Synthetic Biology Journal ›› 2023, Vol. 4 ›› Issue (6): 1281-1299.DOI: 10.12211/2096-8280.2023-056
• Invited Review • Previous Articles Next Articles
Advances and applications of gene editing and transcriptional regulation in electroactive microorganisms
CHEN Yaru1,2, CAO Yingxiu1,2, SONG Hao1,2
- 1.School of Chemical Engineering and Technology,Tianjin University,Tianjin 300072,China
2.Key Laboratory of Systems Bioengineering (Ministry of Education),Frontier Science Center for Synthetic Biology,Tianjin University,Tianjin 300072,China
-
Received:2023-08-19Revised:2023-09-18Online:2024-01-19Published:2023-12-31 -
Contact:CAO Yingxiu, SONG Hao
电活性微生物基因编辑与转录调控技术进展与应用
陈雅如1,2, 曹英秀1,2, 宋浩1,2
- 1.天津大学,化工学院,天津 300072
2.天津大学,合成生物学前沿科学中心,系统生物工程教育部重点实验室,天津 300072
-
通讯作者:曹英秀,宋浩 -
作者简介:陈雅如 (1995—),女,博士研究生。研究方向为电活性微生物,基因编辑与调控。E-mail:yaruchen2018207303@tju.edu.cn曹英秀 (1986—),女,副教授,博士生导师。研究方向为高性能生物燃料细胞工厂设计与重构。E-mail:caoyingxiu@tju.edu.cn宋浩 (1973—),男,教授,博士生导师。研究方向为电能细胞合成生物学,微生物光/电合成。E-mail:hsong@tju.edu.cn -
基金资助:国家重点研发计划(2018YFA0901300);国家自然科学基金(32071411)
CLC Number:
Cite this article
CHEN Yaru, CAO Yingxiu, SONG Hao. Advances and applications of gene editing and transcriptional regulation in electroactive microorganisms[J]. Synthetic Biology Journal, 2023, 4(6): 1281-1299.
陈雅如, 曹英秀, 宋浩. 电活性微生物基因编辑与转录调控技术进展与应用[J]. 合成生物学, 2023, 4(6): 1281-1299.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2023-056
| 1 | LOVLEY D R, HOLMES D E. Electromicrobiology: the ecophysiology of phylogenetically diverse electroactive microorganisms[J]. Nature Reviews Microbiology, 2022, 20(1): 5-19. |
| 2 | JIANG Y, ZENG R J. Bidirectional extracellular electron transfers of electrode-biofilm: mechanism and application[J]. Bioresource Technology, 2019, 271: 439-448. |
| 3 | LOGAN B E, ROSSI R, RAGAB A, et al. Electroactive microorganisms in bioelectrochemical systems[J]. Nature Reviews Microbiology, 2019, 17(5): 307-319. |
| 4 | CHU N, LIANG Q J, HAO W, et al. Microbial electrochemical sensor for water biotoxicity monitoring[J]. Chemical Engineering Journal, 2021, 404: 127053. |
| 5 | LIU C J, YU H, ZHANG B C, et al. Engineering whole-cell microbial biosensors: design principles and applications in monitoring and treatment of heavy metals and organic pollutants[J]. Biotechnology Advances, 2022, 60: 108019. |
| 6 | LÜ J, REN G P, HU Q C, et al. Microbial biofilm-based hydrovoltaic technology[J]. Trends in Biotechnology, 2023, 41(9): 1155-1167. |
| 7 | LOVLEY D R, HOLMES D E. Protein nanowires: the electrification of the microbial world and maybe our own[J]. Journal of Bacteriology, 2020, 202(20): e00331-20. |
| 8 | CHEN H, DONG F Y, MINTEER S D. The progress and outlook of bioelectrocatalysis for the production of chemicals, fuels and materials[J]. Nature Catalysis, 2020, 3(3): 225-244. |
| 9 | CESTELLOS-BLANCO S, ZHANG H, KIM J M, et al. Photosynthetic semiconductor biohybrids for solar-driven biocatalysis[J]. Nature Catalysis, 2020, 3(3): 245-255. |
| 10 | CAO B C, ZHAO Z P, PENG L L, et al. Silver nanoparticles boost charge-extraction efficiency in Shewanella microbial fuel cells[J]. Science, 2021, 373(6561): 1336-1340. |
| 11 | YU Y Y, WANG Y Z, FANG Z, et al. Single cell electron collectors for highly efficient wiring-up electronic abiotic/biotic interfaces[J]. Nature Communications, 2020, 11: 4087. |
| 12 | TABARES M, DULAY H, REGUERA G. Geobacter sulfurreducens[J]. Trends in Microbiology, 2020, 28(4): 327-328. |
| 13 | MENG F K, ELLIS T. The second decade of synthetic biology: 2010-2020[J]. Nature Communications, 2020, 11: 5174. |
| 14 | SHARAN S K, THOMASON L C, KUZNETSOV S G, et al. Recombineering: a homologous recombination-based method of genetic engineering[J]. Nature Protocols, 2009, 4(2): 206-223. |
| 15 | FAN Y Y, TANG Q A, LI Y, et al. Rapid and highly efficient genomic engineering with a novel iEditing device for programming versatile extracellular electron transfer of electroactive bacteria[J]. Environmental Microbiology, 2021, 23(2): 1238-1255. |
| 16 | COPPI M V, LEANG C, SANDLER S J, et al. Development of a genetic system for Geobacter sulfurreducens [J]. Applied and Environmental Microbiology, 2001, 67(7): 3180-3187. |
| 17 | CHEN W Z, ZHANG Y, ZHANG Y F, et al. CRISPR/Cas9-based genome editing in Pseudomonas aeruginosa and cytidine deaminase-mediated base editing in Pseudomonas species[J]. iScience, 2018, 6: 222-231. |
| 18 | LI Y, LI Y Y, CHEN Y R, et al. Coupling riboflavin de novo biosynthesis and cytochrome expression for improving extracellular electron transfer efficiency in Shewanella oneidensis [J]. Biotechnology and Bioengineering, 2022, 119(10): 2806-2818. |
| 19 | WU Z Y, HUANG Y T, CHAO W C, et al. Reversal of carbapenem-resistance in Shewanella algae by CRISPR/Cas9 genome editing[J]. Journal of Advanced Research, 2019, 18: 61-69. |
| 20 | LIU T, YU Y Y, DENG X P, et al. Enhanced Shewanella biofilm promotes bioelectricity generation[J]. Biotechnology and Bioengineering, 2015, 112(10): 2051-2059. |
| 21 | LI F, LI Y X, CAO Y X, et al. Modular engineering to increase intracellular NAD(H/+) promotes rate of extracellular electron transfer of Shewanella oneidensis [J]. Nature Communications, 2018, 9: 3637. |
| 22 | ZHENG W T, XIA Y D, WANG X E, et al. Precise genome engineering in Pseudomonas using phage-encoded homologous recombination and the Cascade-Cas3 system[J]. Nature Protocols, 2023, 18(9): 2642-2670. |
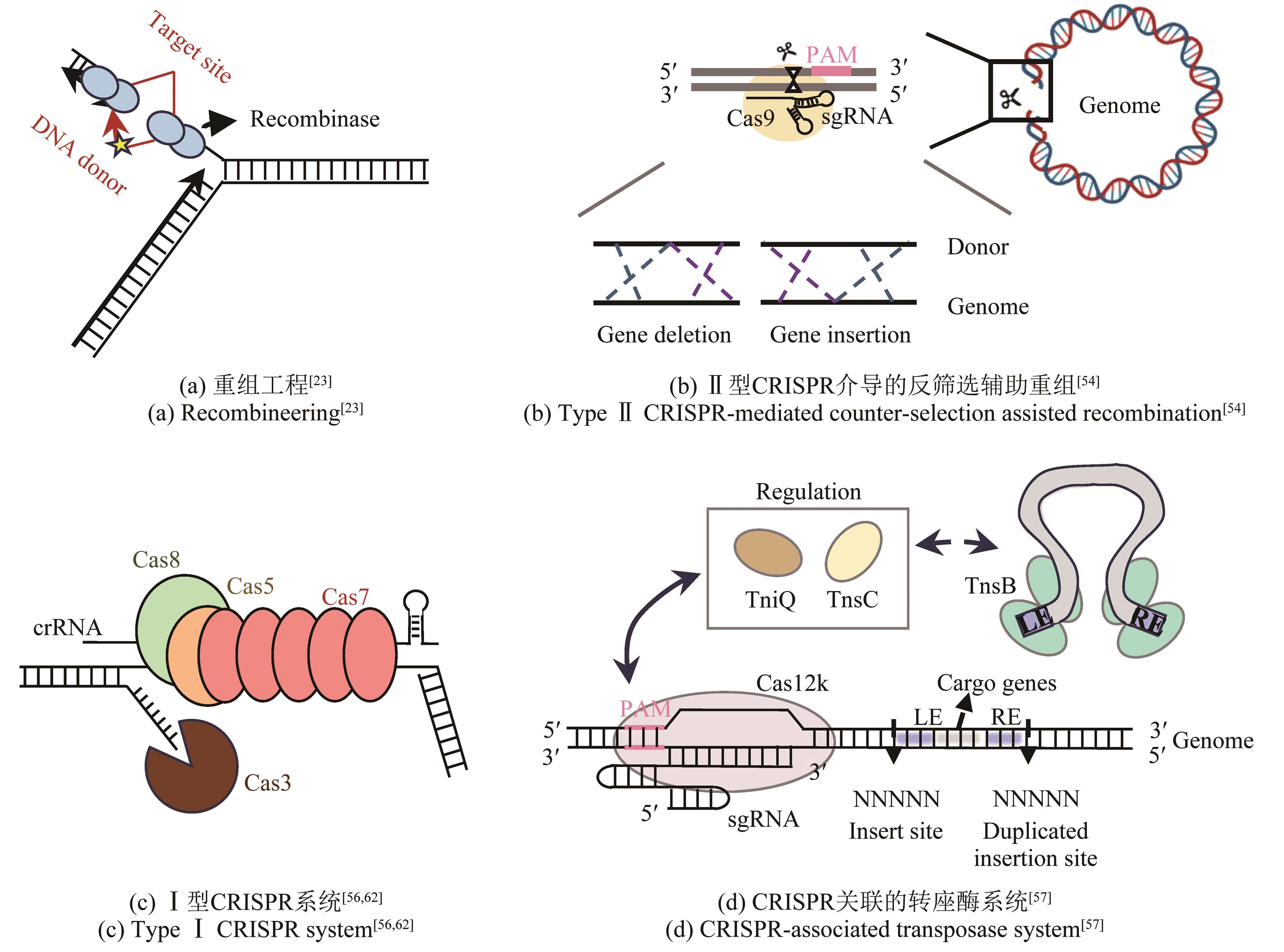
| 23 | CORTS A D, THOMASON L C, GILL R T, et al. A new recombineering system for precise genome-editing in Shewanella oneidensis strain MR-1 using single-stranded oligonucleotides[J]. Scientific Reports, 2019, 9: 39. |
| 24 | HMELO L R, BORLEE B R, ALMBLAD H, et al. Precision-engineering the Pseudomonas aeruginosa genome with two-step allelic exchange[J]. Nature Protocols, 2015, 10(11): 1820-1841. |
| 25 | THORMANN K M, SAVILLE R M, SHUKLA S, et al. Induction of rapid detachment in Shewanella oneidensis MR-1 biofilms[J]. Journal of Bacteriology, 2005, 187(3): 1014-1021. |
| 26 | CHAN C H, LEVAR C E, ZACHAROFF L, et al. Scarless genome editing and stable inducible expression vectors for Geobacter sulfurreducens [J]. Applied and Environmental Microbiology, 2015, 81(20): 7178-7186. |
| 27 | GRAF N, ALTENBUCHNER J. Development of a method for markerless gene deletion in Pseudomonas putida [J]. Applied and Environmental Microbiology, 2011, 77(15): 5549-5552. |
| 28 | PÓSFAI G, KOLISNYCHENKO V, BERECZKI Z, et al. Markerless gene replacement in Escherichia coli stimulated by a double-strand break in the chromosome[J]. Nucleic Acids Research, 1999, 27(22): 4409-4415. |
| 29 | MARTÍNEZ-GARCÍA E, DE LORENZO V. Engineering multiple genomic deletions in Gram-negative bacteria: analysis of the multi-resistant antibiotic profile of Pseudomonas putida KT2440[J]. Environmental Microbiology, 2011, 13(10): 2702-2716. |
| 30 | LUO X, YANG Y W, LING W, et al. Pseudomonas putida KT2440 markerless gene deletion using a combination of λ Red recombineering and Cre/loxP site-specific recombination[J]. FEMS Microbiology Letters, 2016, 363(4): fnw014. |
| 31 | VELÁZQUEZ E, AL-RAMAHI Y, TELLECHEA-LUZARDO J, et al. Targetron-assisted delivery of exogenous DNA sequences into Pseudomonas putida through CRISPR-aided counterselection[J]. ACS Synthetic Biology, 2021, 10(10): 2552-2565. |
| 32 | BIRD L J, KUNDU B B, TSCHIRHART T, et al. Engineering wired life: synthetic biology for electroactive bacteria[J]. ACS Synthetic Biology, 2021, 10(11): 2808-2823. |
| 33 | MURPHY K C. Use of bacteriophage λ recombination functions to promote gene replacement in Escherichia coli [J]. Journal of Bacteriology, 1998, 180(8): 2063-2071. |
| 34 | SAWITZKE J A, THOMASON L C, COSTANTINO N, et al. Recombineering: in vivo genetic engineering in E. coli, S. enterica, and beyond[J]. Methods in Enzymology, 2007, 421: 171-199. |
| 35 | PINES G, FREED E F, WINKLER J D, et al. Bacterial recombineering: genome engineering via phage-based homologous recombination[J]. ACS Synthetic Biology, 2015, 4(11): 1176-1185. |
| 36 | SWINGLE B, BAO Z M, MARKEL E, et al. Recombineering using RecTE from Pseudomonas syringae [J]. Applied and Environmental Microbiology, 2010, 76(15): 4960-4968. |
| 37 | BAO Z M, CARTINHOUR S, SWINGLE B. Substrate and target sequence length influence RecTEPsy recombineering efficiency in Pseudomonas syringae [J]. PLoS One, 2012, 7(11): e50617. |
| 38 | APARICIO T, JENSEN S I, NIELSEN A T, et al. The Ssr protein (T1E_1405) from Pseudomonas putida DOT-T1E enables oligonucleotide-based recombineering in platform strain P. putida EM42[J]. Biotechnology Journal, 2016, 11(10): 1309-1319. |
| 39 | WANG H H, ISAACS F J, CARR P A, et al. Programming cells by multiplex genome engineering and accelerated evolution[J]. Nature, 2009, 460(7257): 894-898. |
| 40 | CARROLL D. Genome engineering with targetable nucleases[J]. Annual Review of Biochemistry, 2014, 83: 409-439. |
| 41 | RAN F A, HSU P D, WRIGHT J, et al. Genome engineering using the CRISPR-Cas9 system[J]. Nature Protocols, 2013, 8(11): 2281-2308. |
| 42 | STERNBERG S H, REDDING S, JINEK M, et al. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9[J]. Nature, 2014, 507(7490): 62-67. |
| 43 | PETERS J M, COLAVIN A, SHI H D, et al. A comprehensive, CRISPR-based functional analysis of essential genes in bacteria[J]. Cell, 2016, 165(6): 1493-1506. |
| 44 | FAN Y Y, TANG Q A, LI F H, et al. Enhanced bioreduction of radionuclides by driving microbial extracellular electron pumping with an engineered CRISPR platform[J]. Environmental Science & Technology, 2021, 55(17): 11997-12008. |
| 45 | WIRTH N T, KOZAEVA E, NIKEL P I. Accelerated genome engineering of Pseudomonas putida by Ⅰ-SceⅠ-mediated recombination and CRISPR-Cas9 counterselection[J]. Microbial Biotechnology, 2020, 13(1): 233-249. |
| 46 | CORTS A D, THOMASON L C, GILL R T, et al. Efficient and precise genome editing in Shewanella with recombineering and CRISPR/Cas9-mediated counter-selection[J]. ACS Synthetic Biology, 2019, 8(8): 1877-1889. |
| 47 | COOK T B, RAND J M, NURANI W, et al. Genetic tools for reliable gene expression and recombineering in Pseudomonas putida [J]. Journal of Industrial Microbiology & Biotechnology, 2018, 45(7): 517-527. |
| 48 | WU Z X, CHEN Z Q, GAO X Y, et al. Combination of ssDNA recombineering and CRISPR-Cas9 for Pseudomonas putida KT2440 genome editing[J]. Applied Microbiology and Biotechnology, 2019, 103(6): 2783-2795. |
| 49 | ZHOU Y Y, LIN L, WANG H, et al. Development of a CRISPR/Cas9n-based tool for metabolic engineering of Pseudomonas putida for ferulic acid-to-polyhydroxyalkanoate bioconversion[J]. Communications Biology, 2020, 3: 98. |
| 50 | SUN J, WANG Q Z, JIANG Y, et al. Genome editing and transcriptional repression in Pseudomonas putida KT2440 via the type Ⅱ CRISPR system[J]. Microbial Cell Factories, 2018, 17(1): 41. |
| 51 | GARST A D, BASSALO M C, PINES G, et al. Genome-wide mapping of mutations at single-nucleotide resolution for protein, metabolic and genome engineering[J]. Nature Biotechnology, 2017, 35(1): 48-55. |
| 52 | PETERS J E, MAKAROVA K S, SHMAKOV S, et al. Recruitment of CRISPR-Cas systems by Tn7-like transposons[J]. Proceedings of the National Academy of Sciences of the United States of America, 2017, 114(35): E7358-E7366. |
| 53 | ASIN-GARCIA E, MARTIN-PASCUAL M, GARCIA-MORALES L, et al. ReScribe: an unrestrained tool combining multiplex recombineering and minimal-PAM ScCas9 for genome recoding Pseudomonas putida [J]. ACS Synthetic Biology, 2021, 10(10): 2672-2688. |
| 54 | CHEN Y R, CHENG M J, FENG X R, et al. Genome editing by CRISPR/Cas12 recognizing AT-rich PAMs in Shewanella oneidensis MR-1[J]. ACS Synthetic Biology, 2022, 11(9): 2947-2955. |
| 55 | LIN Z L, LI H H, HE L, et al. Efficient genome editing for Pseudomonas aeruginosa using CRISPR-Cas12a[J]. Gene, 2021, 790: 145693. |
| 56 | CSÖRGŐ B, LEÓN L M, CHAU-LY I J, et al. A compact Cascade-Cas3 system for targeted genome engineering[J]. Nature Methods, 2020, 17(12): 1183-1190. |
| 57 | CHENG Z H, WU J, LIU J Q, et al. Repurposing CRISPR RNA-guided integrases system for one-step, efficient genomic integration of ultra-long DNA sequences[J]. Nucleic Acids Research, 2022, 50(13): 7739-7750. |
| 58 | MAKAROVA K S, WOLF Y I, IRANZO J, et al. Evolutionary classification of CRISPR-Cas systems: a burst of class 2 and derived variants[J]. Nature Reviews Microbiology, 2020, 18(2): 67-83. |
| 59 | XU Z L, LI M, LI Y R, et al. Native CRISPR-cas-mediated genome editing enables dissecting and sensitizing clinical multidrug-resistant P. aeruginosa [J]. Cell Reports, 2019, 29(6): 1707-1717.e3. |
| 60 | DWARAKANATH S, BRENZINGER S, GLEDITZSCH D, et al. Interference activity of a minimal TypeⅠCRISPR-Cas system from Shewanella putrefaciens [J]. Nucleic Acids Research, 2015, 43(18): 8913-8923. |
| 61 | CHEN Y R, CHENG M J, SONG H, et al. Type Ⅰ-F CRISPR-PAIR platform for multi-mode regulation to boost extracellular electron transfer in Shewanella oneidensis [J]. iScience, 2022, 25(6): 104491. |
| 62 | CHEN Y X, LIU J Q, ZHI S Y, et al. Repurposing type Ⅰ-F CRISPR-Cas system as a transcriptional activation tool in human cells[J]. Nature Communications, 2020, 11: 3136. |
| 63 | XU Z L, LI Y R, CAO H L, et al. A transferrable and integrative type Ⅰ-F Cascade for heterologous genome editing and transcription modulation[J]. Nucleic Acids Research, 2021, 49(16): e94. |
| 64 | KLOMPE S E, VO P L H, HALPIN-HEALY T S, et al. Transposon-encoded CRISPR-Cas systems direct RNA-guided DNA integration[J]. Nature, 2019, 571(7764): 219-225. |
| 65 | STRECKER J, LADHA A, GARDNER Z, et al. RNA-guided DNA insertion with CRISPR-associated transposases[J]. Science, 2019, 365(6448): 48-53. |
| 66 | WANG Y, LIU Y, ZHENG P, et al. Microbial base editing: a powerful emerging technology for microbial genome engineering[J]. Trends in Biotechnology, 2021, 39(2): 165-180. |
| 67 | CHENG L, MIN D, HE R L, et al. Developing a base-editing system to expand the carbon source utilization spectra of Shewanella oneidensis MR-1 for enhanced pollutant degradation[J]. Biotechnology and Bioengineering, 2020, 117(8): 2389-2400. |
| 68 | ABDULLAH, WANG P J, HAN T R, et al. Adenine base editing system for Pseudomonas and prediction workflow for protein dysfunction via ABE[J]. ACS Synthetic Biology, 2022, 11(4): 1650-1657. |
| 69 | MOLLA K A, YANG Y N. CRISPR/cas-mediated base editing: technical considerations and practical applications[J]. Trends in Biotechnology, 2019, 37(10): 1121-1142. |
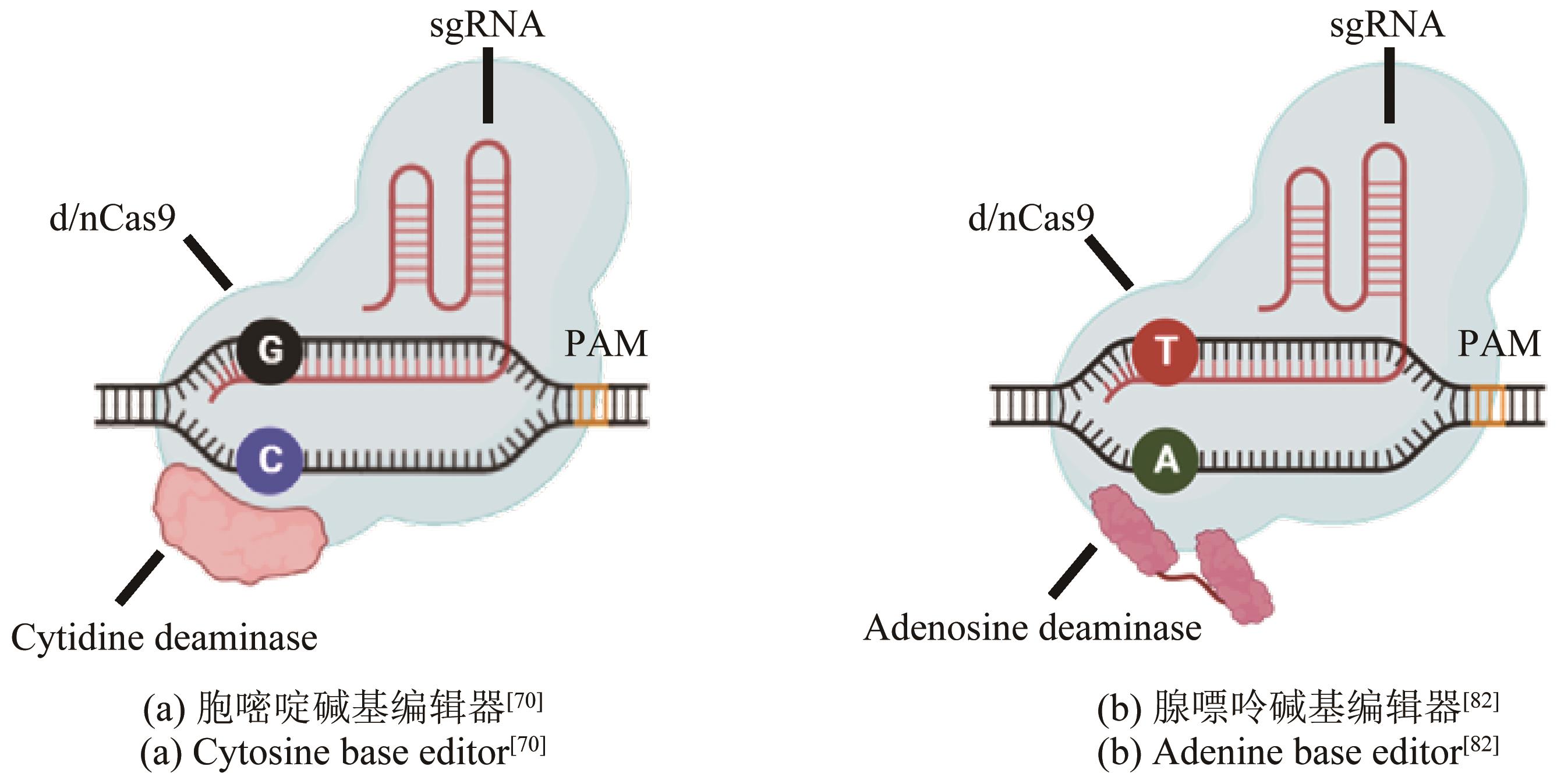
| 70 | KOMOR A C, KIM Y B, PACKER M S, et al. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage[J]. Nature, 2016, 533(7603): 420-424. |
| 71 | NISHIDA K, ARAZOE T, YACHIE N, et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems[J]. Science, 2016, 353(6305): aaf8729. |
| 72 | KUSCU C, PARLAK M, TUFAN T R, et al. CRISPR-STOP: gene silencing through base-editing-induced nonsense mutations[J]. Nature Methods, 2017, 14(7): 710-712. |
| 73 | WANG Y, LIU Y, LIU J, et al. MACBETH: multiplex automated Corynebacterium glutamicum base editing method[J]. Metabolic Engineering, 2018, 47: 200-210. |
| 74 | CHEN Y R, FANG L X, YING X A, et al. Development of whole genome-scale base editing toolbox to promote efficiency of extracellular electron transfer in Shewanella oneidensis MR-1[J]. Advanced Biology, 2022, 6(3): 2101296. |
| 75 | HE R L, WU J E, CHENG Z H, et al. Biomolecular insights into extracellular pollutant reduction pathways of Geobacter sulfurreducens using a base editor system[J]. Environmental Science & Technology, 2022, 56(17): 12247-12256. |
| 76 | YUE S J, HUANG P, LI S, et al. Developing a CRISPR-assisted base-editing system for genome engineering of Pseudomonas chlororaphis [J]. Microbial Biotechnology, 2022, 15(9): 2324-2336. |
| 77 | SUN J, LU L B, LIANG T X, et al. CRISPR-assisted multiplex base editing system in Pseudomonas putida KT2440[J]. Frontiers in Bioengineering and Biotechnology, 2020, 8: 905. |
| 78 | WEGRZYN R D, LEE A H, JENKINS A L, et al. Genome editing: insights from chemical biology to support safe and transformative therapeutic applications[J]. ACS Chemical Biology, 2018, 13(2): 333-342. |
| 79 | JAIN S, XUN G H, ABESTEH S, et al. Precise regulation of Cas9-mediated genome engineering by anti-CRISPR-based inducible CRISPR controllers[J]. ACS Synthetic Biology, 2021, 10(6): 1320-1327. |
| 80 | SAITO M, XU P Y, FAURE G, et al. Fanzor is a eukaryotic programmable RNA-guided endonuclease[J]. Nature, 2023, 620(7974): 660-668. |
| 81 | MATSOUKAS I G. Commentary: programmable base editing of A·T to G·C in genomic DNA without DNA cleavage[J]. Frontiers in Genetics, 2018, 9: 21. |
| 82 | GAUDELLI N M, KOMOR A C, REES H A, et al. Programmable base editing of A·T to G·C in genomic DNA without DNA cleavage[J]. Nature, 2017, 551(7681): 464-471. |
| 83 | WANG X J, LIU Z W, LI G L, et al. Efficient gene silencing by adenine base editor-mediated start codon mutation[J]. Molecular Therapy, 2020, 28(2): 431-440. |
| 84 | ZHANG Y, ZHANG H Y, WANG Z P, et al. Programmable adenine deamination in bacteria using a Cas9-adenine-deaminase fusion[J]. Chemical Science, 2020, 11(6): 1657-1664. |
| 85 | WANG T L, ZHANG J W, WEI L, et al. Developing a PAM-flexible CRISPR-mediated dual-deaminase base editor to regulate extracellular electron transport in Shewanella oneidensis [J]. ACS Synthetic Biology, 2023, 12(6): 1727-1738. |
| 86 | LI C, ZHANG R, MENG X B, et al. Targeted, random mutagenesis of plant genes with dual cytosine and adenine base editors[J]. Nature Biotechnology, 2020, 38(7): 875-882. |
| 87 | APARICIO T, DE LORENZO V, MARTÍNEZ-GARCÍA E. CRISPR/Cas9-enhanced ssDNA recombineering for Pseudomonas putida [J]. Microbial Biotechnology, 2019, 12(5): 1076-1089. |
| 88 | MCCARTY N S, GRAHAM A E, STUDENÁ L, et al. Multiplexed CRISPR technologies for gene editing and transcriptional regulation[J]. Nature Communications, 2020, 11: 1281. |
| 89 | CHEN Y R, CHENG M J, LI Y, et al. Highly efficient multiplex base editing: one-shot deactivation of eight genes in Shewanella oneidensis MR-1[J]. Synthetic and Systems Biotechnology, 2023, 8(1): 1-10. |
| 90 | VOLKE D C, MARTINO R A, KOZAEVA E, et al. Modular (de)construction of complex bacterial phenotypes by CRISPR/nCas9-assisted, multiplex cytidine base-editing[J]. Nature Communications, 2022, 13: 3026. |
| 91 | WU J, CHENG Z H, MIN D, et al. CRISPRi system as an efficient, simple platform for rapid identification of genes involved in pollutant transformation by Aeromonas hydrophila [J]. Environmental Science & Technology, 2020, 54(6): 3306-3315. |
| 92 | CAO Y X, LI X F, LI F, et al. CRISPRi-sRNA: transcriptional-translational regulation of extracellular electron transfer in Shewanella oneidensis [J]. ACS Synthetic Biology, 2017, 6(9): 1679-1690. |
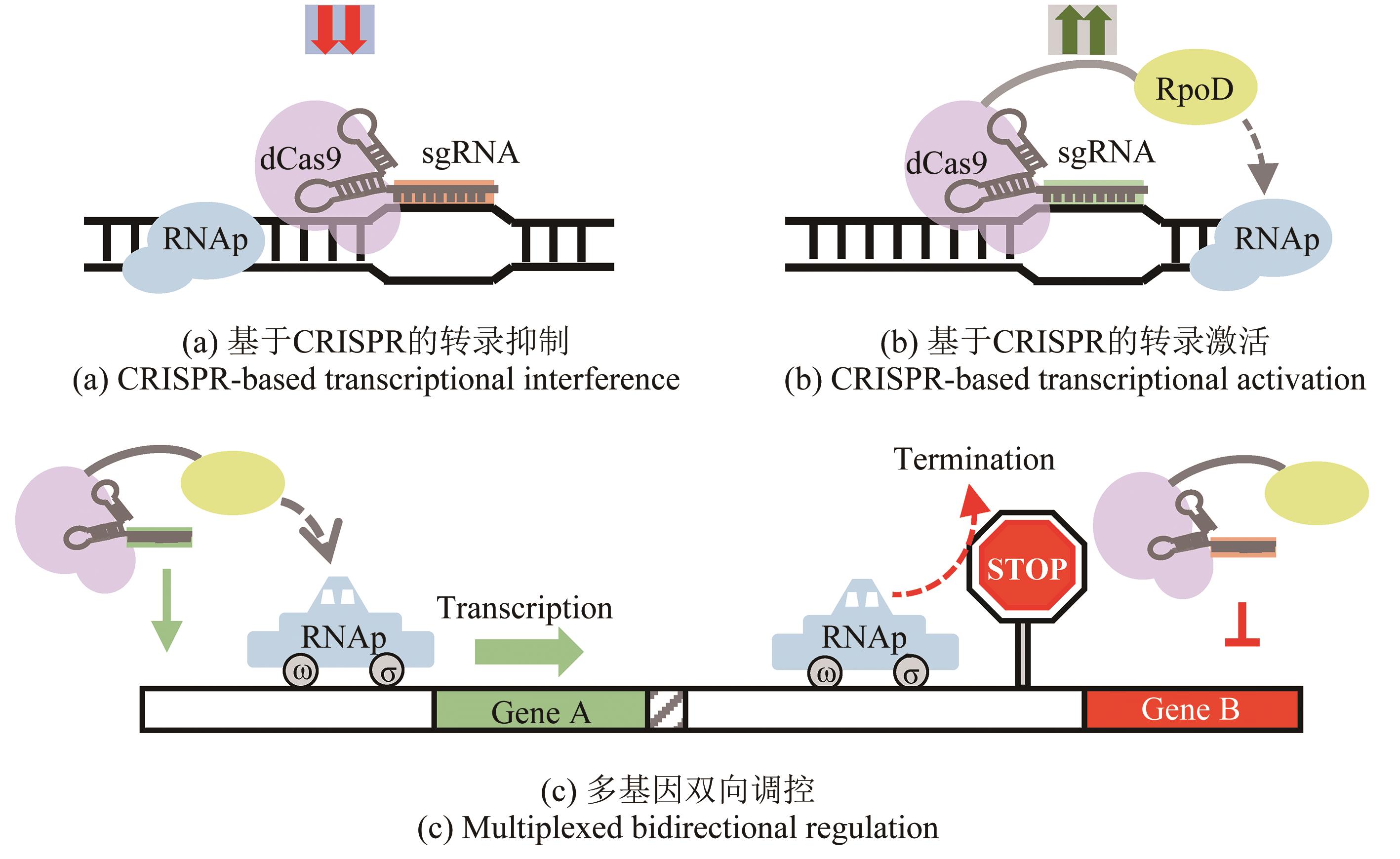
| 93 | CHEN Y R, NIU X L, CHENG M J, et al. CRISPR/dCas9-RpoD-mediated simultaneous transcriptional activation and repression in Shewanella oneidensis MR-1[J]. ACS Synthetic Biology, 2022, 11(6): 2184-2192. |
| 94 | LARSON M H, GILBERT L A, WANG X W, et al. CRISPR interference (CRISPRi) for sequence-specific control of gene expression[J]. Nature Protocols, 2013, 8(11): 2180-2196. |
| 95 | LI J E, TANG Q A, LI Y, et al. Rediverting electron flux with an engineered CRISPR-ddAsCpf1 system to enhance the pollutant degradation capacity of Shewanella oneidensis [J]. Environmental Science & Technology, 2020, 54(6): 3599-3608. |
| 96 | QI L S, LARSON M H, GILBERT L A, et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression[J]. Cell, 2013, 152(5): 1173-1183. |
| 97 | LIU Y, WAN X Y, WANG B J. Engineered CRISPRa enables programmable eukaryote-like gene activation in bacteria[J]. Nature Communications, 2019, 10: 3693. |
| 98 | BROWNING D F, BUSBY S J W. Local and global regulation of transcription initiation in bacteria[J]. Nature Reviews Microbiology, 2016, 14(10): 638-650. |
| 99 | TANENBAUM M E, GILBERT L A, QI L S, et al. A protein-tagging system for signal amplification in gene expression and fluorescence imaging[J]. Cell, 2014, 159(3): 635-646. |
| 100 | HO H I, FANG J R, CHEUNG J, et al. Programmable CRISPR-Cas transcriptional activation in bacteria[J]. Molecular Systems Biology, 2020, 16(7): e9427. |
| 101 | CZAJKA J J, BANERJEE D, ENG T, et al. Tuning a high performing multiplexed-CRISPRi Pseudomonas putida strain to further enhance indigoidine production[J]. Metabolic Engineering Communications, 2022, 15: e00206. |
| 102 | LI F, TANG R, ZHANG B C, et al. Systematic full-cycle engineering microbial biofilms to boost electricity production in Shewanella oneidensis [J]. Research, 2023, 6: 81. |
| 103 | BHOKISHAM N, VANARSDALE E, STEPHENS K T, et al. A redox-based electrogenetic CRISPR system to connect with and control biological information networks[J]. Nature Communications, 2020, 11: 2427. |
| 104 | SCHUETZ B, SCHICKLBERGER M, KUERMANN J, et al. Periplasmic electron transfer via the c-type cytochromes MtrA and FccA of Shewanella oneidensis MR-1[J]. Applied and Environmental Microbiology, 2009, 75(24): 7789-7796. |
| 105 | YU H, LU Y J, LAN F, et al. Engineering outer membrane vesicles to increase extracellular electron transfer of Shewanella oneidensis [J]. ACS Synthetic Biology, 2023, 12(6): 1645-1656. |
| 106 | SHI L, DONG H L, REGUERA G, et al. Extracellular electron transfer mechanisms between microorganisms and minerals[J]. Nature Reviews Microbiology, 2016, 14(10): 651-662. |
| 107 | THANASSI D G, BLISKA J B, CHRISTIE P J. Surface organelles assembled by secretion systems of Gram-negative bacteria: diversity in structure and function[J]. FEMS Microbiology Reviews, 2012, 36(6): 1046-1082. |
| 108 | WANG F B, GU Y Q, O'BRIEN J P, et al. Structure of microbial nanowires reveals stacked hemes that transport electrons over micrometers[J]. Cell, 2019, 177(2): 361-369.e10. |
| 109 | YALCIN S E, MALVANKAR N S. The blind men and the filament: understanding structures and functions of microbial nanowires[J]. Current Opinion in Chemical Biology, 2020, 59: 193-201. |
| 110 | LOVLEY D R, WALKER D J F. Geobacter protein nanowires[J]. Frontiers in Microbiology, 2019, 10: 2078. |
| 111 | LEANG C, MALVANKAR N S, FRANKS A E, et al. Engineering Geobacter sulfurreducens to produce a highly cohesive conductive matrix with enhanced capacity for current production[J]. Energy & Environmental Science, 2013, 6(6): 1901-1908. |
| 112 | PIRBADIAN S, BARCHINGER S E, LEUNG K M, et al. Shewanella oneidensis MR-1 nanowires are outer membrane and periplasmic extensions of the extracellular electron transport components[J]. Proceedings of the National Academy of Sciences of the United States of America, 2014, 111(35): 12883-12888. |
| 113 | CATANIA C, KARBELKAR A A, FURST A L. Engineering the interface between electroactive bacteria and electrodes[J]. Joule, 2021, 5(4): 743-747. |
| 114 | HEIJNE A TER, PEREIRA M A, PEREIRA J, et al. Electron storage in electroactive biofilms[J]. Trends in Biotechnology, 2021, 39(1): 34-42. |
| 115 | STURM G, RICHTER K, DOETSCH A, et al. A dynamic periplasmic electron transfer network enables respiratory flexibility beyond a thermodynamic regulatory regime[J]. The ISME Journal, 2015, 9(8): 1802-1811. |
| 116 | FANG L X, LI Y Y, LI Y, et al. Transcriptome analysis to identify crucial genes for reinforcing flavins-mediated extracellular electron transfer in Shewanella oneidensis [J]. Frontiers in Microbiology, 2022, 13: 852527. |
| 117 | MIN D, CHENG L, ZHANG F, et al. Enhancing extracellular electron transfer of Shewanella oneidensis MR-1 through coupling improved flavin synthesis and metal-reducing conduit for pollutant degradation[J]. Environmental Science & Technology, 2017, 51(9): 5082-5089. |
| 118 | YI Y C, NG I S. Redirection of metabolic flux in Shewanella oneidensis MR-1 by CRISPRi and modular design for 5-aminolevulinic acid production[J]. Bioresources and Bioprocessing, 2021, 8: 13. |
| 119 | SCHUSTER A, ERASIMUS H, FRITAH S, et al. RNAi/CRISPR screens: from a pool to a valid hit[J]. Trends in Biotechnology, 2019, 37(1): 38-55. |
| 120 | ALPER H, MOXLEY J, NEVOIGT E, et al. Engineering yeast transcription machinery for improved ethanol tolerance and production[J]. Science, 2006, 314(5805): 1565-1568. |
| 121 | WARNER J R, REEDER P J, KARIMPOUR-FARD A, et al. Rapid profiling of a microbial genome using mixtures of barcoded oligonucleotides[J]. Nature Biotechnology, 2010, 28(8): 856-862. |
| 122 | BAO Z H, HAMEDIRAD M, XUE P, et al. Genome-scale engineering of Saccharomyces cerevisiae with single-nucleotide precision[J]. Nature Biotechnology, 2018, 36(6): 505-508. |
| 123 | YAO L, SHABESTARY K, BJÖRK S M, et al. Pooled CRISPRi screening of the cyanobacterium Synechocystis sp. PCC 6803 for enhanced industrial phenotypes[J]. Nature Communications, 2020, 11: 1666. |
| 124 | WANG T M, GUAN C G, GUO J H, et al. Pooled CRISPR interference screening enables genome-scale functional genomics study in bacteria with superior performance[J]. Nature Communications, 2018, 9: 2475. |
| 125 | LIAN J Z, SCHULTZ C, CAO M F, et al. Multi-functional genome-wide CRISPR system for high throughput genotype-phenotype mapping[J]. Nature Communications, 2019, 10: 5794. |
| 126 | XIAO X, LIU Q Y, LI T T, et al. A high-throughput dye-reducing photometric assay for evaluating microbial exoelectrogenic ability[J]. Bioresource Technology, 2017, 241: 743-749. |
| 127 | YANG Z C, CHENG Y Y, ZHANG F, et al. Rapid detection and enumeration of exoelectrogenic bacteria in lake sediments and a wastewater treatment plant using a coupled WO3 nanoclusters and most probable number method[J]. Environmental Science & Technology Letters, 2016, 3(4): 133-137. |
| [1] | DONG Ying, MA Mengdan, HUANG Weiren. Progress in the miniaturization of CRISPR-Cas systems [J]. Synthetic Biology Journal, 2025, 6(1): 105-117. |
| [2] | CHEN Ziling, XIANG Yangfei. Integrated development of organoid technology and synthetic biology [J]. Synthetic Biology Journal, 2024, 5(4): 795-812. |
| [3] | CHEN Yingying, LIU Yang, SHI Junjie, MA Junying, JU Jianhua. CRISPR/Cas systems and their applications in gene editing with filamentous fungi [J]. Synthetic Biology Journal, 2024, 5(3): 672-693. |
| [4] | HUI Zhen, TANG Xiaoyu. Applications of the CRISPR/Cas9 editing system in the study of microbial natural products [J]. Synthetic Biology Journal, 2024, 5(3): 658-671. |
| [5] | DU Yao, GAO Hongdan, LIU Jiakun, LIU Xiaorong, XING Zhihao, ZHANG Tao, MA Dongli. Research progress of the CRISPR-Cas system in the detecting pathogen nucleic acids [J]. Synthetic Biology Journal, 2024, 5(1): 202-216. |
| [6] | XU Zhimeng, XIE Zhen. Research progress and biotechnological applications of the prime editing [J]. Synthetic Biology Journal, 2024, 5(1): 1-15. |
| [7] | ZHU Huawei, LI Yin. Biophotovoltaics: an environmentally friendly technology for solar energy utilization [J]. Synthetic Biology Journal, 2023, 4(6): 1259-1280. |
| [8] | Mengdan MA, Mengyu SHANG, Yuchen LIU. Application and prospect of CRISPR-Cas9 system in tumor biology [J]. Synthetic Biology Journal, 2023, 4(4): 703-719. |
| [9] | Tiantian WANG, Hong ZHU, Chen YANG. Development of CRISPRa for metabolic engineering applications in cyanobacteria [J]. Synthetic Biology Journal, 2023, 4(4): 824-839. |
| [10] | Jicong LIN, Gen ZOU, Hongmin LIU, Yongjun WEI. Application of CRISPR/Cas genome editing technology in the synthesis of secondary metabolites of filamentous fungi [J]. Synthetic Biology Journal, 2023, 4(4): 738-755. |
| [11] | Hailong LV, Jian WANG, Hao LV, Jin WANG, Yong XU, Dayong GU. Synthetic biology for next-generation genetic diagnostics [J]. Synthetic Biology Journal, 2023, 4(2): 318-332. |
| [12] | Mengdan MA, Yuchen LIU. Potential application of synthetic biology in disease information recording and real-time monitoring [J]. Synthetic Biology Journal, 2023, 4(2): 301-317. |
| [13] | Ke LIU, Guihong LIN, Kun LIU, Wei ZHOU, Fengqing WANG, Dongzhi WEI. Mining, engineering and functional expansion of CRISPR/Cas systems [J]. Synthetic Biology Journal, 2023, 4(1): 47-66. |
| [14] | Yingjia PAN, Siyang XIA, Chang DONG, Jin CAI, Jiazhang LIAN. Mutator-driven continuous genome evolution of Saccharomyces cerevisiae [J]. Synthetic Biology Journal, 2023, 4(1): 225-240. |
| [15] | Xiaolong TENG, Shuobo SHI. Optimization and development of CRISPR/Cas9 systems for genome editing [J]. Synthetic Biology Journal, 2023, 4(1): 67-85. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||