapeutics[J]. Stem Cell Reviews and Reports, 2023, 19(2): 309-321. |
| [19] | TRENTESAUX C, YAMADA T, KLEIN O D, et al. Harnessing synthetic biology to engineer organoids and tissues[J]. Cell Stem Cell, 2023, 30(1): 10-19. |
| [20] | SOLÉ R V, MONTAÑEZ R, DURAN-NEBREDA S. Synthetic circuit designs for earth terraformation[J]. Biology Direct, 2015, 10(1): 37. |
| [21] | SHIH R M, CHEN Y Y. Engineering principles for synthetic biology circuits in cancer immunotherapy[J]. Cancer Immunology Research, 2022, 10(1): 6-11. |
| [22] | WEBER J S, YANG J C, ATKINS M B, et al. Toxicities of immunotherapy for the practitioner[J]. Journal of Clinical Oncology, 2015, 33(18): 2092-2099. |
| [23] | THOMPSON J A. New NCCN guidelines: recognition and management of immunotherapy-related toxicity[J]. Journal of the National Comprehensive Cancer Network, 2018, 16(5S): 594-596. |
| [24] | NEELAPU S S, TUMMALA S, KEBRIAEI P, et al. Chimeric antigen receptor T-cell therapy - assessment and management of toxicities[J]. Nature Reviews Clinical Oncology, 2018, 15(1): 47-62. |
| [25] | VAN ERP E A, VAN KAMPEN M R, VAN KASTEREN P B, et al. Viral infection of human natural killer cells[J]. Viruses, 2019, 11(3): 243. |
| [26] | LE BERT N, TAN A T, KUNASEGARAN K, et al. SARS-CoV-2-specific T cell immunity in cases of COVID-19 and SARS, and uninfected controls[J]. Nature, 2020, 584(7821): 457-462. |
| [27] | DARVIN P, TOOR S M, SASIDHARAN NAIR V, et al. Immune checkpoint inhibitors: recent progress and potential biomarkers[J]. Experimental & Molecular Medicine, 2018, 50(12): 1-11. |
| [28] | KEIR M E, BUTTE M J, FREEMAN G J, et al. PD-1 and its ligands in tolerance and immunity[J]. Annual Review of Immunology, 2008, 26: 677-704. |
| [29] | CURRAN K J, PEGRAM H J, BRENTJENS R J. Chimeric antigen receptors for T cell immunotherapy: current understanding and future directions[J]. The Journal of Gene Medicine, 2012, 14(6): 405-415. |
| [30] | LINETTE G P, STADTMAUER E A, MAUS M V, et al. Cardiovascular toxicity and titin cross-reactivity of affinity-enhanced T cells in myeloma and melanoma[J]. Blood, 2013, 122(6): 863-871. |
| [31] | WAT J, BARMETTLER S. Hypogammaglobulinemia after chimeric antigen receptor (CAR) T-cell therapy: characteristics, management, and future directions[J]. The Journal of Allergy and Clinical Immunology: in Practice, 2022, 10(2): 460-466. |
| [32] | ARNETH B. Tumor microenvironment[J]. Medicina, 2020, 56(1): 15. |
| [33] | ZARETSKY J M, GARCIA-DIAZ A, SHIN D S, et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma[J]. The New England Journal of Medicine, 2016, 375(9): 819-829. |
| [34] | SHIN D S, ZARETSKY J M, ESCUIN-ORDINAS H, et al. Primary resistance to PD-1 blockade mediated by JAK1/2 mutations[J]. Cancer Discovery, 2017, 7(2): 188-201. |
| [35] | SHAH N N, FRY T J. Mechanisms of resistance to CAR T cell therapy[J]. Nature Reviews Clinical Oncology, 2019, 16(6): 372-385. |
| [36] | SIEGLER E L, ZHU Y N, WANG P, et al. Off-the-shelf CAR-NK cells for cancer immunotherapy[J]. Cell Stem Cell, 2018, 23(2): 160-161. |
| [37] | LIN J K, LERMAN B J, BARNES J I, et al. Cost effectiveness of chimeric antigen receptor T-cell therapy in relapsed or refractory pediatric B-cell acute lymphoblastic leukemia[J]. Journal of Clinical Oncology, 2018, 36(32): 3192-3202. |
| [38] | DEPIL S, DUCHATEAU P, GRUPP S A, et al. 'Off-the-shelf' allogeneic CAR T cells: development and challenges[J]. Nature Reviews Drug Discovery, 2020, 19(3): 185-199. |
| [39] | LIN H L, CHENG J L, MU W, et al. Advances in universal CAR-T cell therapy[J]. Frontiers in Immunology, 2021, 12: 744823. |
| [40] | ENDY D. Foundations for engineering biology[J]. Nature, 2005, 438(7067): 449-453. |
| [41] | ANDRIANANTOANDRO E, BASU S, KARIG D K, et al. Synthetic biology: new engineering rules for an emerging discipline[J]. Molecular Systems Biology, 2006, 2: 2006.0028. |
| [42] | GROUP B F, BAKER D, CHURCH G, et al. Engineering life: building a fab for biology[J]. Scientific American, 2006, 294(6): 44-51. |
| [43] | THIEL G, KAUFMANN A, RÖSSLER O G. G-protein-coupled designer receptors - new chemical-genetic tools for signal transduction research[J]. Biological Chemistry, 2013, 394(12): 1615-1622. |
| [44] | CONKLIN B R, HSIAO E C, CLAEYSEN S, et al. Engineering GPCR signaling pathways with RASSLs[J]. Nature Methods, 2008, 5(8): 673-678. |
| [45] | MCCUDDEN C R, HAINS M D, KIMPLE R J, et al. G-protein signaling: back to the future[J]. Cellular and Molecular Life Sciences, 2005, 62(5): 551-577. |
| [46] | WICKMAN K, CLAPHAM D E. Ion channel regulation by G proteins[J]. Physiological Reviews, 1995, 75(4): 865-885. |
| [47] | VAN BIESEN T, LUTTRELL L M, HAWES B E, et al. Mitogenic signaling via G protein-coupled receptors[J]. Endocrine Reviews, 1996, 17(6): 698-714. |
| [48] | SPIEGEL A M, SHENKER A, WEINSTEIN L S. Receptor-effector coupling by G proteins: implications for normal and abnormal signal transduction[J]. Endocrine Reviews, 1992, 13(3): 536-565. |
| [49] | COWARD P, WADA H G, FALK M S, et al. Controlling signaling with a specifically designed Gi-coupled receptor[J]. Proceedings of the National Academy of Sciences of the United States of America, 1998, 95(1): 352-357. |
| [50] | LEFKOWITZ R J, SHENOY S K. Transduction of receptor signals by beta-arrestins[J]. Science, 2005, 308(5721): 512-517. |
| [51] | SHENOY S K, LEFKOWITZ R J. β-Arrestin-mediated receptor trafficking and signal transduction[J]. Trends in Pharmacological Sciences, 2011, 32(9): 521-533. |
| [52] | REITER E, AHN S, SHUKLA A K, et al. Molecular mechanism of β-arrestin-biased agonism at seven-transmembrane receptors[J]. Annual Review of Pharmacology and Toxicology, 2012, 52: 179-197. |
| [53] | BARNEA G, STRAPPS W, HERRADA G, et al. The genetic design of signaling cascades to record receptor activation[J]. Proceedings of the National Academy of Sciences of the United States of America, 2008, 105(1): 64-69. |
| [54] | KIPNISS N H, DINGAL P C D P, ABBOTT T R, et al. Engineering cell sensing and responses using a GPCR-coupled CRISPR-Cas system[J]. Nature Communications, 2017, 8: 2212. |
| [55] | ANDERSSON E R, SANDBERG R, LENDAHL U. Notch signaling: simplicity in design, versatility in function[J]. Development, 2011, 138(17): 3593-3612. |
| [56] | WANG H F, ZANG C Z, LIU X S, et al. The role of Notch receptors in transcriptional regulation[J]. Journal of Cellular Physiology, 2015, 230(5): 982-988. |
| [57] | GORDON W R, VARDAR-ULU D, HISTEN G, et al. Structural basis for autoinhibition of Notch[J]. Nature Structural & Molecular Biology, 2007, 14(4): 295-300. |
| [58] | GORDON W R, ZIMMERMAN B, HE L, et al. Mechanical allostery: evidence for a force requirement in the proteolytic activation of Notch[J]. Developmental Cell, 2015, 33(6): 729-736. |
| [59] | STRUHL G, ADACHI A. Nuclear access and action of Notch in vivo [J]. Cell, 1998, 93(4): 649-660. |
| [60] | MORSUT L, ROYBAL K T, XIONG X, et al. Engineering customized cell sensing and response behaviors using synthetic Notch receptors[J]. Cell, 2016, 164(4): 780-791. |
| [61] | MORAGA I, SPANGLER J, MENDOZA J L, et al. Multifarious determinants of cytokine receptor signaling specificity[J]. Advances in Immunology, 2014, 121: 1-39. |
| [62] | MORAGA I, WERNIG G, WILMES S, et al. Tuning cytokine receptor signaling by re-orienting dimer geometry with surrogate ligands[J]. Cell, 2015, 160(6): 1196-1208. |
| [63] | MORAGA I, SPANGLER J B, MENDOZA J L, et al. Synthekines are surrogate cytokine and growth factor agonists that compel signaling through non-natural receptor dimers[J]. eLife, 2017, 6: e22882. |
| [64] | REMY I, WILSON I A, MICHNICK S W. Erythropoietin receptor activation by a ligand-induced conformation change[J]. Science, 1999, 283(5404): 990-993. |
| [65] | CONSTANTINESCU S N, KEREN T, RUSS W P, et al. The erythropoietin receptor transmembrane domain mediates complex formation with viral anemic and polycythemic gp55 proteins[J]. Journal of Biological Chemistry, 2003, 278(44): 43755-43763. |
| [66] | WITTHUHN B A, QUELLE F W, SILVENNOINEN O, et al. JAK2 associates with the erythropoietin receptor and is tyrosine phosphorylated and activated following stimulation with erythropoietin[J]. Cell, 1993, 74(2): 227-236. |
| [67] | SCHELLER L, STRITTMATTER T, FUCHS D, et al. Generalized extracellular molecule sensor platform for programming cellular behavior[J]. Nature Chemical Biology, 2018, 14(7): 723-729. |
| [68] | DARINGER N M, DUDEK R M, SCHWARZ K A, et al. Modular extracellular sensor architecture for engineering mammalian cell-based devices[J]. ACS Synthetic Biology, 2014, 3(12): 892-902. |
| [69] | HARTFIELD R M, SCHWARZ K A, MULDOON J J, et al. Multiplexing engineered receptors for multiparametric evaluation of environmental ligands[J]. ACS Synthetic Biology, 2017, 6(11): 2042-2055. |
| [70] | SCHWARZ K A, DARINGER N M, DOLBERG T B, et al. Rewiring human cellular input–output using modular extracellular sensors[J]. Nature Chemical Biology, 2017, 13(2): 202-209. |
| [71] | URBAN D J, ROTH B L. DREADDs (designer receptors exclusively activated by designer drugs): chemogenetic tools with therapeutic utility[J]. Annual Review of Pharmacology and Toxicology, 2015, 55: 399-417. |
| [72] | TODA S, BLAUCH L R, TANG S K Y, et al. Programming self-organizing multicellular structures with synthetic cell-cell signaling[J]. Science, 2018, 361(6398): 156-162. |
| [73] | BURRILL D R, SILVER P A. Making cellular memories[J]. Cell, 2010, 140(1): 13-18. |
| [74] | BROPHY J A N, MAGALLON K J, DUAN L N, et al. Synthetic genetic circuits as a means of reprogramming plant roots[J]. Science, 2022, 377(6607): 747-751. |
| [75] | WAN X Y, VOLPETTI F, PETROVA E, et al. Cascaded amplifying circuits enable ultrasensitive cellular sensors for toxic metals[J]. Nature Chemical Biology, 2019, 15(5): 540-548. |
| [76] | WEINBERG B H, CHO J H, AGARWAL Y, et al. High-performance chemical- and light-inducible recombinases in mammalian cells and mice[J]. Nature Communications, 2019, 10: 4845. |
| [77] | POTVIN-TROTTIER L, LORD N D, VINNICOMBE G, et al. Synchronous long-term oscillations in a synthetic gene circuit[J]. Nature, 2016, 538(7626): 514-517. |
| [78] | BERTSCHI A, WANG P L, GALVAN S, et al. Combinatorial protein dimerization enables precise multi-input synthetic computations[J]. Nature Chemical Biology, 2023, 19(6): 767-777. |
| [79] | CARRINGTON J C, DOUGHERTY W G. A viral cleavage site cassette: identification of amino acid sequences required for tobacco etch virus polyprotein processing[J]. Proceedings of the National Academy of Sciences of the United States of America, 1988, 85(10): 3391-3395. |
| [80] | TÖZSÉR J, TROPEA J E, CHERRY S, et al. Comparison of the substrate specificity of two potyvirus proteases[J]. The FEBS Journal, 2005, 272(2): 514-523. |
| [81] | BARTENSCHLAGER R. The NS3/4A proteinase of the hepatitis C virus: unravelling structure and function of an unusual enzyme and a prime target for antiviral therapy[J]. Journal of Viral Hepatitis, 1999, 6(3): 165-181. |
| [82] | GAO X J, CHONG L S, KIM M S, et al. Programmable protein circuits in living cells[J]. Science, 2018, 361(6408): 1252-1258. |
| [83] | COX A D, FESIK S W, KIMMELMAN A C, et al. Drugging the undruggable RAS: mission possible?[J]. Nature Reviews Drug Discovery, 2014, 13(11): 828-851. |
| [84] | DOWNWARD J. Targeting RAS signalling pathways in cancer therapy[J]. Nature Reviews Cancer, 2003, 3(1): 11-22. |
| [85] | FINK T, LONZARIĆ J, PRAZNIK A, et al. Design of fast proteolysis-based signaling and logic circuits in mammalian cells[J]. Nature Chemical Biology, 2019, 15(2): 115-122. |
| [86] | GAYET R V, ILIA K, RAZAVI S, et al. Autocatalytic base editing for RNA-responsive translational control[J]. Nature Communications, 2023, 14: 1339. |
| [87] | SNEPPEN K, KRISHNA S, SEMSEY S. Simplified models of biological networks[J]. Annual Review of Biophysics, 2010, 39: 43-59. |
| [88] | LIM W A, LEE C M, TANG C. Design principles of regulatory networks: searching for the molecular algorithms of the cell[J]. Molecular Cell, 2013, 49(2): 202-212. |
| [89] | INNISS M C, SILVER P A. Building synthetic memory[J]. Current Biology, 2013, 23(17): R812-R816. |
| [90] | FERRELL J E. Self-perpetuating states in signal transduction: positive feedback, double-negative feedback and bistability[J]. Current Opinion in Cell Biology, 2002, 14(2): 140-148. |
| [91] | PADIRAC A, FUJII T, RONDELEZ Y. Bottom-up construction of in vitro switchable memories[J]. Proceedings of the National Academy of Sciences of the United States of America, 2012, 109(47): E3212-E3220. |
| [92] | GOODWIN B B C. Temporal Organization in Cells: A Dynamic Theory of Cellular Control Processes[M]. London: Academic Press, 1963. |
| [93] | ZHOU Z, LIU Y T, FENG Y S, et al. Engineering longevity-design of a synthetic gene oscillator to slow cellular aging[J]. Science, 2023, 380(6643): 376-381. |
| [94] | KHAMMASH M H. Perfect adaptation in biology[J]. Cell Systems, 2021, 12(6): 509-521. |
| [95] | LANGAN R A, BOYKEN S E, NG A H, et al. De novo design of bioactive protein switches[J]. Nature, 2019, 572(7768): 205-210. |
| [96] | QUIJANO-RUBIO A, YEH H W, PARK J, et al. De novo design of modular and tunable protein biosensors[J]. Nature, 2021, 591(7850): 482-487. |
| [97] | ZHENG X H, WU Y Q, BI J C, et al. The use of supercytokines, immunocytokines, engager cytokines, and other synthetic cytokines in immunotherapy[J]. Cellular & Molecular Immunology, 2022, 19(2): 192-209. |
| [98] | VAZQUEZ-LOMBARDI R, LOETSCH C, ZINKL D, et al. Potent antitumour activity of interleukin-2-Fc fusion proteins requires Fc-mediated depletion of regulatory T-cells[J]. Nature Communications, 2017, 8: 15373. |
| [99] | DE LUCA R, SOLTERMANN A, PRETTO F, et al. Potency-matched dual cytokine-antibody fusion proteins for cancer therapy[J]. Molecular Cancer Therapeutics, 2017, 16(11): 2442-2451. |
| [100] | SEIF M, EINSELE H, LÖFFLER J. CAR T cells beyond cancer: hope for immunomodulatory therapy of infectious diseases[J]. Frontiers in Immunology, 2019, 10: 2711. |
| [101] | PRESTI E LO, DIELI F, MERAVIGLIA S. Tumor-infiltrating γδ T lymphocytes: pathogenic role, clinical significance, and differential programing in the tumor microenvironment[J]. Frontiers in Immunology, 2014, 5: 607. |
| [102] | TASIAN S K, GARDNER R A. CD19-redirected chimeric antigen receptor-modified T cells: a promising immunotherapy for children and adults with B-cell acute lymphoblastic leukemia (ALL)[J]. Therapeutic Advances in Hematology, 2015, 6(5): 228-241. |
| [103] | PARK J S, RHAU B, HERMANN A, et al. Synthetic control of mammalian-cell motility by engineering chemotaxis to an orthogonal bioinert chemical signal[J]. Proceedings of the National Academy of Sciences of the United States of America, 2014, 111(16): 5896-5901. |
| [104] | MAALEJ K M, MERHI M, INCHAKALODY V P, et al. CAR-cell therapy in the era of solid tumor treatment: current challenges and emerging therapeutic advances[J]. Molecular Cancer, 2023, 22(1): 20. |
| [105] | ALBELDA S M. CAR T cell therapy for patients with solid tumours: key lessons to learn and unlearn[J]. Nature Reviews Clinical Oncology, 2024, 21(1): 47-66. |
| [106] | BAI Z L, FENG B, MCCLORY S E, et al. Single-cell CAR T atlas reveals type 2 function in 8-year leukaemia remission[J]. Nature, 2024, 634(8034): 702-711. |
| [107] | BUI T A, MEI H Q, SANG R, et al. Advancements and challenges in developing in vivo CAR T cell therapies for cancer treatment[J]. eBioMedicine, 2024, 106: 105266. |
| [108] | XU J, LIU L, PARONE P, et al. In-vivo B-cell maturation antigen CAR T-cell therapy for relapsed or refractory multiple myeloma[J]. Lancet, 2025, 406(10500): 228-231. |
| [109] | MOHAMMADI M, NAJAFI H, MOHAMMADI P. CAR T-cell therapy in renal cell carcinoma: opportunities, challenges, and new strategies to overcome[J]. Medical Oncology, 2025, 42(6): 179. |
| [110] | LIU E L, MARIN D, BANERJEE P, et al. Use of CAR-transduced natural killer cells in CD19-positive lymphoid tumors[J]. The New England Journal of Medicine, 2020, 382(6): 545-553. |
| [111] | MYERS J A, MILLER J S. Exploring the NK cell platform for cancer immunotherapy[J]. Nature Reviews Clinical Oncology, 2021, 18(2): 85-100. |
| [112] | XIE G Z, DONG H, LIANG Y, et al. CAR-NK cells: a promising cellular immunotherapy for cancer[J]. eBioMedicine, 2020, 59: 102975. |
| [113] | KLICHINSKY M, RUELLA M, SHESTOVA O, et al. Human chimeric antigen receptor macrophages for cancer immunotherapy[J]. Nature Biotechnology, 2020, 38(8): 947-953. |
| [114] | ZHANG W L, LIU L, SU H F, et al. Chimeric antigen receptor macrophage therapy for breast tumours mediated by targeting the tumour extracellular matrix[J]. British Journal of Cancer, 2019, 121(10): 837-845. |
| [115] | KHAWAR M B, SUN H B. CAR-NK cells: from natural basis to design for kill[J]. Frontiers in Immunology, 2021, 12: 707542. |
| [116] | CHU J, DENG Y, BENSON D M, et al. CS1-specific chimeric antigen receptor (CAR)-engineered natural killer cells enhance in vitro and in vivo antitumor activity against human multiple myeloma[J]. Leukemia, 2014, 28(4): 917-927. |
| [117] | BOUTILIER A J, ELSAWA S F. Macrophage polarization states in the tumor microenvironment[J]. International Journal of Molecular Sciences, 2021, 22(13): 6995. |
| [118] | CHUNG H K, ZOU X Z, BAJAR B T, et al. A compact synthetic pathway rewires cancer signaling to therapeutic effector release[J]. Science, 2019, 364(6439): eaat6982. |
| [119] | WEI P, WONG W W, PARK J S, et al. Bacterial virulence proteins as tools to rewire kinase pathways in yeast and immune cells[J]. Nature, 2012, 488(7411): 384-388. |
| [120] | MAUDE S L, LAETSCH T W, BUECHNER J, et al. Tisagenlecleucel in children and young adults with B-cell lymphoblastic leukemia[J]. The New England Journal of Medicine, 2018, 378(5): 439-448. |
| [121] | PAN Q Z, WENG D S, LIU J Y, et al. Phase 1 clinical trial to assess safety and efficacy of NY-ESO-1-specific TCR T cells in HLA-a 02: 01 patients with advanced soft tissue sarcoma[J]. Cell Reports Medicine, 2023, 4(8): 101133. |
| [122] | ALLA S S M, TEKURU Y, LOKESH M S, et al. Talimogene laherparepvec (T-VEC) as a treatment for melanoma: a systematic review[J]. Journal of Oncology Pharmacy Practice, 2025, 31(3): 481-487. |
| [123] | CHO J H, OKUMA A, SOFJAN K, et al. Engineering advanced logic and distributed computing in human CAR immune cells[J]. Nature Communications, 2021, 12: 792. |
| [124] | VAKULSKAS C A, DEVER D P, RETTIG G R, et al. A high-fidelity Cas9 mutant delivered as a ribonucleoprotein complex enables efficient gene editing in human hematopoietic stem and progenitor cells[J]. Nature Medicine, 2018, 24(8): 1216-1224. |
| [125] | LU J R, JIANG G. The journey of CAR-T therapy in hematological malignancies[J]. Molecular Cancer, 2022, 21(1): 194. |
| [126] | SHIN J E, RIESSELMAN A J, KOLLASCH A W, et al. Protein design and variant prediction using autoregressive generative models[J]. Nature Communications, 2021, 12: 2403. |
| [127] | MARCHISIO M A, STELLING J. Computational design of synthetic gene circuits with composable parts[J]. Bioinformatics, 2008, 24(17): 1903-1910. |
| [128] | HALD ALBERTSEN C, KULKARNI J A, WITZIGMANN D, et al. The role of lipid components in lipid nanoparticles for vaccines and gene therapy[J]. Advanced Drug Delivery Reviews, 2022, 188: 114416. |
| [129] | ISSER A, LIVINGSTON N K, SCHNECK J P. Biomaterials to enhance antigen-specific T cell expansion for cancer immunotherapy[J]. Biomaterials, 2021, 268: 120584. |
| [1] | SONG Kainan, ZHANG Liwen, WANG Chao, TIAN Pingfang, LI Guangyue, PAN Guohui, XU Yuquan. Advances in small-molecule biopesticides and their biosynthesis [J]. Synthetic Biology Journal, 2025, 6(5): 1203-1223. |
| [2] | YU Wenwen, LV Xueqin, LI Zhaofeng, LIU Long. Plant synthetic biology and bioproduction of human milk oligosaccharides [J]. Synthetic Biology Journal, 2025, 6(5): 992-997. |
| [3] | YAN Zhaotao, ZHOU Pengfei, WANG Yangzhong, ZHANG Xin, XIE Wenyan, TIAN Chenfei, WANG Yong. Plant synthetic biology: new opportunities for large-scale culture of plant cells [J]. Synthetic Biology Journal, 2025, 6(5): 1107-1125. |
| [4] | SUN Yang, CHEN Lichao, SHI Yanyun, WANG Ke, LV Dandan, XU Xiumei, ZHANG Lixin. Strategies and prospects of synthetic biology in crop photosynthesis [J]. Synthetic Biology Journal, 2025, 6(5): 1025-1040. |
| [5] | ZHAO Xinyu, SHENG Qi, LIU Kaifang, LIU Jia, LIU Liming. Construction of microbial cell factories for aspartate-family feed amino acids [J]. Synthetic Biology Journal, 2025, 6(5): 1184-1202. |
| [6] | HE Yangyu, YANG Kai, WANG Weilin, HUANG Qian, QIU Ziying, SONG Tao, HE Liushang, YAO Jinxin, GAN Lu, HE Yuchi. Design and practice of plant synthetic biology theme in the International Genetically Engineered Machine Competition [J]. Synthetic Biology Journal, 2025, 6(5): 1243-1254. |
| [7] | ZHANG Xuebo, ZHU Chengshu, CHEN Ruiyun, JIN Qingzi, LIU Xiao, XIONG Yan, CHEN Daming. Policy planning and industrial development of agricultural synthetic biology [J]. Synthetic Biology Journal, 2025, 6(5): 1224-1242. |
| [8] | LIU Jie, GAO Yu, MA Yongshuo, SHANG Yi. Progress and challenges of synthetic biology in agriculture [J]. Synthetic Biology Journal, 2025, 6(5): 998-1024. |
| [9] | ZHENG Lei, ZHENG Qiteng, ZHANG Tianjiao, DUAN Kun, ZHANG Ruifu. Engineering rhizosphere synthetic microbial communities to enhance crop nutrient use efficiency [J]. Synthetic Biology Journal, 2025, 6(5): 1058-1071. |
| [10] | LI Chao, ZHANG Huan, YANG Jun, WANG Ertao. Research advances in nitrogen fixation synthetic biology [J]. Synthetic Biology Journal, 2025, 6(5): 1041-1057. |
| [11] | WEI Jiaxiu, JI Peiyun, JIE Qingyu, HUANG Qiuyan, YE Hao, DAI Junbiao. Construction and application of plant artificial chromosomes [J]. Synthetic Biology Journal, 2025, 6(5): 1093-1106. |
| [12] | FANG Xinyi, SUN Lichao, HUO Yixin, WANG Ying, YUE Haitao. Trends and challenges in microbial synthesis of higher alcohols [J]. Synthetic Biology Journal, 2025, 6(4): 873-898. |
| [13] | WU Xiaoyan, SONG Qi, XU Rui, DING Chenjun, CHEN Fang, GUO Qing, ZHANG Bo. A comparative analysis of global research and development competition in synthetic biology [J]. Synthetic Biology Journal, 2025, 6(4): 940-955. |
| [14] | ZHANG Jiankang, WANG Wenjun, GUO Hongju, BAI Beichen, ZHANG Yafei, YUAN Zheng, LI Yanhui, LI Hang. Development and application of a high-throughput microbial clone picking workstation based on machine vision [J]. Synthetic Biology Journal, 2025, 6(4): 956-971. |
| [15] | LI Quanfei, CHEN Qian, LIU Hao, HE Kundong, PAN Liang, LEI Peng, GU Yi’an, SUN Liang, LI Sha, QIU Yibin, WANG Rui, XU Hong. Synthetic biology and applications of high-adhesion protein materials [J]. Synthetic Biology Journal, 2025, 6(4): 806-828. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||