Synthetic Biology Journal ›› 2021, Vol. 2 ›› Issue (6): 902-919.DOI: 10.12211/2096-8280.2021-049
• Invited Review • Previous Articles Next Articles
Progress in artificial metabolic pathways for biosynthesis of organic alcohols & acids
CAO Chenkai, LI Jialong, ZHANG Kechun
- Key Laboratory of Coastal Environment and Resources of Zhejiang Province,School of Engineering,Westlake University,Hangzhou 310024,Zhejiang,China
-
Received:2021-04-23Revised:2021-11-19Online:2022-01-21Published:2021-12-31 -
Contact:ZHANG Kechun
人工代谢途径合成有机醇有机酸的研究进展
曹晨凯, 李佳隆, 张科春
- 西湖大学工学院,浙江省海岸带环境与资源研究重点实验室,浙江 杭州 310024
-
通讯作者:张科春 -
作者简介:曹晨凯 (1996—),男,博士研究生。研究方向为代谢工程。E-mail:caochenkai@westlake.edu.cn李佳隆 (1996—),男,博士研究生。研究方向为代谢工程。E-mail:lijialong@westlake.edu.cn张科春 (1978—),特聘研究员,博士生导师。研究方向为应用合成生物、绿色化学、材料科学和工程优化的方法,设计绿色新化工生产路线和开发环保新材料,为循环经济向前发展提供新的解决方案。E-mail:zhangkechun@westlake.edu.cn -
基金资助:国家自然科学基金(22078267)
CLC Number:
Cite this article
CAO Chenkai, LI Jialong, ZHANG Kechun. Progress in artificial metabolic pathways for biosynthesis of organic alcohols & acids[J]. Synthetic Biology Journal, 2021, 2(6): 902-919.
曹晨凯, 李佳隆, 张科春. 人工代谢途径合成有机醇有机酸的研究进展[J]. 合成生物学, 2021, 2(6): 902-919.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2021-049
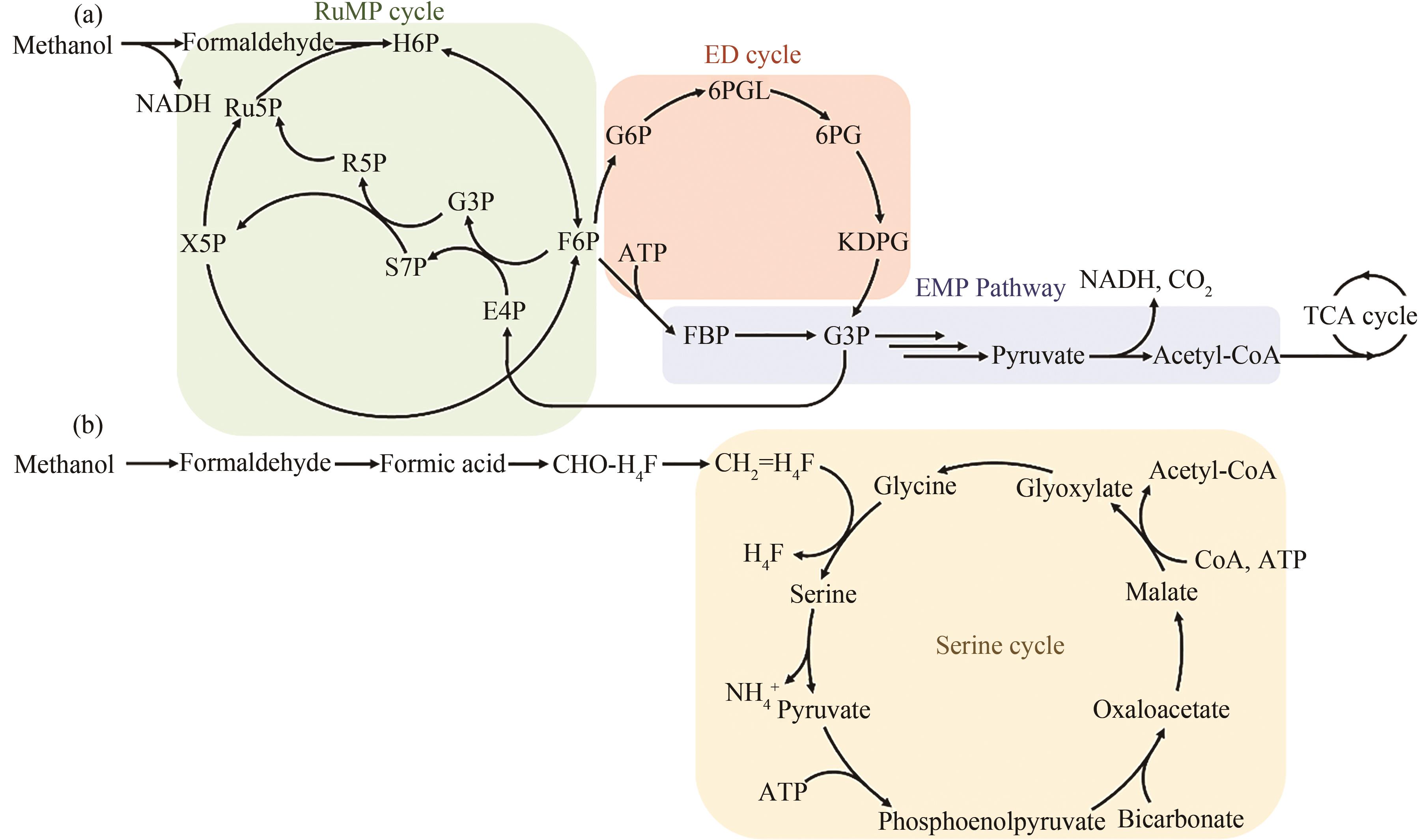
Fig. 1 C1 compound assimilation through ribulose-5-phosphate (RuMP), ED (Entner-Doudoroff), EMP (Embden-Meyerhof-Parnas) pathway (a) and serine cycle (b)[19]
| 菌株 | 方法 | 培养基 | 产物 | 产量/(g/L) | 时间 | 文献 |
|---|---|---|---|---|---|---|
| 甲醇芽孢杆菌 | 补料分批发酵 | 含200 mmol/L甲醇的MVcM培养基 | γ-氨基丁酸 | 9.0 | 2016 | [ |
| α-变形杆菌甲基杆菌AM1 | 摇瓶发酵 | 含20 mmol/L乙胺的基础培养基 | L-丁醇 | 25.5×10-3 | 2016 | [ |
| 扭脱甲基杆菌AM1 | 补料分批发酵 | 含有3 g/L甲醇的矿物盐培养基 | 2-羟基异丁酸 | 2.1 | 2016 | [ |
| 扭脱甲基杆菌AM1 | 摇瓶发酵 | 含125 mmol/L甲醇的基础培养基 | 3-羟基丙酸 | 69.8×10-3 | 2017 | [ |
| α-变形杆菌甲基杆菌AM1 | 补料分批发酵 | 含5%(体积分数)甲醇的基础培养基 | 甲羟戊酸 | 2.3 | 2018 | [ |
| 毕赤酵母 | 试管发酵 | 酵母/蛋白胨/葡萄糖培养基 | D-乳酸 | 3.5 | 2019 | [ |
Tab. 1 Summary of methylotrophic production data
| 菌株 | 方法 | 培养基 | 产物 | 产量/(g/L) | 时间 | 文献 |
|---|---|---|---|---|---|---|
| 甲醇芽孢杆菌 | 补料分批发酵 | 含200 mmol/L甲醇的MVcM培养基 | γ-氨基丁酸 | 9.0 | 2016 | [ |
| α-变形杆菌甲基杆菌AM1 | 摇瓶发酵 | 含20 mmol/L乙胺的基础培养基 | L-丁醇 | 25.5×10-3 | 2016 | [ |
| 扭脱甲基杆菌AM1 | 补料分批发酵 | 含有3 g/L甲醇的矿物盐培养基 | 2-羟基异丁酸 | 2.1 | 2016 | [ |
| 扭脱甲基杆菌AM1 | 摇瓶发酵 | 含125 mmol/L甲醇的基础培养基 | 3-羟基丙酸 | 69.8×10-3 | 2017 | [ |
| α-变形杆菌甲基杆菌AM1 | 补料分批发酵 | 含5%(体积分数)甲醇的基础培养基 | 甲羟戊酸 | 2.3 | 2018 | [ |
| 毕赤酵母 | 试管发酵 | 酵母/蛋白胨/葡萄糖培养基 | D-乳酸 | 3.5 | 2019 | [ |
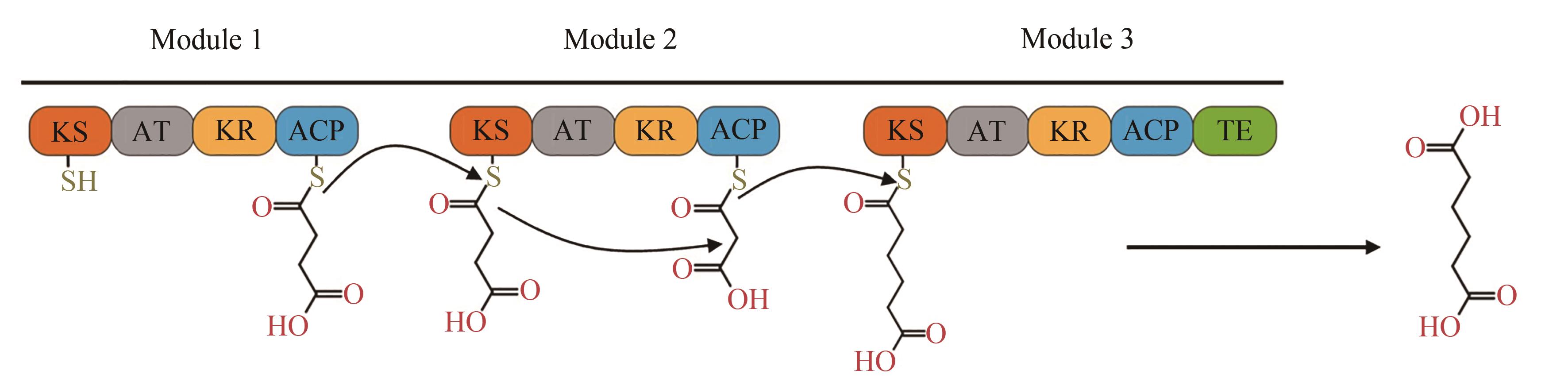
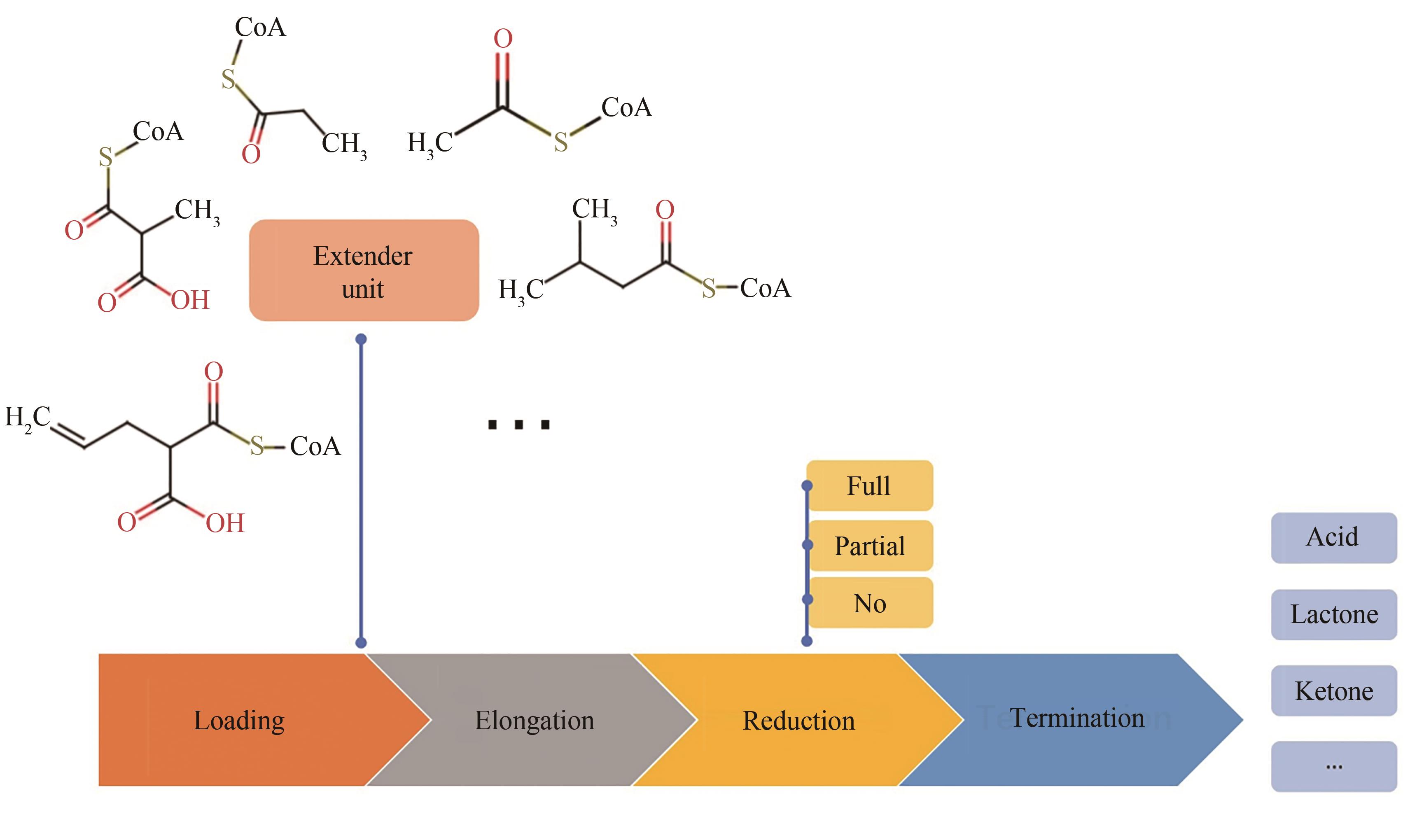
Fig. 6 PKS pathway for production of adipic acid[There are 3 respective modules. Substrate is transferred from previous module to next one via thioester exchange reaction, and finally released by thioesterase (TEp) domain]
| 1 | 丁明珠, 李炳志, 王颖, 等. 合成生物学重要研究方向进展[J]. 合成生物学, 2020, 1(1): 7-28. |
| DING M Z, LI B Z, WANG Y, et al. Significant research progress in synthetic biology[J]. Synthetic Biology Journal, 2020, 1(1): 7-28. | |
| 2 | PACALA S, SOCOLOW R. Stabilization wedges: Solving the climate problem for the next 50 years with current technologies[J]. Science, 2004, 305(5686): 968-972. |
| 3 | O'NEILL B C, OPPENHEIMER M. Climate change. Dangerous climate impacts and the Kyoto protocol[J]. Science, 2002, 296(5575): 1971-1972. |
| 4 | 任光. 我国煤制甲醇的工业现状及发展趋势分析[J]. 化肥设计, 2016, 54(5): 5-7. |
| REN G. Analysis on present situation and development trend of coal methanol industry in China[J]. Chemical Fertilizer Design, 2016, 54(5): 5-7. | |
| 5 | 高姣丽, 潘子鹤, 成怀刚, 等. CO2催化制甲酸反应设备的现状与研究进展[J]. 化工机械, 2020, 47(6): 737-741. |
| GAO J L, PAN Z H, CHENG H G, et al. Status and research progress of reaction equipment for preparation of formic acid catalyzed by CO2 [J]. Chemical Engineering & Machinery, 2020, 47(6): 737-741. | |
| 6 | CHEN X, LIU Y, WU J W. Sustainable production of formic acid from biomass and carbon dioxide[J]. Molecular Catalysis, 2020, 483: 110716. |
| 7 | LI H, OPGENORTH P H, WERNICK D G, et al. Integrated electromicrobial conversion of CO2 to higher alcohols[J]. Science, 2012, 335(6076): 1596. |
| 8 | SAWATDEENARUNAT C, NGUYEN D, SURENDRA K C, et al. Anaerobic biorefinery: current status, challenges, and opportunities[J]. Bioresource Technology, 2016, 215: 304-313. |
| 9 | CLOMBURG J M, CRUMBLEY A M, GONZALEZ R. Industrial biomanufacturing: the future of chemical production [J]. Science, 2017, 355(6320): aag0804. |
| 10 | ZENG A P. New bioproduction systems for chemicals and fuels: Needs and new development[J]. Biotechnology Advances, 2019, 37(4): 508-518. |
| 11 | CHEN F Y H, JUNG H W, TSUEI C Y, et al. Converting Escherichia coli to a synthetic methylotroph growing solely on methanol[J]. Cell, 2020, 182(4): 933-946. |
| 12 | BODEN R, CUNLIFFE M, SCANLAN J, et al. Complete genome sequence of the aerobic marine methanotroph Methylomonas methanica MC09[J]. Journal of Bacteriology, 2011, 193(24): 7001-7002. |
| 13 | PEEL D, QUAYLE J R. Microbial growth on C1 compounds (I): Isolation and characterization of Pseudomonas AM 1[J]. Biochemical Journal, 1961, 81(3): 465-469. |
| 14 | STROM T, FERENCI T, QUAYLE J R. The carbon assimilation pathways of Methylococcus capsulatus, Pseudomonas methanica and Methylosinus trichosporium (OB3B) during growth on methane[J]. The Biochemical Journal, 1974, 144(3): 465-476. |
| 15 | WHITAKER W B, JONES J A, BENNETT R K, et al. Engineering the biological conversion of methanol to specialty chemicals in Escherichia coli [J]. Metabolic Engineering, 2017, 39: 49-59. |
| 16 | KALYUZHNAYA M G, YANG S, ROZOVA O N, et al. Highly efficient methane biocatalysis revealed in a methanotrophic bacterium[J]. Nature Communications, 2013, 4(1): 2785. |
| 17 | MEYER F, KELLER P, HARTL J, et al. Methanol-essential growth of Escherichia coli [J]. Nature Communications, 2018, 9(1): 1508. |
| 18 | WOOLSTON B M, KING J R, REITER M, et al. Improving formaldehyde consumption drives methanol assimilation in engineered E. coli [J]. Nature Communications, 2018, 9(1): 2387-2398. |
| 19 | YU H, LIAO J C. A modified serine cycle in Escherichia coli coverts methanol and CO2 to two-carbon compounds[J]. Nature Communications, 2018, 9(1): 3992. |
| 20 | 郭姝媛, 吴良焕, 刘香健, 等. 微生物中一碳代谢网络构建的进展与挑战[J/OL]. 合成生物学: 1-24, doi: 10.12211/2096-8280.2021-079 . . |
| GUO Shuyuan, WU Lianghuan, LIU Xiangjian, et al. The biotransformation outlook of C1-based metabolic networks in methylotrophy[J/OL]. Synthetic Biology Journal: 1-24, doi: 10.12211/2096-8280.2021-079 . . | |
| 21 | GOYAL N, ZHOU Z, KARIMI I A. Metabolic processes of Methanococcus maripaludis and potential applications[J]. Microbial Cell Factories, 2016, 15(1): 107. |
| 22 | SOHN Y J, SON J, JO S Y, et al. Chemoautotroph Cupriavidus necator as a potential game-changer for global warming and plastic waste problem: a review[J]. Bioresource Technology, 2021, 340: 125693. |
| 23 | KOZLOWSKI J T, DAVIS R J. Heterogeneous catalysts for the guerbet coupling of alcohols[J]. ACS Catalysis, 2013, 3(7): 1588-1600. |
| 24 | LIAO J C, MI L, PONTRELLI S, et al. Fuelling the future: microbial engineering for the production of sustainable biofuels [J]. Nature Reviews Microbiology, 2016, 14(5): 288-304. |
| 25 | IRLA M, NÆRDAL I, BRAUTASET T, et al. Methanol-based γ-aminobutyric acid (GABA) production by genetically engineered Bacillus methanolicus strains[J]. Industrial Crops and Products, 2017, 106: 12-20. |
| 26 | HU B, YANG Y M, BECK D A C, et al. Comprehensive molecular characterization of Methylobacterium extorquens AM1 adapted for 1-butanol tolerance[J]. Biotechnology for Biofuels, 2016, 9(1): 84-97. |
| 27 | ROHDE M T, TISCHER S, HARMS H, et al. Production of 2-hydroxyisobutyric acid from methanol by Methylobacterium extorquens AM1 expressing (R)-3-hydroxybutyryl-CoA isomerizing enzymes [J]. Applied & Environmental Microbiology, 2017, 83(3): AEM.02622-16. |
| 28 | YANG Y M, CHEN W J, YANG J, et al. Production of 3-hydroxypropionic acid in engineered Methylobacterium extorquens AM1 and its reassimilation through a reductive route[J]. Microbial Cell Factories, 2017, 16(1): 179. |
| 29 | CUI L Y, WANG S S, GUAN C G, et al. Breeding of methanol-tolerant Methylobacterium extorquens AM1 by atmospheric and room temperature plasma mutagenesis combined with adaptive laboratory evolution[J]. Biotechnology Journal, 2018, 13(6): e1700679. |
| 30 | YAMADA R, OGURA K, KIMOTO Y, et al. Toward the construction of a technology platform for chemicals production from methanol: D-lactic acid production from methanol by an engineered yeast Pichia pastoris [J]. World Journal of Microbiology and Biotechnology, 2019, 35(2): 37-45. |
| 31 | GODDARD A W. Cortical and subcortical gamma amino acid butyric acid deficits in anxiety and stress disorders: clinical implications[J]. World Journal of Psychiatry, 2016, 6(1): 43-53. |
| 32 | BOGORAD I W, CHEN C T, THEISEN M K, et al. Building carbon-carbon bonds using a biocatalytic methanol condensation cycle[J]. Proceedings of the National Academy of Sciences of the United States of America, 2014, 111(45): 15928-15933. |
| 33 | TILMAN D, SOCOLOW R, FOLEY J A, et al. Beneficial biofuels—the food, energy, and environment trilemma[J]. Science, 2009, 325(5938): 270-271. |
| 34 | GRAHAM-ROWE D. Agriculture: beyond food versus fuel[J]. Nature, 2011, 474(7352): S6-S8. |
| 35 | PERLACK R D, WRIGHT L L, TURHOLLOW A F, et al. Biomass as feedstock for a bioenergy and bioproducts industry: The technical feasibility of a billion-ton annual supply[R]. Office of Scientific and Technical Information (OSTI), 2005. |
| 36 | SOMERVILLE C. The billion-ton biofuels vision[J]. Science, 2006, 312(5778): 1277. |
| 37 | RAGAUSKAS A J, WILLIAMS C K, DAVISON B H, et al. The path forward for biofuels and biomaterials[J]. Science, 2006, 311(5760): 484-489. |
| 38 | GALL D L, RALPH J, DONOHUE T J, et al. Biochemical transformation of lignin for deriving valued commodities from lignocellulose[J]. Current Opinion in Biotechnology, 2017, 45: 120-126. |
| 39 | QIN L, LI W C, LIU L, et al. Inhibition of lignin-derived phenolic compounds to cellulase[J]. Biotechnology for Biofuels, 2016, 9: 70. |
| 40 | LI W C, ZHU J Q, ZHAO X, et al. Improving co-fermentation of glucose and xylose by adaptive evolution of engineering xylose-fermenting Saccharomyces cerevisiae and different fermentation strategies[J]. Renewable Energy, 2019, 139: 1176-1183. |
| 41 | LIU H, ZHU J Q, LI X, et al. Hybridization improves inhibitor tolerance of xylose-fermenting Saccharomyces cerevisiae [J]. BioResources, 2017, 12(3): 4737-4753. |
| 42 | JEFFRIES T W. Engineering yeasts for xylose metabolism[J]. Current Opinion in Biotechnology, 2006, 17(3): 320-326. |
| 43 | MATSUSHIKA A, INOUE H, KODAKI T, et al. Ethanol production from xylose in engineered Saccharomyces cerevisiae strains: current state and perspectives [J]. Applied Microbiology and Biotechnology, 2009, 84(1): 37-53. |
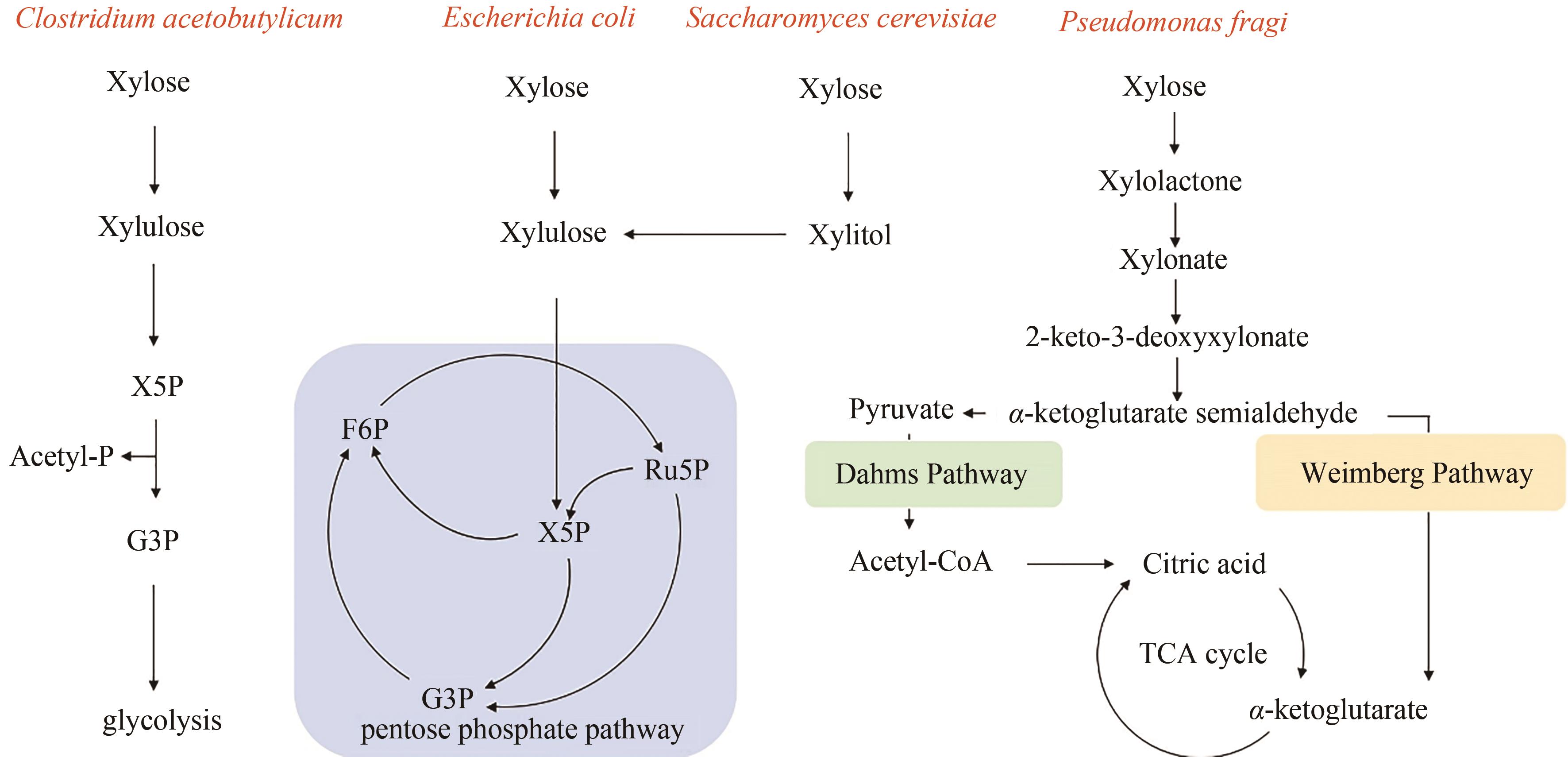
| 44 | LIU L X, ZHANG L, TANG W, et al. Phosphoketolase pathway for xylose catabolism in Clostridium acetobutylicum revealed by 13C metabolic flux analysis[J]. Journal of Bacteriology, 2012, 194(19): 5413-5422. |
| 45 | TANAKA K, KOMIYAMA A, SONOMOTO K, et al. Two different pathways for D-xylose metabolism and the effect of xylose concentration on the yield coefficient of L-lactate in mixed-acid fermentation by the lactic acid bacterium Lactococcus lactis IO-1[J]. Applied Microbiology and Biotechnology, 2002, 60(1/2): 160-167. |
| 46 | SCALCINATI G, OTERO J M, VLEET J R H VAN, et al. Evolutionary engineering of Saccharomyces cerevisiae for efficient aerobic xylose consumption[J]. FEMS Yeast Research, 2012, 12(5): 582-597. |
| 47 | WEIMBERG R. Pentose oxidation by Pseudomonas fragi [J]. Journal of Biological Chemistry, 1961, 236(3): 629-635. |
| 48 | DAHMS A S. 3-Deoxy-D-pentulosonic acid aldolase and its role in a new pathway of D-xylose degradation [J]. Biochemical and Biophysical Research Communications, 1974, 60(4): 1433-9. |
| 49 | KUHAD R C, GUPTA R, KHASA Y P, et al. Bioethanol production from pentose sugars: current status and future prospects[J]. Renewable and Sustainable Energy Reviews, 2011, 15(9): 4950-4962. |
| 50 | KIM S R, PARK Y C, JIN Y S, et al. Strain engineering of Saccharomyces cerevisiae for enhanced xylose metabolism[J]. Biotechnology Advances, 2013, 31(6): 851-861. |
| 51 | QI X, ZHA J, LIU G-G, et al. Heterologous xylose isomerase pathway and evolutionary engineering improve xylose utilization in Saccharomyces cerevisiae [J]. Frontiers in Microbiology, 2015, 6: 1165. |
| 52 | GOPINARAYANAN V E, NAIR N U. Pentose metabolism in Saccharomyces cerevisiae: the need to engineer global regulatory systems[J]. Biotechnology Journal, 2019, 14(1): e1800364. |
| 53 | KWAK S, JIN Y S. Production of fuels and chemicals from xylose by engineered Saccharomyces cerevisiae: a review and perspective [J]. Microbial Cell Factories, 2017, 16(1): 82-96. |
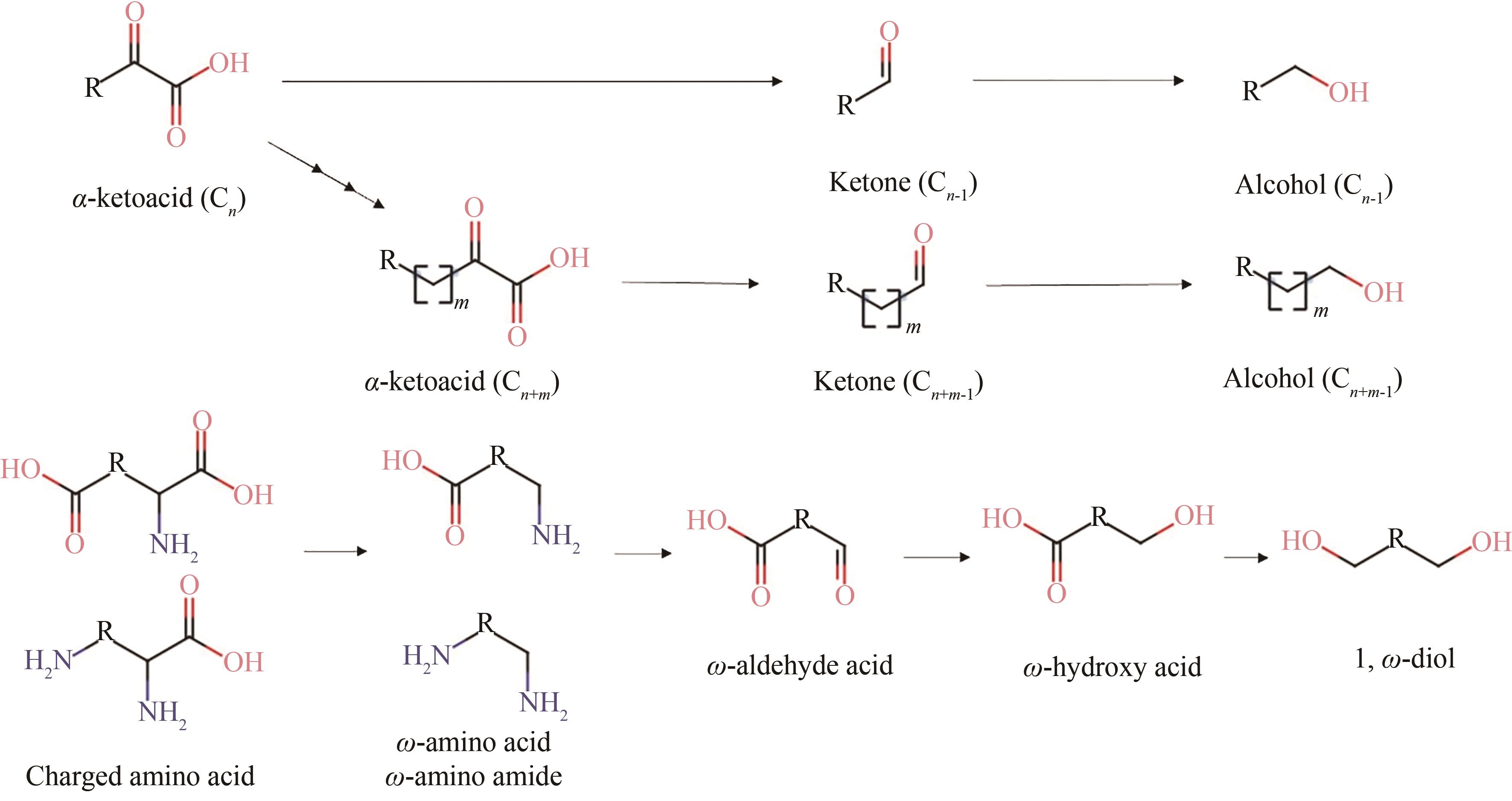
| 54 | ZHA J, SHEN M H, HU M L, et al. Enhanced expression of genes involved in initial xylose metabolism and the oxidative pentose phosphate pathway in the improved xylose-utilizing Saccharomyces cerevisiae through evolutionary engineering[J]. Journal of Industrial Microbiology & Biotechnology, 2014, 41(1): 27-39. |
| 55 | HOU J, SHEN Y, JIAO C L, et al. Characterization and evolution of xylose isomerase screened from the bovine rumen metagenome in Saccharomyces cerevisiae [J]. Journal of Bioscience and Bioengineering, 2016, 121(2): 160-165. |
| 56 | KATAHIRA S, MURAMOTO N, MORIYA S, et al. Screening and evolution of a novel protist xylose isomerase from the termite Reticulitermes speratus for efficient xylose fermentation in Saccharomyces cerevisiae [J]. Biotechnology for Biofuels, 2017, 10(1): 203-220. |
| 57 | SONDEREGGER M, SAUER U. Evolutionary engineering of Saccharomyces cerevisiae for anaerobic growth on xylose[J]. Applied and Environmental Microbiology, 2003, 69(4): 1990-1998. |
| 58 | MADHAVAN A, TAMALAMPUDI S, USHIDA K, et al. Xylose isomerase from polycentric fungus Orpinomyces: gene sequencing, cloning, and expression in Saccharomyces cerevisiae for bioconversion of xylose to ethanol[J]. Applied Microbiology and Biotechnology, 2009, 82(6): 1067-1078. |
| 59 | KUYPER M, TOIRKENS M J, DIDERICH J A, et al. Evolutionary engineering of mixed-sugar utilization by a xylose-fermenting Saccharomyces cerevisiae strain[J]. FEMS Yeast Research, 2005, 5(10): 925-934. |
| 60 | HOU J, QIU C X, SHEN Y, et al. Engineering of Saccharomyces cerevisiae for the efficient co-utilization of glucose and xylose[J]. FEMS Yeast Research, 2017, 17(4): fox034. |
| 61 | WANG X, JIN M J, BALAN V, et al. Comparative metabolic profiling revealed limitations in xylose-fermenting yeast during co-fermentation of glucose and xylose in the presence of inhibitors[J]. Biotechnology and Bioengineering, 2014, 111(1): 152-164. |
| 62 | LÜ Y J, WANG X, MA Q, et al. Proteomic analysis reveals complex metabolic regulation in Saccharomyces cerevisiae cells against multiple inhibitors stress[J]. Applied Microbiology and Biotechnology, 2014, 98(5): 2207-2221. |
| 63 | ZHA J, LI B Z, SHEN M H, et al. Optimization of CDT-1 and XYL1 expression for balanced co-production of ethanol and xylitol from cellobiose and xylose by engineered Saccharomyces cerevisiae [J]. PLoS One, 2013, 8(7): e68317. |
| 64 | DUNN K L, RAO C V. High-throughput sequencing reveals adaptation-induced mutations in pentose-fermenting strains of Zymomonas mobilis [J]. Biotechnology and Bioengineering, 2015, 112(11): 2228-2240. |
| 65 | DUNN K L, RAO C V. Expression of a xylose-specific transporter improves ethanol production by metabolically engineered Zymomonas mobilis [J]. Applied Microbiology and Biotechnology, 2014, 98(15): 6897-6905. |
| 66 | YOUNG E M, TONG A, BUI H, et al. Rewiring yeast sugar transporter preference through modifying a conserved protein motif[J]. Proceedings of the National Academy of Sciences of the United States of America, 2014, 111(1): 131-136. |
| 67 | SARKAR P, MUKHERJEE M, GOSWAMI G, et al. Adaptive laboratory evolution induced novel mutations in Zymomonas mobilis ATCC ZW658: a potential platform for co-utilization of glucose and xylose[J]. Journal of Industrial Microbiology & Biotechnology, 2020, 47(3): 329-341. |
| 68 | SARKAR P, GOSWAMI G, MUKHERJEE M, et al. Heterologous expression of xylose specific transporter improves xylose utilization by recombinant Zymomonas mobilis strain in presence of glucose[J]. Process Biochemistry, 2021, 102: 190-198. |
| 69 | TAI Y S, XIONG M, JAMBUNATHAN P, et al. Engineering nonphosphorylative metabolism to generate lignocellulose-derived products[J]. Nature Chemical Biology, 2016, 12(4): 247-253. |
| 70 | LIU H W, LU T. Autonomous production of 1,4-butanediol via a de novo biosynthesis pathway in engineered Escherichia coli [J]. Metabolic Engineering, 2015, 29: 135-141. |
| 71 | WANG J, JAIN R, SHEN X L, et al. Rational engineering of diol dehydratase enables 1,4-butanediol biosynthesis from xylose[J]. Metabolic Engineering, 2017, 40: 148-156. |
| 72 | WANG J, SHEN X L, JAIN R, et al. Establishing a novel biosynthetic pathway for the production of 3,4-dihydroxybutyric acid from xylose in Escherichia coli [J]. Metabolic Engineering, 2017, 41: 39-45. |
| 73 | WANG J, SHEN X L, LIN Y H, et al. Investigation of the synergetic effect of xylose metabolic pathways on the production of glutaric acid[J]. ACS Synthetic Biology, 2018, 7(1): 24-29. |
| 74 | BAI W Q, TAI Y S, WANG J Y, et al. Engineering nonphosphorylative metabolism to synthesize mesaconate from lignocellulosic sugars in Escherichia coli [J]. Metabolic Engineering, 2016, 38: 285-292. |
| 75 | WEI N, QUARTERMAN J, KIM S R, et al. Enhanced biofuel production through coupled acetic acid and xylose consumption by engineered yeast[J]. Nature Communications, 2013, 4: 2580. |
| 76 | MEILGAARD M C. Flavor chemistry of beer (II): Flavor and threshold of 239 aroma volatiles [J]. Techquartmasterbrewassocam, 1975, 12: 22-8. |
| 77 | EHRLICH F. Über die Bedingungen der Fuselölbildung und über ihren Zusammenhang mit dem Eiweißaufbau der Hefe [J]. Berichte der Deutschen Chemischen Gesellschaft, 1907, 40(1): 1027-1047. |
| 78 | HOHMANN S. Characterisation of PDC2, a gene necessary for high level expression of pyruvate decarboxylase structural genes in Saccharomyces cerevlsiae [J]. Molecular and General Genetics MGG, 1993, 241(5/6): 657-666. |
| 79 | HOHMANN S. Characterization of PDC6, a third structural gene for pyruvate decarboxylase in Saccharomyces cerevisiae [J]. Journal of Bacteriology, 1991, 173(24): 7963-7969. |
| 80 | HOHMANN S, CEDERBERG H. Autoregulation may control the expression of yeast pyruvate decarboxylase structural genes PDC1 and PDC5 [J]. European Journal of Biochemistry, 1990, 188(3): 615-621. |
| 81 | VURALHAN Z, MORAIS M A, TAI S L, et al. Identification and characterization of phenylpyruvate decarboxylase genes in Saccharomyces cerevisiae [J]. Applied and Environmental Microbiology, 2003, 69(8): 4534-4541. |
| 82 | VURALHAN Z, LUTTIK M A, TAI S L, et al. Physiological characterization of the ARO10-dependent, broad-substrate-specificity 2-oxo acid decarboxylase activity of Saccharomyces cerevisiae [J]. Applied and Environmental Microbiology, 2005, 71(6): 3276-3284. |
| 83 | IRAQUI I, VISSERS S, ANDRÉ B, et al. Transcriptional induction by aromatic amino acids in Saccharomyces cerevisiae [J]. Molecular and Cellular Biology, 1999, 19(5): 3360-3371. |
| 84 | MOJZITA D, HOHMANN S. Pdc2 coordinates expression of the THI regulon in the yeast Saccharomyces cerevisiae [J]. Molecular Genetics and Genomics, 2006, 276(2): 147-161. |
| 85 | NOSAKA K, ONOZUKA M, KONNO H, et al. Genetic regulation mediated by thiamin pyrophosphate-binding motif in Saccharomyces cerevisiae [J]. Molecular Microbiology, 2005, 58(2): 467-479. |
| 86 | SMIT G, SMIT B A, ENGELS W J M. Flavour formation by lactic acid bacteria and biochemical flavour profiling of cheese products[J]. FEMS Microbiology Reviews, 2005, 29(3): 591-610. |
| 87 | ATSUMI S, HANAI T, LIAO J C. Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels[J]. Nature, 2008, 451(7174): 86-89. |
| 88 | KÖNIG S. Subunit structure, function and organisation of pyruvate decarboxylases from various organisms[J]. Biochimica et Biophysica Acta, 1998, 1385(2): 271-286. |
| 89 | CONNOR M R, CANN A F, LIAO J C. 3-Methyl-1-butanol production in Escherichia coli: random mutagenesis and two-phase fermentation[J]. Applied Microbiology and Biotechnology, 2010, 86(4): 1155-1164. |
| 90 | ATSUMI S, WU T Y, ECKL E M, et al. Engineering the isobutanol biosynthetic pathway in Escherichia coli by comparison of three aldehyde reductase/alcohol dehydrogenase genes[J]. Applied Microbiology and Biotechnology, 2010, 85(3): 651-657. |
| 91 | CANN A F, LIAO J C. Production of 2-methyl-1-butanol in engineered Escherichia coli [J]. Applied Microbiology and Biotechnology, 2008, 81(1): 89-98. |
| 92 | ZHANG K C, SAWAYA M R, EISENBERG D S, et al. Expanding metabolism for biosynthesis of nonnatural alcohols[J]. Proceedings of the National Academy of Sciences of the United States of America, 2008, 105(52): 20653-20658. |
| 93 | XIONG M, DENG J, WOODRUFF A P, et al. A bio-catalytic approach to aliphatic ketones[J]. Scientific Reports, 2012, 2: 311. |
| 94 | WENDISCH V F. Metabolic engineering advances and prospects for amino acid production[J]. Metabolic Engineering, 2020, 58: 17-34. |
| 95 | LIN P P, MI L, MORIOKA A H, et al. Consolidated bioprocessing of cellulose to isobutanol using Clostridium thermocellum [J]. Metabolic Engineering, 2015, 31: 44-52. |
| 96 | LI S S, WEN J P, JIA X Q. Engineering Bacillus subtilis for isobutanol production by heterologous Ehrlich pathway construction and the biosynthetic 2-ketoisovalerate precursor pathway overexpression[J]. Applied Microbiology and Biotechnology, 2011, 91(3): 577-589. |
| 97 | MATSUDA F, ISHII J, KONDO T, et al. Increased isobutanol production in Saccharomyces cerevisiae by eliminating competing pathways and resolving cofactor imbalance [J]. Microbial Cell Factories, 2013, 12: 119-129. |
| 98 | WANG J, LI C Y, ZOU Y S, et al. Bacterial synthesis of C3-C5 diols via extending amino acid catabolism[J]. Proceedings of the National Academy of Sciences of the United States of America, 2020, 117(32): 19159-19167. |
| 99 | CHAE T U, AHN J H, KO Y S, et al. Metabolic engineering for the production of dicarboxylic acids and diamines[J]. Metabolic Engineering, 2020, 58: 2-16. |
| 100 | CHAE T U, KO Y S, HWANG K S, et al. Metabolic engineering of Escherichia coli for the production of four-, five- and six-carbon lactams[J]. Metabolic Engineering, 2017, 41: 82-91. |
| 101 | YU J L, XIA X X, ZHONG J J, et al. A novel synthetic pathway for glutarate production in recombinant Escherichia coli [J]. Process Biochemistry, 2017, 59: 167-171. |
| 102 | YU J L, XIA X X, ZHONG J J, et al. Enhanced production of C5 dicarboxylic acids by aerobic-anaerobic shift in fermentation of engineered Escherichia coli [J]. Process Biochemistry, 2017, 62: 53-58. |
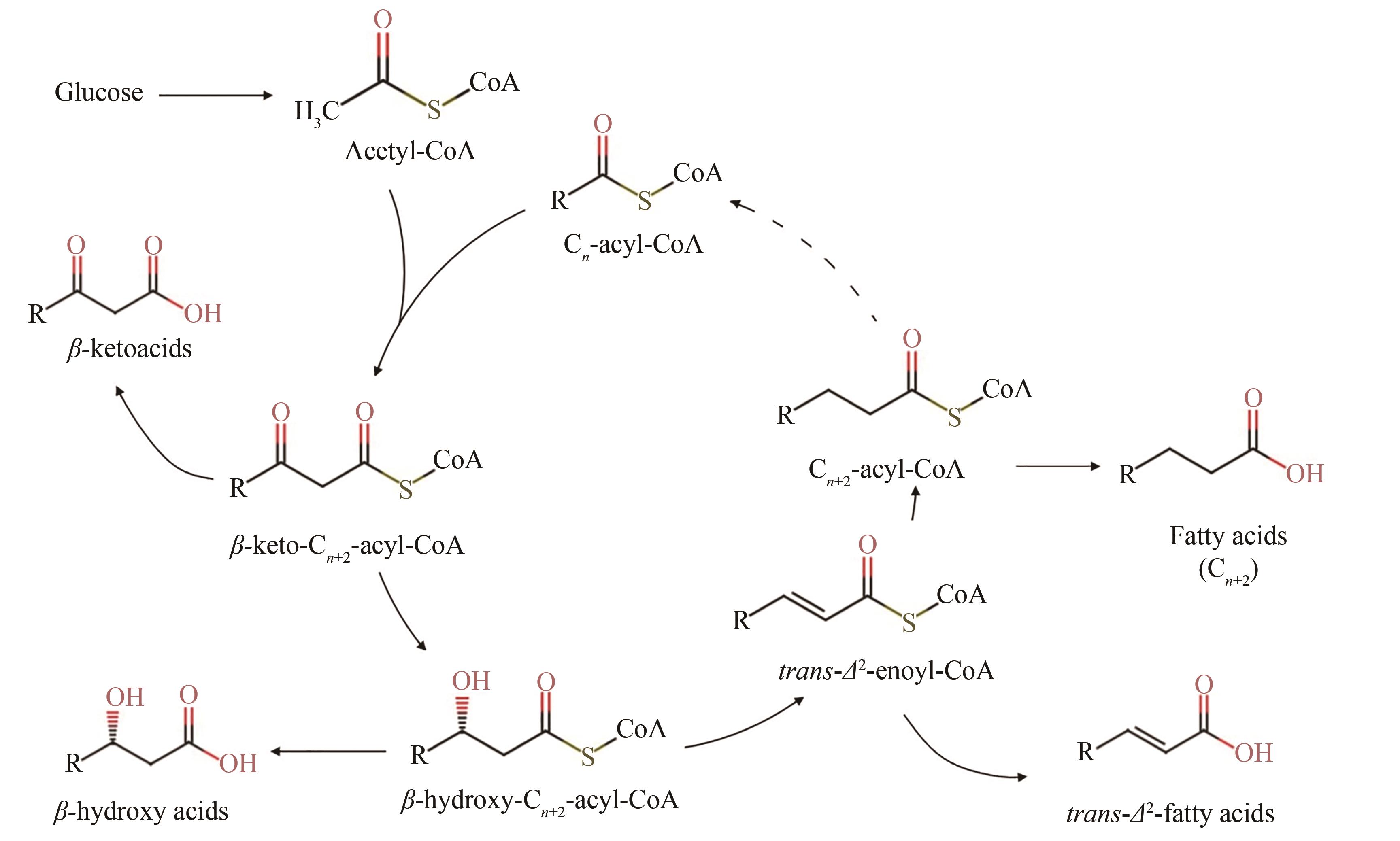
| 103 | SCHULZ H. Beta oxidation of fatty acids[J]. Biochimica et Biophysica Acta, 1991, 1081(2): 109-120. |
| 104 | KATIYAR S S, PORTER J W. Mechanism of fatty acid synthesis[J]. Life Sciences, 1977, 20(5): 737-759. |
| 105 | CLOMBURG J M, VICK J E, BLANKSCHIEN M D, et al. A synthetic biology approach to engineer a functional reversal of the β-oxidation cycle[J]. ACS Synthetic Biology, 2012, 1(11): 541-554. |
| 106 | GULEVICH A Y, SKOROKHODOVA A Y, SUKHOZHENKO A V, et al. Metabolic engineering of Escherichia coli for 1-butanol biosynthesis through the inverted aerobic fatty acid β-oxidation pathway[J]. Biotechnology Letters, 2012, 34(3): 463-469. |
| 107 | LIAN J Z, ZHAO H M. Reversal of the β-oxidation cycle in Saccharomyces cerevisiae for production of fuels and chemicals[J]. ACS Synthetic Biology, 2015, 4(3): 332-341. |
| 108 | ZHUANG Q Q, WANG Q, LIANG Q F, et al. Synthesis of polyhydroxyalkanoates from glucose that contain medium-chain-length monomers via the reversed fatty acid β-oxidation cycle in Escherichia coli [J]. Metabolic Engineering, 2014, 24: 78-86. |
| 109 | CINTOLESI A, CLOMBURG J M, GONZALEZ R. In silico assessment of the metabolic capabilities of an engineered functional reversal of the β-oxidation cycle for the synthesis of longer-chain (C≥4) products[J]. Metabolic Engineering, 2014, 23: 100-115. |
| 110 | DELLOMONACO C, CLOMBURG J M, MILLER E N, et al. Engineered reversal of the β-oxidation cycle for the synthesis of fuels and chemicals[J]. Nature, 2011, 476(7360): 355-359. |
| 111 | CLOMBURG J M, BLANKSCHIEN M D, VICK J E, et al. Integrated engineering of β-oxidation reversal and ω-oxidation pathways for the synthesis of medium chain ω-functionalized carboxylic acids[J]. Metabolic Engineering, 2015, 28: 202-212. |
| 112 | CHEONG S, CLOMBURG J M, GONZALEZ R. Energy-and carbon-efficient synthesis of functionalized small molecules in bacteria using non-decarboxylative Claisen condensation reactions[J]. Nature Biotechnology, 2016, 34(5): 556-561. |
| 113 | STEEN E J, KANG Y, BOKINSKY G, et al. Microbial production of fatty-acid-derived fuels and chemicals from plant biomass[J]. Nature, 2010, 463(7280): 559-562. |
| 114 | LENNEN R M, BRADEN D J, WEST R A, et al. A process for microbial hydrocarbon synthesis: overproduction of fatty acids in Escherichia coli and catalytic conversion to alkanes[J]. Biotechnology and Bioengineering, 2010, 106(2): 193-202. |
| 115 | CAI W L, ZHANG W J. Engineering modular polyketide synthases for production of biofuels and industrial chemicals[J]. Current Opinion in Biotechnology, 2018, 50: 32-38. |
| 116 | ZHANG Z, PAN H X, TANG G L. New insights into bacterial type II polyketide biosynthesis[J]. F1000Research, 2017, 6: 172. |
| 117 | SHIMIZU Y, OGATA H, GOTO S. Type III polyketide synthases: functional classification and phylogenomics[J]. ChemBioChem, 2017, 18(1): 50-65. |
| 118 | KHOSLA C, HERSCHLAG D, CANE D E, et al. Assembly line polyketide synthases: Mechanistic insights and unsolved problems[J]. Biochemistry, 2014, 53(18): 2875-2883. |
| 119 | ZHANG W J, LIU J. Recent advances in understanding and engineering polyketide synthesis[J]. F1000Research, 2016, 5: 208. |
| 120 | WEISSMAN K J. Genetic engineering of modular PKSs: from combinatorial biosynthesis to synthetic biology [J]. Natural Product Reports, 2016, 33(2): 203-230. |
| 121 | WONG F T, KHOSLA C. Combinatorial biosynthesis of polyketides-a perspective[J]. Current Opinion in Chemical Biology, 2012, 16(1/2): 117-123. |
| 122 | HERTWECK C. Decoding and reprogramming complex polyketide assembly lines: prospects for synthetic biology[J]. Trends in Biochemical Sciences, 2015, 40(4): 189-199. |
| 123 | KALKREUTER E, CROWETIPTON J M, LOWELL A N, et al. Engineering the substrate specificity of a modular polyketide synthase for installation of consecutive non-natural extender units[J]. Journal of the American Chemical Society, 2019, 141(5): 1961-1969. |
| 124 | ENG C H, YUZAWA S, WANG G, et al. Alteration of polyketide stereochemistry from anti to syn by a ketoreductase domain exchange in a type I modular polyketide synthase subunit[J]. Biochemistry, 2016, 55(12): 1677-1680. |
| 125 | ZARGAR A, LAL R, VALENCIA L, et al. Chemoinformatic-guided engineering of polyketide synthases[J]. Journal of the American Chemical Society, 2020, 142(22): 9896-9901. |
| 126 | LÜNNE F, NIEHAUS E M, LIPINSKI S, et al. Identification of the polyketide synthase PKS7 responsible for the production of lecanoric acid and ethyl lecanorate in Claviceps purpurea [J]. Fungal Genetics and Biology, 2020, 145: 103481. |
| 127 | HAGEN A, POUST S, ROND T D, et al. Engineering a polyketide synthase for in vitro production of adipic acid[J]. ACS Synthetic Biology, 2016, 5(1): 21-27. |
| 128 | FARRELL A E, PLEVIN R J, TURNER B T, et al. Ethanol can contribute to energy and environmental goals [J]. Science, 2006, 311(5760): 506-508. |
| [1] | GUO Shuyuan, ZHANG Qiannan, Gulikezi· MAIMAITIREXIATI, YANG Yiqun, YU Tao. Advances in microbial production of liquid biofuels [J]. Synthetic Biology Journal, 2025, 6(1): 18-44. |
| [2] | CHAI Meng, WANG Fengqing, WEI Dongzhi. Synthesis of organic acids from lignocellulose by biotransformation [J]. Synthetic Biology Journal, 2024, 5(6): 1242-1263. |
| [3] | SHAO Mingwei, SUN Simian, YANG Shimao, CHEN Guoqiang. Bioproduction based on extremophiles [J]. Synthetic Biology Journal, 2024, 5(6): 1419-1436. |
| [4] | ZHAO Liang, LI Zhenshuai, FU Liping, LYU Ming, WANG Shi’an, ZHANG Quan, LIU Licheng, LI Fuli, LIU Ziyong. Progress in biomanufacturing of lipids and single cell protein from one-carbon compounds [J]. Synthetic Biology Journal, 2024, 5(6): 1300-1318. |
| [5] | ZHU Fanghuan, CEN Xuecong, CHEN Zhen. Research progress of diols production by microbes [J]. Synthetic Biology Journal, 2024, 5(6): 1367-1385. |
| [6] | YU Wei, GAO Jiaoqi, ZHOU Yongjin. Bioconversion of one carbon feedstocks for producing organic acids [J]. Synthetic Biology Journal, 2024, 5(5): 1169-1188. |
| [7] | CHEN Xiwei, ZHANG Huaran, ZOU Yi. Biosynthesis and metabolic engineering of fungal non-ribosomal peptides [J]. Synthetic Biology Journal, 2024, 5(3): 571-592. |
| [8] | XIE Huang, ZHENG Yilei, SU Yiting, RUAN Jingyi, LI Yongquan. An overview on reconstructing the biosynthetic system of actinomycetes for polyketides production [J]. Synthetic Biology Journal, 2024, 5(3): 612-630. |
| [9] | ZHAO Jingyu, ZHANG Jian, QI Qingsheng, WANG Qian. Research progress in biosensors based on bacterial two-component systems [J]. Synthetic Biology Journal, 2024, 5(1): 38-52. |
| [10] | SUN Huili, CUI Jinyu, LUAN Guodong, LYU Xuefeng. Progress of cyanobacterial synthetic biotechnology for efficient light-driven carbon fixation and ethanol production [J]. Synthetic Biology Journal, 2023, 4(6): 1161-1177. |
| [11] | YAN Xiongying, WANG Zhen, LOU Jiyun, ZHANG Haoyu, HUANG Xingyu, WANG Xia, YANG Shihui. Progress in the construction of microbial cell factories for efficient biofuel production [J]. Synthetic Biology Journal, 2023, 4(6): 1082-1121. |
| [12] | XIAO Yan, LIU Yajun, FENG Yin′gang, CUI Qiu. Progress in synthetic biology research of Clostridium thermocellum for biomass energy applications [J]. Synthetic Biology Journal, 2023, 4(6): 1055-1081. |
| [13] | CHENG Zhenzhen, ZHANG Jian, GAO Cong, LIU Liming, CHEN Xiulai. Progress in metabolic engineering of microorganisms for the utilization of formate [J]. Synthetic Biology Journal, 2023, 4(4): 756-778. |
| [14] | LIU Jiayu, YANG Zhihan, YANG Lei, ZHU Liying, ZHU Zhengming, JIANG Ling. Advances in the development of Clostridium tyrobutyricum cell factories driven by synthetic biotechnology [J]. Synthetic Biology Journal, 2022, 3(6): 1174-1200. |
| [15] | GUO Shuyuan, WU Lianghuan, LIU Xiangjian, WANG Bo, YU Tao. Developing C1-based metabolic network in methylotrophy for biotransformation [J]. Synthetic Biology Journal, 2022, 3(1): 116-137. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||