Synthetic Biology Journal ›› 2022, Vol. 3 ›› Issue (3): 567-586.DOI: 10.12211/2096-8280.2021-013
• Invited Review • Previous Articles Next Articles
Assessment on the pre-reaction state of enzyme: could we understand catalytic activity with near transition-state molecular dynamic simulation?-a review
SIM Byuri, ZHAO Yilei
- State Key Laboratory of Microbial Metabolism,School of Life Sciences and Biotechnology,Shanghai Jiao Tong University,Shanghai 200240,China
-
Received:2021-01-27Revised:2021-02-03Online:2022-07-13Published:2022-06-30 -
Contact:ZHAO Yilei
预反应态模型浅析:催化活性和近过渡态分子模拟
SIM Byuri, 赵一雷
- 上海交通大学生命科学技术学院,微生物代谢国家重点实验室,上海 200240
-
通讯作者:赵一雷 -
作者简介:SIM Byuri (1990—),男,硕士研究生。研究方向为醇脱氢酶CpRCR的共进化突变效应。E-mail:byurisim@sjtu.edu.cn赵一雷 (1972—),男,教授,博士生导师。研究方向为酶催化反应分子机制、蛋白质与核酸化学修饰,计算化学与分子生物信息学在蛋白质工程和生物医学中应用。E-mail:yileizhao@sjtu.edu.cn -
基金资助:国家自然科学基金(31970041);国家重点研发计划(2020YFA0907700)
CLC Number:
Cite this article
SIM Byuri, ZHAO Yilei. Assessment on the pre-reaction state of enzyme: could we understand catalytic activity with near transition-state molecular dynamic simulation?-a review[J]. Synthetic Biology Journal, 2022, 3(3): 567-586.
SIM Byuri, 赵一雷. 预反应态模型浅析:催化活性和近过渡态分子模拟[J]. 合成生物学, 2022, 3(3): 567-586.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://synbioj.cip.com.cn/EN/10.12211/2096-8280.2021-013
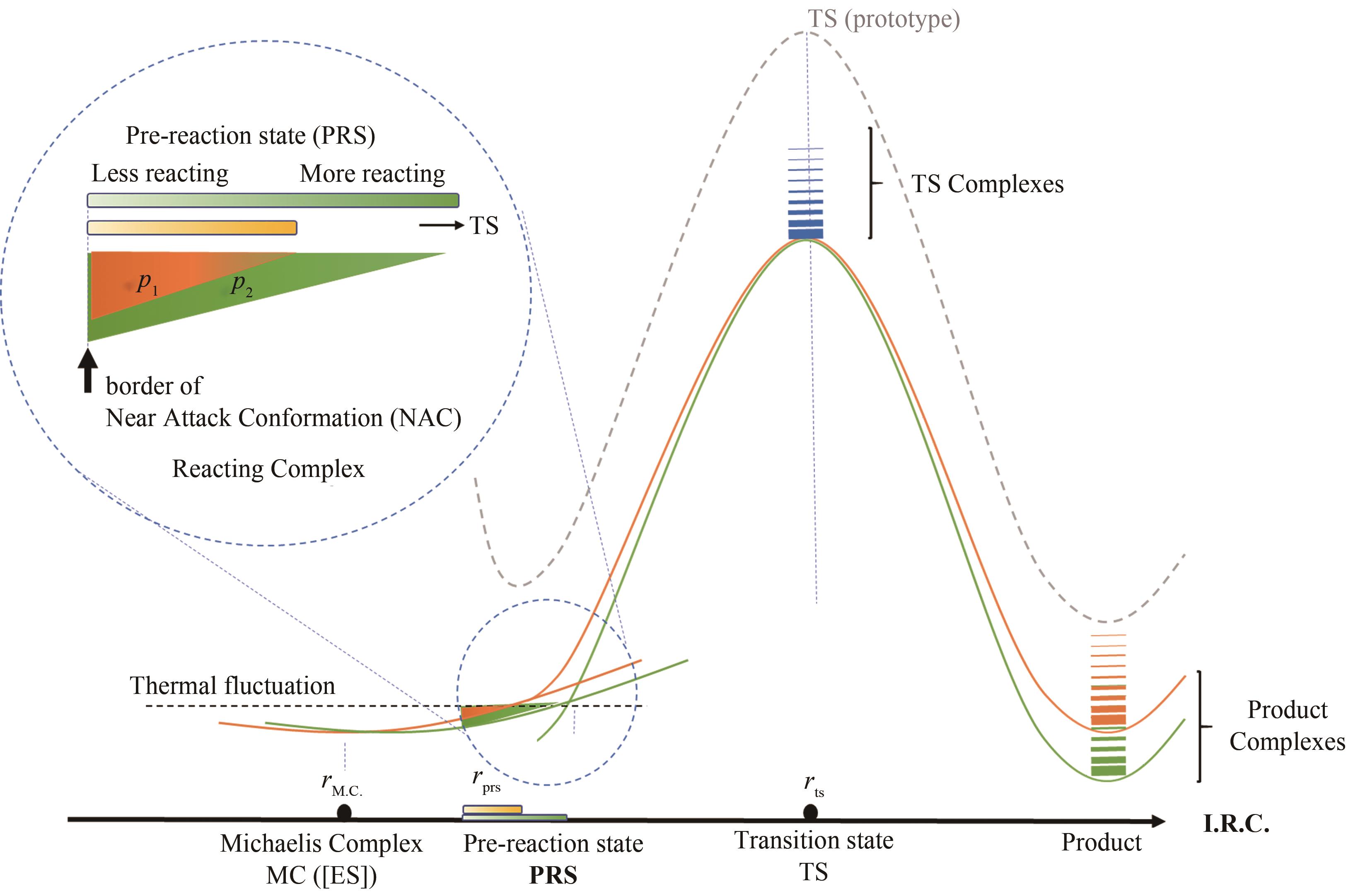
Fig. 2 Diagram for the perturbation of energy profile caused by the mutation of catalytic elements(In the simplest Michaelis's model, pre-reaction state and near attack conformation are identical, using the correlation of active conformation population and enzyme proficiency.)

Fig. 3 Correlation of the turn-over frequency (TOF) and reaction potential energy surface(In multiple steps of a catalytic cycle, pre-reaction state is defined as the active conformation near rate-determining transition state, in which rate is dependent on the protein engineering target such as mutation effect and substrate diversity.)
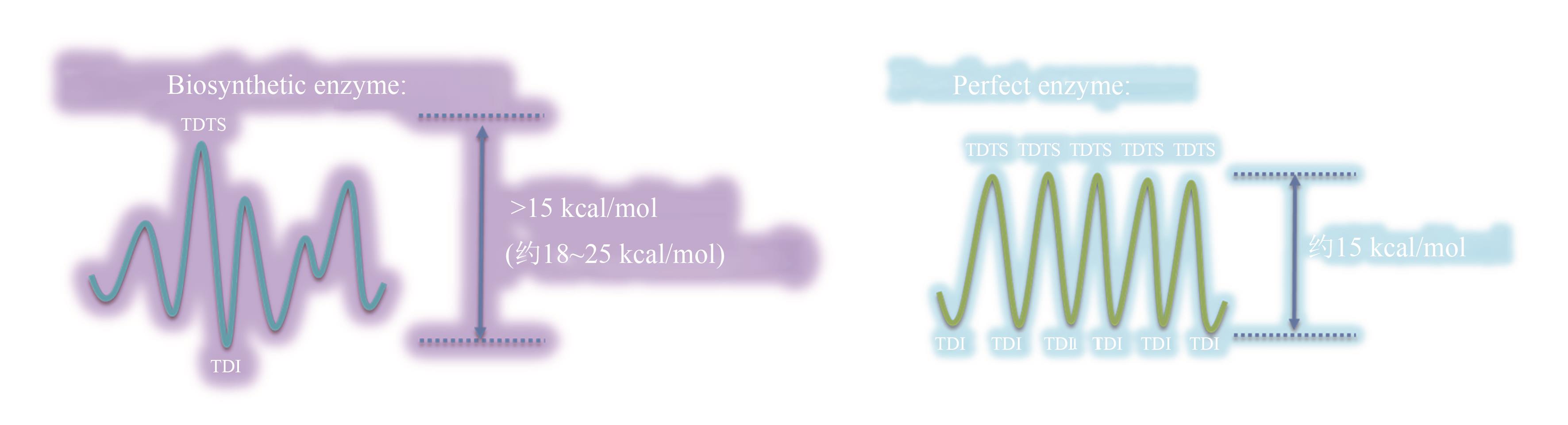
Fig. 4 Imaginary reaction potential energy surface for molecular evolution(Before evolving to a "perfect" enzyme, one point on the reaction potential energy surface controls the overall rate, but many other points are equivalently important for the perfect enzyme.)
| 研究对象 | 预反应态所选择的控制点 | 实例 |
|---|---|---|
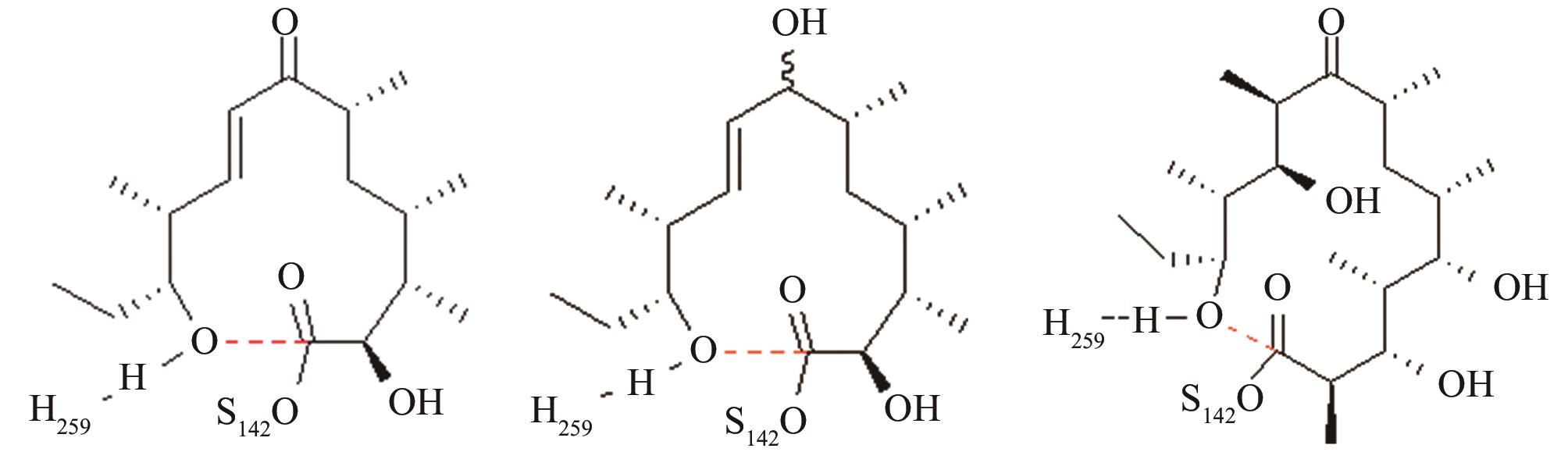
| 红霉素DEBS硫酯酶 | 底物上载后共价复合物水解和环化途径选择 |
与反应位点间隔4~5个化学键的碳原子对反应中心环化和水解活性结构的远程作用[ PRS结构参数:NH259-OH距离、OH-C距离 |
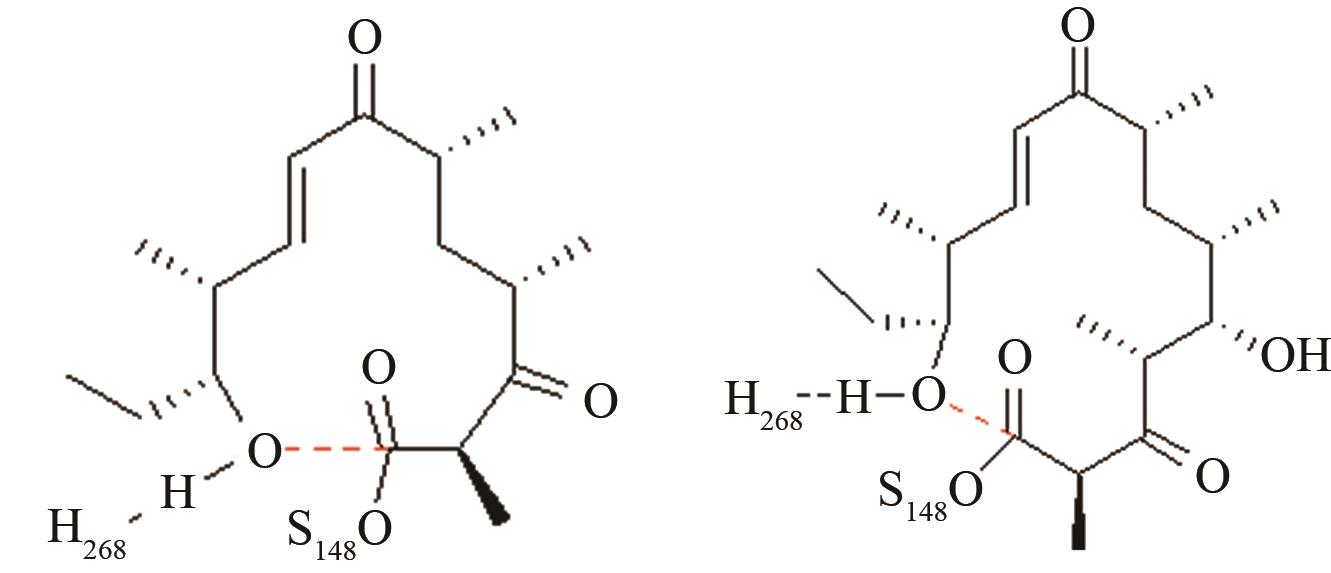
| 苦霉素硫酯酶 | 底物上载后共价复合物水解和环化途径选择 |
12元和14元大环成环对反应中心环化和水解活性结构的影响[ PRS结构参数:NH268-OH距离、OH-C距离 |
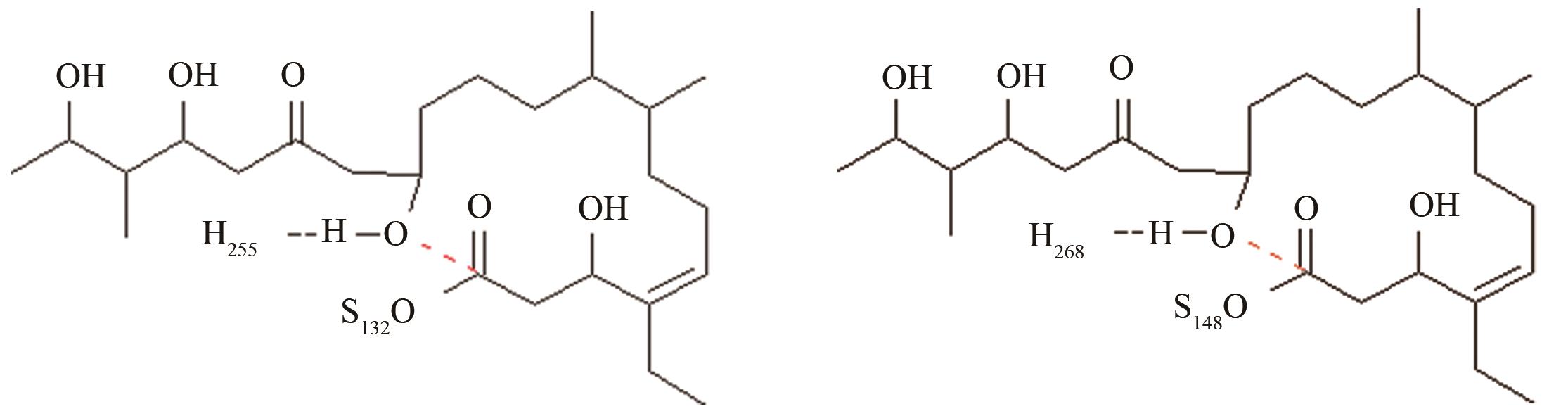
| 变构霉素硫酯酶/苦霉素硫酯酶 | 底物上载后共价复合物水解和环化途径选择 |
变构霉素生物合成时环化释放受阻,利用异源硫酯酶调整环化水解选择性,适配性分析[ 变构霉素硫酯酶PRS结构参数:NH255-OH距离、OH-C距离 苦霉素硫酯酶PRS结构参数:NH268-OH距离、OH-C距离 |
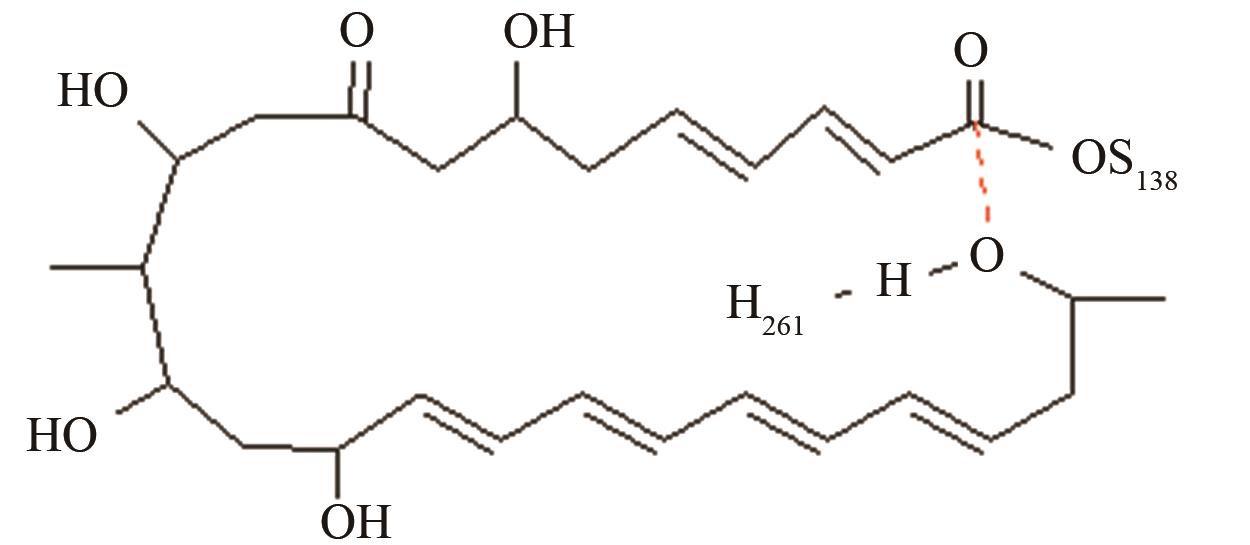
| 匹马霉素硫酯酶 | 底物上载后共价复合物水解和环化途径选择 |
与反应位点间隔11~13个化学键的碳原子修饰对反应中心环化和水解活性结构的远程作用[ PRS结构参数:NH261-OH距离、OH-C距离 |
| 醇脱氢酶KpADH | 质子偶合的负氢原子转移关键过渡态 |
高效突变体与野生型反应中心的活性构象分布变化[ PRS结构参数:HY164-O距离、HNADPH-C距离 |
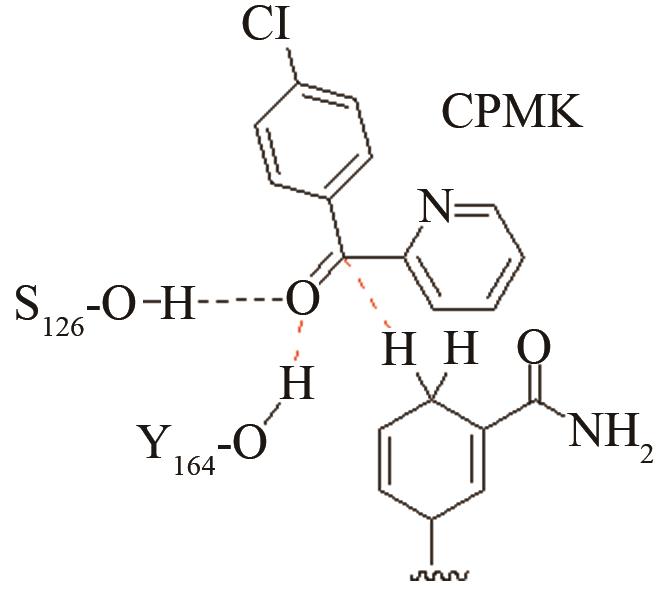
| 醇脱氢酶CpRCR | 质子偶合的负氢原子转移关键过渡态 |
高效突变体与野生型反应中心的活性构象分布变化[ PRS结构参数:HS46-O距离、HNADPH-C距离 |
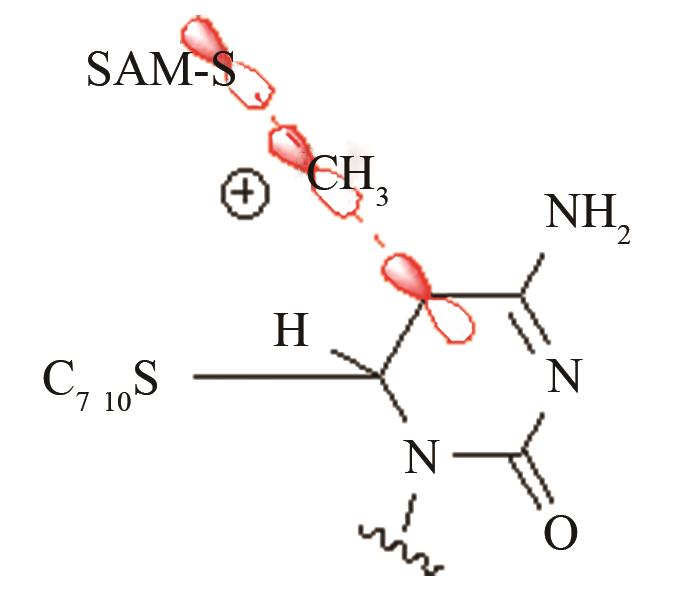
| DNA甲基化酶3A | SAM甲基转移到DNA-酶共价结合中间体 |
白血病高发的R882H远端突变对活性中心SAM甲基转移的影响[ PRS结构参数:CSAM-C距离 |
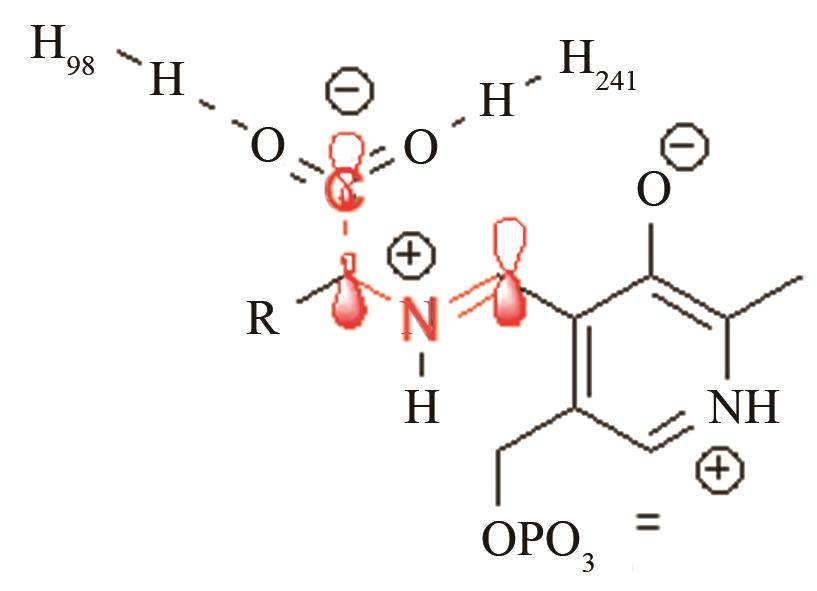
| 脱羧酶TyDC | PLP辅因子与底物的共价加合物C-C键断裂 |
通过H98、H251控制共价加合物的反应前线轨道布局[ PRS结构参数:CCαNSB角度,CCαNSBC4’二面角 |
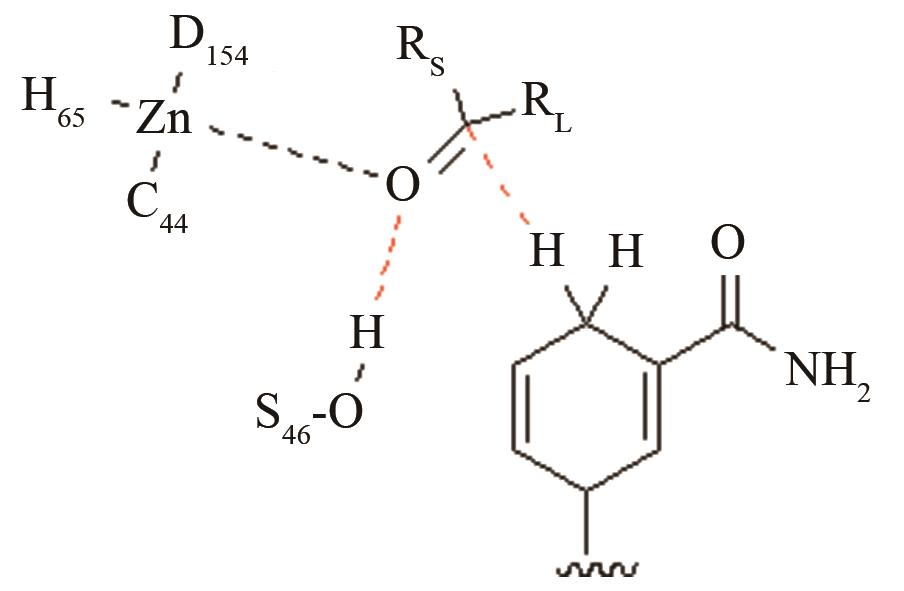
| 酰胺水解酶NiHyuC | 碳四面体氧负离子关键过渡态 |
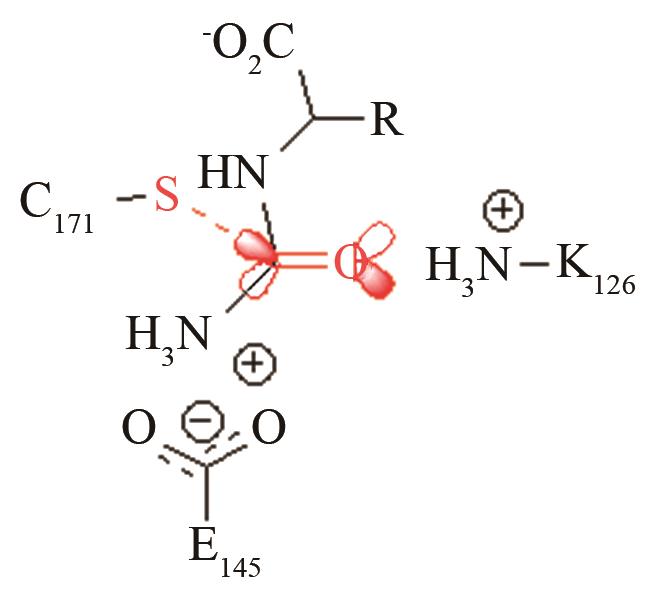
高效突变体对活化后巯基与被进攻的羧基反应前线轨道布局的影响[ PRS结构参数:SC171-C距离、SC171CO进攻角 |
| 普鲁兰多糖酶 | 取代反应SN2过渡态(碳氧键断裂) |
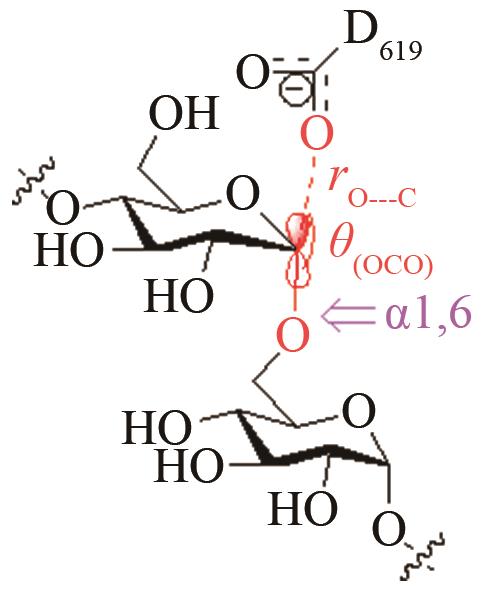
高效突变体对[O-C-O]取代反应前线轨道布局的影响[ PRS结构参数:OD619-C距离、OD619CO夹角 |
| MERS和SARS冠状病毒主蛋白水解酶 | 碳四面体氧负离子关键过渡态 |
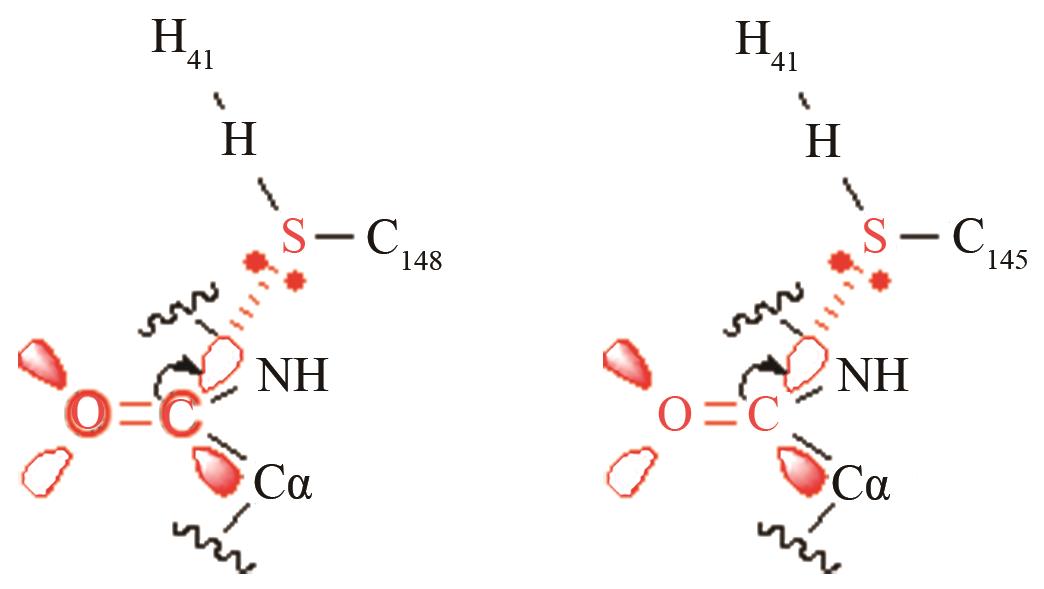
两种病毒主蛋白水解酶的关键反应前线轨道布局[ MERS病毒PRS结构参数:SC148-C距离、SC148CO进攻角 SARS病毒PRS结构参数:SC145-C距离、SC145CO进攻角 |
Tab. 1 Pre-reaction states of the studied enzymes, control points, and geometric parameters[95-98, 101,103,106,108,110,112,114]
| 研究对象 | 预反应态所选择的控制点 | 实例 |
|---|---|---|
| 红霉素DEBS硫酯酶 | 底物上载后共价复合物水解和环化途径选择 |
与反应位点间隔4~5个化学键的碳原子对反应中心环化和水解活性结构的远程作用[ PRS结构参数:NH259-OH距离、OH-C距离 |
| 苦霉素硫酯酶 | 底物上载后共价复合物水解和环化途径选择 |
12元和14元大环成环对反应中心环化和水解活性结构的影响[ PRS结构参数:NH268-OH距离、OH-C距离 |
| 变构霉素硫酯酶/苦霉素硫酯酶 | 底物上载后共价复合物水解和环化途径选择 |
变构霉素生物合成时环化释放受阻,利用异源硫酯酶调整环化水解选择性,适配性分析[ 变构霉素硫酯酶PRS结构参数:NH255-OH距离、OH-C距离 苦霉素硫酯酶PRS结构参数:NH268-OH距离、OH-C距离 |
| 匹马霉素硫酯酶 | 底物上载后共价复合物水解和环化途径选择 |
与反应位点间隔11~13个化学键的碳原子修饰对反应中心环化和水解活性结构的远程作用[ PRS结构参数:NH261-OH距离、OH-C距离 |
| 醇脱氢酶KpADH | 质子偶合的负氢原子转移关键过渡态 |
高效突变体与野生型反应中心的活性构象分布变化[ PRS结构参数:HY164-O距离、HNADPH-C距离 |
| 醇脱氢酶CpRCR | 质子偶合的负氢原子转移关键过渡态 |
高效突变体与野生型反应中心的活性构象分布变化[ PRS结构参数:HS46-O距离、HNADPH-C距离 |
| DNA甲基化酶3A | SAM甲基转移到DNA-酶共价结合中间体 |
白血病高发的R882H远端突变对活性中心SAM甲基转移的影响[ PRS结构参数:CSAM-C距离 |
| 脱羧酶TyDC | PLP辅因子与底物的共价加合物C-C键断裂 |
通过H98、H251控制共价加合物的反应前线轨道布局[ PRS结构参数:CCαNSB角度,CCαNSBC4’二面角 |
| 酰胺水解酶NiHyuC | 碳四面体氧负离子关键过渡态 |
高效突变体对活化后巯基与被进攻的羧基反应前线轨道布局的影响[ PRS结构参数:SC171-C距离、SC171CO进攻角 |
| 普鲁兰多糖酶 | 取代反应SN2过渡态(碳氧键断裂) |
高效突变体对[O-C-O]取代反应前线轨道布局的影响[ PRS结构参数:OD619-C距离、OD619CO夹角 |
| MERS和SARS冠状病毒主蛋白水解酶 | 碳四面体氧负离子关键过渡态 |
两种病毒主蛋白水解酶的关键反应前线轨道布局[ MERS病毒PRS结构参数:SC148-C距离、SC148CO进攻角 SARS病毒PRS结构参数:SC145-C距离、SC145CO进攻角 |
| 20 | ZHOU Huaxiang, WLODEK S T, MCCAMMON J A. Conformation gating as a mechanism for enzyme specificity[J]. Proceedings of the National Academy of Sciences of the United States of America, 1998, 95(16): 9280-9283. |
| 21 | JOHNSON K A. Role of induced fit in enzyme specificity: a molecular forward/reverse switch[J]. Journal of Biological Chemistry, 2008, 283(39): 26297-26301. |
| 22 | BENKOVIC S J, HAMMES G G, HAMMES-SCHIFFER S. Free-energy landscape of enzyme catalysis[J]. Biochemistry, 2008, 47(11):3317-3321. |
| 23 | WEI Guanghong, XI Wenhui, NUSSINOV R, et al. Protein ensembles: how does nature harness thermodynamic fluctuations for life? The diverse functional roles of conformational ensembles in the cell[J]. Chemical Reviews, 2016, 116(11): 6516-6551. |
| 24 | BUCHHOLZ K, KASCHE V, BORNSCHEUER U T. Biocatalysts and enzyme technology[M]. Wiley-VCH, Weihheim, Germany, 2005. |
| 25 | ARNOLD F H. Combinatorial and computational challenges for biocatalyst design[J]. Nature, 2001, 409(6817): 253-257. |
| 26 | BURTON S G, COWAN D A, WOODLEY J M. The search for the ideal biocatalyst[J]. Nature Biotechnology, 2002, 20(1): 37-45. |
| 27 | KRAUT D A, CARROLL K S, HERSCHLAG D. Challenges in enzyme mechanism and energetics[J]. Annual Review of Biochemistry, 2003, 72: 517-571. |
| 28 | RADZICKA A, WOLFENDEN R. A proficient enzyme[J]. Science, 1995, 267(5194): 90-93. |
| 29 | ZHANG Xiyun, HOUK K N. Why enzymes are proficient catalysts: beyond the Pauling paradigm[J]. Accounts of Chemical Research, 2005, 38(5): 379-385. |
| 30 | TANTILLO D J, CHEN Jiangang, HOUK K N. Theozymes and compuzymes: theoretical models for biological catalysis[J]. Current Opinion in Chemical Biology, 1998, 2(6): 743-750. |
| 31 | KISS G, ÇELEBI-ÖLÇÜM N, MORETTI R, et al. Computational enzyme design[J]. Angewandte Chemie International Edition, 2013, 52(22): 5700-5725. |
| 32 | FRUSHICHEVA M P, MILLS M J, SCHOPF P, et al. Computer aided enzyme design and catalytic concepts[J]. Current Opinion in Chemical Biology, 2014, 21: 56-62. |
| 33 | ANDERSON V E. Quantifying energetic contributions to ground state destabilization[J]. Archives of Biochemistry and Biophysics, 2005, 433(1): 27-33. |
| 34 | SCHRAMM V L. Enzymatic transition states and transition state analog design[J]. Annual Review of Biochemistry, 1998, 67: 693-720. |
| 35 | KOLESNIKOVÁ L, LEÓN I, ALONSO E R, et al. Laser ablation assists cyclization reactions of hydantoic acid: a proof for the near-attack conformation theory?[J]. The Journal of Physical Chemistry Letters, 2019, 10(6): 1325-1330. |
| 36 | CROSS P J, ALLISON T M, DOBSON R C J, et al. Engineering allosteric control to an unregulated enzyme by transfer of a regulatory domain[J]. Proceedings of the National Academy of Sciences of the United States of America, 2013, 110(6): 2111-2116. |
| 37 | ALHAMBRA C, GAO Jiali, CORCHADO J C, et al. Quantum mechanical dynamical effects in an enzyme-catalyzed proton transfer reaction[J]. Journal of the American Chemical Society, 1999, 121(10): 2253-2258. |
| 38 | RIBEIRO J M L, TSAI Sui-ting, PRAMANIK D, et al. Kinetics of ligand-protein dissociation from all-atom simulations: are we there yet?[J]. Biochemistry, 2019, 58(3): 156-165. |
| 39 | BARON R, MCCAMMON J A. Molecular recognition and ligand association[J]. Annual Review of Physical Chemistry, 2013, 64: 151-175. |
| 40 | AGARWAL V, LIN S, LUKK T, et al. Structure of the enzyme-acyl carrier protein (ACP) substrate gatekeeper complex required for biotin synthesis[J]. Proceedings of the National Academy of Sciences of the United States of America, 2012, 109(43): 17406-17411. |
| 41 | YING Hanxiao, WANG Jing, SHI Ting, et al. Engineering of lysine cyclodeaminase conformational dynamics for relieving substrate and product inhibitions in the biosynthesis of L-pipecolic acid[J]. Catalysis Science & Technology, 2019, 9(2): 398-405. |
| 42 | BUTTON D K. Biochemical basis for whole-cell uptake kinetics: specific affinity, oligotrophic capacity, and the meaning of the Michaelis constant[J]. Applied and Environmental Microbiology, 1991, 57(7): 2033-2038. |
| 43 | LEVINE R D. Molecular reaction dynamics[M]. Cambridge, U.K: Cambridge University Press, 2005. |
| 44 | GRUEBELE M, ZEWAIL A H. Ultrafast reaction dynamics[J]. Berichte der Bunsengesellschaft für Physikalische Chemie, 1990, 94(11): 1210-1218. |
| 45 | LAIDLER K J, KING M C. Development of transition-state theory[J]. The Journal of Physical Chemistry, 1983, 87(15): 2657-2664. |
| 46 | OSUNA S, JIMÉNEZ-OSÉS G, NOEY E L, et al. Molecular dynamics explorations of active site structure in designed and evolved enzymes[J]. Accounts of Chemical Research, 2015, 48(4): 1080-1089. |
| 47 | LIGHTSTONE F C, BRUICE T C. Ground state conformations and entropic and enthalpic factors in the efficiency of intramolecular and enzymatic reactions (I): Cyclic anhydride formation by substituted glutarates, succinate, and 3,6-endoxo-Δ 4-tetrahydrophthalate monophenyl esters[J]. Journal of the American Chemical Society, 1996, 118(11): 2595-2605. |
| 48 | HUR S, BRUICE T C. The near attack conformation approach to the study of the chorismate to prephenate reaction[J]. Proceedings of the National Academy of Sciences of the United States of America, 2003, 100(21): 12015-12020. |
| 49 | BRUICE T C. A view at the millennium: the efficiency of enzymatic catalysis[J]. Accounts of Chemical Research, 2002, 35(3): 139-148. |
| 50 | LAU E Y, BRUICE T C. Importance of correlated motions in forming highly reactive near attack conformations in catechol O-methyltransferase[J]. Journal of the American Chemical Society, 1998, 120(48): 12387-12394. |
| 51 | LAU E Y, BRUICE T C. Active site dynamics of the HhaI methyltransferase: insights from computer simulation[J]. Journal of Molecular Biology, 1999, 293(1): 9-18. |
| 52 | TORRES R A, SCHIØTT B, BRUICE T C. Molecular dynamics simulations of ground and transition states for the hydride transfer from formate to NAD+ in the active site of formate dehydrogenase[J]. Journal of the American Chemical Society, 1999, 121(36): 8164-8173. |
| 53 | RADKIEWICZ J L, BROOKS C L. Protein dynamics in enzymatic catalysis: exploration of dihydrofolate reductase[J]. Journal of the American Chemical Society, 2000, 122(2): 225-231. |
| 54 | LAITINEN T, ROUVINEN J, PERÄKYLÄ M. Inversion of the roles of the nucleophile and acid/base catalysts in the covalent binding of epoxyalkyl xyloside inhibitor to the catalytic glutamates of endo-1, 4-β-xylanase (XYNII): A molecular dynamics study[J]. Protein Engineering, Design and Selection, 2000, 13(4): 247-252. |
| 55 | LUO Jia, BRUICE T C. Dynamic structures of horse liver alcohol dehydrogenase (HLADH): results of molecular dynamics simulations of HLADH-NAD+-PhCH2OH, HLADH-NAD+-PhCH2O-, and HLADH-NADH-PhCHO[J]. Journal of the American Chemical Society, 2001, 123(48): 11952-11959. |
| 56 | REDDY S Y, KAHN K, ZHENG Yajun, et al. Protein engineering of nitrile hydratase activity of papain: molecular dynamics study of a mutant and wild-type enzyme[J]. Journal of the American Chemical Society, 2002, 124(44): 12979-12990. |
| 57 | SCHIØTT B, BRUICE T C. Reaction mechanism of soluble epoxide hydrolase: insights from molecular dynamics simulations[J]. Journal of the American Chemical Society, 2002, 124(49): 14558-14570. |
| 58 | MAZUMDER-SHIVAKUMAR D, BRUICE T C. Molecular dynamics studies of ground state and intermediate of the hyperthermophilic indole-3-glycerol phosphate synthase[J]. Proceedings of the National Academy of Sciences of the United States of America, 2004, 101(40): 14379-14384. |
| 59 | BRUICE T C. Computational approaches: reaction trajectories, structures, and atomic motions. enzyme reactions and proficiency[J]. Chemical Reviews, 2006, 106(8): 3119-3139. |
| 60 | SENN H M, THIEL W. QM/MM methods for biomolecular systems[J]. Angewandte Chemie International Edition, 2009, 48(7): 1198-1229. |
| 61 | CLAEYSSENS F, RANAGHAN K E, MANBY F R, et al. Multiple high-level QM/MM reaction paths demonstrate transition-state stabilization in chorismate mutase: correlation of barrier height with transition-state stabilization[J]. Chemical Communications, 2005(40): 5068-5070. |
| 62 | ZHANG Xiaodong, Zhang Xiaohua, BRUICE T C. A definitive mechanism for chorismate mutase[J]. Biochemistry, 2005, 44(31): 10443-10448. |
| 63 | GIRALDO J, ROCHE D, ROVIRA X, et al. The catalytic power of enzymes: conformational selection or transition state stabilization?[J]. FEBS Letters, 2006, 580(9): 2170-2177. |
| 64 | RYDE U. How many conformations need to be sampled to obtain converged QM/MM energies? the curse of exponential averaging[J]. Journal of Chemical Theory and Computation, 2017, 13(11): 5745-5752. |
| 65 | COOPER A M, KÄSTNER J. Averaging techniques for reaction barriers in QM/MM simulations[J]. ChemPhysChem, 2014, 15(15): 3264-3269. |
| 66 | LI Yanwei, ZHANG Ruiming, DU Likai, et al. Catalytic mechanism of C-F bond cleavage: insights from QM/MM analysis of fluoroacetate dehalogenase[J]. Catalysis Science & Technology, 2016, 6(1): 73-80. |
| 67 | DIETERICH J M, WERNER H J, MATA R A, et al. Reductive half-reaction of aldehyde oxidoreductase toward acetaldehyde: ab initio and free energy quantum mechanical/molecular mechanical calculations[J]. Journal of Chemical Physics, 2010, 132(3):035101. |
| 1 | CANTON B, LABNO A, ENDY D. Refinement and standardization of synthetic biological parts and devices[J]. Nature Biotechnology, 2008, 26(7): 787-793. |
| 2 | 崔颖璐, 吴边. 符合工程化需求的生物元件设计[J]. 中国科学院院刊, 2018, 33(11): 1150-1157. |
| CUI Yinglu, WU Bian. Biological components design for engineering requirements[J]. Bulletin of Chinese Academy of Sciences, 2018, 33(11): 1150-1157. | |
| 3 | ZEYMER C, HILVERT D. Directed evolution of protein catalysts[J]. Annual Review of Biochemistry, 2018, 87: 131-157. |
| 4 | REETZ M T. KAHAKEAW D, LOHMER R. Addressing the numbers problem in directed evolution[J]. ChemBioChem, 2008, 9(11):1797-1804. |
| 5 | HUANG P S, BOYKEN S E, BAKER D. The coming of age of de novo protein design[J]. Nature, 2016, 537(7620): 320-327. |
| 6 | KLEPEIS J L, LINDORFF-LARSEN K, DROR R O, et al. Long-timescale molecular dynamics simulations of protein structure and function[J]. Current Opinion in Structural Biology, 2009, 19(2): 120-127. |
| 7 | AHMADI S, BARRIOS HERRERA L, CHEHELAMIRANI M, et al. Multiscale modeling of enzymes: QM-cluster, QM/MM, and QM/MM/MD: a tutorial review[J]. International Journal of Quantum Chemistry, 2018, 118(9): e25558. |
| 8 | SALOMON-FERRER R, CASE D A, WALKER R C. An overview of the Amber biomolecular simulation package[J]. Wiley Interdisciplinary Reviews: Computational Molecular Science, 2013, 3(2): 198-210. |
| 9 | HOUK K N, LIU Fang. Holy grails for computational organic chemistry and biochemistry[J]. Accounts of Chemical Research, 2017, 50(3): 539-543. |
| 10 | KRYLOV A, WINDUS T L, BARNES T, et al. Perspective: Computational chemistry software and its advancement as illustrated through three grand challenge cases for molecular science[J]. The Journal of Chemical Physics, 2018, 149(18): 180901. |
| 11 | GRIMME S, SCHREINER P R. Computational chemistry: The fate of current methods and future challenges[J]. Angewandte Chemie International Edition, 2018, 57(16): 4170-4176. |
| 68 | LONSDALE R, REETZ M T. Reduction of α, β-unsaturated ketones by old yellow enzymes: mechanistic insights from quantum mechanics/molecular mechanics calculations[J]. Journal of the American Chemical Society, 2015, 137(46): 14733-14742. |
| 69 | VASILEVSKAYA T, KHRENOVA M G, NEMUKHIN A V, et al. Methodological aspects of QM/MM calculations: a case study on matrix metalloproteinase-2[J]. Journal of Computational Chemistry, 2016, 37(19): 1801-1809. |
| 70 | ASTRUC D. Organometallic chemistry and catalysis[M]. New York: Springer, 2007. |
| 71 | ROONEY J J. The extended Eyring kinetic equation and the compensation effect in catalysis[J]. Journal of Molecular Catalysis A: Chemical, 1998, 129(2/3): 131-134. |
| 72 | KOZUCH S, SHAIK S. How to conceptualize catalytic cycles? The energetic span model[J]. Accounts of Chemical Research, 2011, 44(2): 101-110. |
| 73 | AMATORE C, JUTAND A. Mechanistic and kinetic studies of palladium catalytic systems[J]. Journal of Organometallic Chemistry, 1999, 576(1/2): 254-278. |
| 74 | CHRISTIANSEN J A. The elucidation of reaction mechanisms by the method of intermediates in quasi-stationary concentrations[J]. Advances in Catalysis, 1953, 5: 311-353. |
| 75 | KOZUCH S, SHAIK S. A combined kinetic-quantum mechanical model for assessment of catalytic cycles: application to cross-coupling and Heck reactions[J]. Journal of the American Chemical Society, 2006, 128(10): 3355-3365. |
| 76 | CAMPBELL C T. Finding the rate-determining step in a mechanism: comparing DeDonder relations with the "degree of rate control"[J]. Journal of Catalysis, 2001, 204(2): 520-524. |
| 77 | KOZUCH S, SHAIK S. Kinetic-quantum chemical model for catalytic cycles: The Haber-Bosch process and the effect of reagent concentration[J]. The Journal of Physical Chemistry A, 2008, 112(26): 6032-6041. |
| 78 | KOZUCH S, SHAIK S. Defining the optimal inductive and steric requirements for a cross-coupling catalyst using the energetic span model[J]. Journal of Molecular Catalysis A: Chemical, 2010, 324(1/2): 120-126. |
| 79 | STEGELMANN C, ANDREASEN A, CAMPBELL C T. Degree of rate control: how much the energies of intermediates and transition states control rates[J]. Journal of the American Chemical Society, 2009, 131(23): 8077-8082. |
| 12 | CHOWDHURY R, MARANAS C D. From directed evolution to computational enzyme engineering-a review[J]. AIChE Journal, 2020, 66(3): e16847. |
| 13 | OSUNA S. The challenge of predicting distal active site mutations in computational enzyme design[J]. WIREs Computational Molecular Science, 2020, e1502. |
| 14 | MAGALHÃES R P, FERNANDES H S, SOUSA S F. Modelling enzymatic mechanisms with QM/MM approaches: Current status and future challenges[J]. Israel Journal of Chemistry, 2020, 60(7): 655-666. |
| 15 | PAULING L. Nature of forces between large molecules of biological interest[J]. Nature, 1948, 161(4097): 707-709. |
| 16 | MICHAELIS L, MENTEN M L, JOHNSON K A, et al. The original Michaelis constant: translation of the 1913 Michaelis-Menten paper[J]. Biochemistry, 2011, 50(39): 8264-8269. |
| 17 | TOMCZAK J M, WĘGLARZ-TOMCZAK E. Estimating kinetic constants in the Michaelis-Menten model from one enzymatic assay using Approximate Bayesian Computation[J]. FEBS Letters, 2019, 593(19): 2742-2750. |
| 18 | HOUK K N, LEACH A G, KIM S P, et al. Binding affinities of host-guest, protein-ligand, and protein-transition-state complexes[J]. Angewandte Chemie International Edition, 2003, 42(40): 4872-4897. |
| 19 | SCHRAMM V L. Enzymatic transition states and transition state analogues[J]. Current Opinion in Structural Biology, 2005, 15(6): 604-613. |
| 80 | ROMERO P A, ARNOLD F H. Exploring protein fitness landscapes by directed evolution[J]. Nature Reviews Molecular Cell Biology, 2009, 10(12): 866-876. |
| 81 | VOIGT C A, MAYO S L, ARNOLD F H, et al. Computational method to reduce the search space for directed protein evolution[J]. Proceedings of the National Academy of Sciences of the United States of America, 2001, 98(7): 3778-3783. |
| 82 | TRACEWELL C A, ARNOLD F H. Directed enzyme evolution: climbing fitness peaks one amino acid at a time[J]. Current Opinion in Chemical Biology, 2009, 13(1): 3-9. |
| 83 | UHE A, KOZUCH S, SHAIK S. Automatic analysis of computed catalytic cycles[J]. Journal of Computational Chemistry, 2011, 32(5): 978-985. |
| 84 | SOLEL E, TARANNAM N, KOZUCH S. Catalysis: energy is the measure of all things[J]. Chemical Communications, 2019, 55(37): 5306-5322. |
| 85 | EINSIEDLER M, JAMIESON C S, MASKERI M A, et al. Fungal dioxygenase AsqJ is promiscuous and bimodal: substrate-directed formation of quinolones versus quinazolinones[J]. Angewandte Chemie International Edition, 2021, 60(15): 8297-8302. |
| 86 | LIU Fengjiao, YANG Zhongyue, YU Yanmin, et al. Bimodal Evans-Polanyi relationships in dioxirane oxidations of sp3 C—H: non-perfect synchronization in generation of delocalized radical intermediates[J]. Journal of the American Chemical Society, 2017, 139(46):16650-16656. |
| 87 | WILLIAMS G J. Engineering polyketide synthases and nonribosomal peptide synthetases[J]. Current Opinion in Structural Biology, 2013, 23(4): 603-612. |
| 88 | LARSEN E M, WILSON M R, TAYLOR R E. Conformation-activity relationships of polyketide natural products[J]. Natural Product Reports, 2015, 32(8): 1183-1206. |
| 89 | MEIER J L, BURKART M D. The chemical biology of modular biosynthetic enzymes[J]. Chemical Society Reviews, 2009, 38(7): 2012-2045. |
| 90 | SMITH S, TSAI S C. The type I fatty acid and polyketide synthases: a tale of two megasynthases[J]. Natural Product Reports, 2007, 24(5): 1041-1072. |
| 91 | KEATINGE-CLAY A T. The structures of type I polyketide synthases[J]. Natural Product Reports, 2012, 29(10): 1050-1073. |
| 92 | KHOSLA C, TANG Yingyuan, CHEN A Y, et al. Structure and mechanism of the 6-deoxyerythronolide B synthase[J]. Annual Reviews of Biochemistry, 2007, 76: 195-221. |
| 93 | ALDRICH C C, VENKATRAMAN L, SHERMAN D H, et al. Chemoenzymatic synthesis of the polyketide macrolactone 10-deoxymethynolide[J]. Journal of the American Chemical Society, 2005, 127(25): 8910-8911. |
| 94 | HE Weiguo, WU Jiaquan, KHOSLA C, et al. Macrolactonization to 10-deoxymethynolide catalyzed by the recombinant thioesterase of the picromycin/methymycin polyketide synthase[J]. Bioorganic & Medicinal Chemistry Letters, 2006, 16(2): 391-394. |
| 95 | CHEN Xiongping, SHI Ting, WANG Xiaolei, et al. Theoretical studies on the mechanism of thioesterase-catalyzed macrocyclization in erythromycin biosynthesis[J]. ACS Catalysis, 2016, 6(7):4369-4378. |
| 96 | SHI Ting, LIU Lanxuan, TAO Wentao, et al. Theoretical studies on the catalytic mechanism and substrate diversity for macrocyclization of pikromycin thioesterase[J]. ACS Catalysis, 2018, 8(5): 4323-4332. |
| 97 | LIU Lei, TAO Wentao, BAI Linquan, et al. Why does tautomycetin thioesterase prefer hydrolysis to macrocyclization? Theoretical study on its catalytic mechanism[J]. Catalysis Science & Technology, 2019, 9(22): 6391-6403. |
| 98 | FAN Shuobing, WANG Rufan, LI Chen, et al. Insight into structural characteristics of protein-substrate interaction in pimaricin thioesterase[J]. International Journal of Molecular Sciences, 2019, 20(4):877. |
| 99 | HALL M, BOMMARIUS A S. Enantioenriched compounds via enzyme-catalyzed redox reactions[J]. Chemical Reviews, 2011, 111(7): 4088-4110. |
| 100 | KULIG J, SIMON R C, ROSE C A, et al. Stereoselective synthesis of bulky 1,2-diols with alcohol dehydrogenases[J]. Catalysis Science & Technology, 2012, 2(8): 1580. |
| 101 | ZHOU Jieyu, WANG Yue, XU Guochao, et al. Structural insight into enantioselective inversion of an alcohol dehydrogenase reveals a "polar gate" in stereorecognition of diaryl ketones[J]. Journal of the American Chemical Society, 2018, 140(39): 12645-12654. |
| 102 | NOEY E L, TIBREWAL N, JIMÉNEZ-OSÉS G, et al. Origins of stereoselectivity in evolved ketoreductases[J]. Proceedings of the National Academy of Sciences of the United States of America, 2015, 112(51): E7065-E7072. |
| 103 | NIE Yao, WANG Shanshan, XU Yan, et al. Enzyme engineering based on X-ray structures and kinetic profiling of substrate libraries: alcohol dehydrogenases for stereospecific synthesis of a broad range of chiral alcohols[J]. ACS Catalysis, 2018, 8(6): 5145-5152. |
| 104 | MESSERSCHMIDT D M, KNOWLES B B, SOLTER D. DNA methylation dynamics during epigenetic reprogramming in the germline and preimplantation embryos[J]. Genes & Development, 2014, 28(8): 812-828. |
| 105 | YANG Li, LIU Yanan, ZHANG Na, et al. Novel impact of the DNMT3A R882H mutation on GSH metabolism in a K562 cell model established by TALENs[J]. Oncotarget, 2017, 8(18): 30395-30409. |
| 106 | LIU Lanxuan, SHI Ting, HOUK K N, et al. Understanding the R882H mutation effects of DNA methyltransferase DNMT3A: a combination of molecular dynamics simulations and QM/MM calculations[J]. RSC Advance, 2019, 9:31425-31434. |
| 107 | DUNATHAN H C. Conformation and reaction specificity in pyridoxal phosphate enzymes[J]. Proceedings of the National Academy of Sciences of the United States, 1966, 55(4): 712-716. |
| 108 | NI Jie, XU Guochao, DAI Wei, et al. Hyperconjugation promoted by hydrogen bonding between His98/His241 and a carboxyl group contributes to tyrosine decarboxylase catalysis[J]. Catalysis Science & Technology, 2019, 9(22): 6222-6226. |
| 109 | HERAS-YAZQUEZ F, CLEMENTE-JIMENEZ J, MARTINEZ-RODRIGUEZ S, et al. Optically pure α-amino acids production by the "hydantoinase process"[J]. Recent Patents Biotechnology, 2008, 2(1): 35-46. |
| 110 | LIU Yafei, XU Guocao, ZHOU Jieyu, et al. Structure-guided engineering of D-carbamoylase reveals a key loop at substrate entrance tunnel[J]. ACS Catalysis, 2020, 10(21): 12393-12402. |
| 111 | BERTOLDO C, ANTRANIKIAN G. Starch-hydrolyzing enzymes from thermophilic archaea and bacteria[J]. Current Opinion in Chemical Biology, 2002, 6(2): 151-160. |
| 112 | WANG Xinye, JING Xiaoran, Deng Yi, et al. Evolutionary coupling saturation mutagenesis: coevolution‐guided identification of distant sites influencing Bacillus naganoensis pullulanase activity[J]. FEBS Letters, 2020, 594(5): 799-812. |
| 113 | TARANTO A G, CARVALHO P, AVERY M A. QM/QM studies for Michael reaction in coronavirus main protease (3CLPro)[J]. Journal of Molecular Graphics and Modelling, 2008, 27(3): 275-285. |
| 114 | WANG Hao, He Shuai, DENG Weilong, et al. Comprehensive insights into the catalytic mechanism of middle east respiratory syndrome 3C-like protease and severe acute respiratory syndrome 3C-like protease[J]. ACS Catalysis, 2020, 10: 5871-5890. |
| [1] | Chengxin ZHANG. Challenges and opportunities in text mining-based protein function annotation [J]. Synthetic Biology Journal, 2025, (): 1-14. |
| [2] | Qian LI, E. Ferrell James, Yuping CHEN. Cytoplasmic concentration: an old question and a new parameter in cell biology [J]. Synthetic Biology Journal, 2025, (): 1-18. |
| [3] | Baiyi Jiang, Long Qian. Application and prospect of live cell molecular recorder in cell lineage tracing [J]. Synthetic Biology Journal, 2025, (): 1-17. |
| [4] | WEN Yanhua, LIU Hedong, CAO Chunlai, WU Ruibo. Applications of protein engineering in pharmaceutical industry [J]. Synthetic Biology Journal, 2025, 6(1): 65-86. |
| [5] | DONG Ying, MA Mengdan, HUANG Weiren. Progress in the miniaturization of CRISPR-Cas systems [J]. Synthetic Biology Journal, 2025, 6(1): 105-117. |
| [6] | Rixin ZHANG, Xiao-jun TIAN. The Cell 'Economics Paradox' in Synthetic Gene Circuits [J]. Synthetic Biology Journal, 2025, (): 1-14. |
| [7] | Xiaotian TAN, Ruihan LI, Hui YANG. Antibody probes in biomolecular sensing: transitioning from carbon-based computing to silicon-based computing [J]. Synthetic Biology Journal, 2025, (): 1-9. |
| [8] | SHAO Mingwei, SUN Simian, YANG Shimao, CHEN Guoqiang. Bioproduction based on extremophiles [J]. Synthetic Biology Journal, 2024, 5(6): 1419-1436. |
| [9] | LIU Jianming, ZHANG Chijian, ZHANG Bing, ZENG Anping. Clostridium pasteurianum as an industrial chassis for efficient production of 1,3-propanediol: from metabolic engineering to fermentation and product separation [J]. Synthetic Biology Journal, 2024, 5(6): 1386-1403. |
| [10] | LIU Yining, PU Wei, YANG Jinxing, WANG Yu. Recent advances in the biosynthesis of ω-amino acids and lactams [J]. Synthetic Biology Journal, 2024, 5(6): 1350-1366. |
| [11] | ZHAO Liang, LI Zhenshuai, FU Liping, LYU Ming, WANG Shi’an, ZHANG Quan, LIU Licheng, LI Fuli, LIU Ziyong. Progress in biomanufacturing of lipids and single cell protein from one-carbon compounds [J]. Synthetic Biology Journal, 2024, 5(6): 1300-1318. |
| [12] | Kainan SONG, Liwen ZHANG, Chao WANG, Pingfang TIAN, Guangyue LI, Guohui PAN, Yuquan XU. Advances in small-molecule biopesticides and their biosynthesis [J]. Synthetic Biology Journal, 2024, (): 1-21. |
| [13] | Jinhang YI, Yulin TANG, Chunyu LI, Heyun WU, Qian MA, Xixian XIE. Applications and advances in the research of biosynthesis of amino acid derivatives as key ingredients in cosmetics [J]. Synthetic Biology Journal, 2024, (): 1-36. |
| [14] | . [J]. Synthetic Biology Journal, 2024, 5(5): 909-912. |
| [15] | ZHANG Shouqi, WANG Tao, KONG Yao, ZOU Jiasheng, LIU Yuanning, XU Zhengren. Chemoenzymatic synthesis of natural products: evolution of synthetic methodology and strategy [J]. Synthetic Biology Journal, 2024, 5(5): 913-940. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||